Genome-wide association and RNA-seq analyses reveal a potential gene related to linolenic acid in soybean seeds
- Published
- Accepted
- Received
- Academic Editor
- Paula Soares
- Subject Areas
- Biotechnology, Molecular Biology, Plant Science
- Keywords
- Soybean, Linolenic acid, GWAS, RNA-seq, MALDI-TOF IMS, Fatty acid biosynthesis
- Copyright
- © 2023 Qin et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2023. Genome-wide association and RNA-seq analyses reveal a potential gene related to linolenic acid in soybean seeds. PeerJ 11:e16138 https://doi.org/10.7717/peerj.16138
Abstract
Linolenic acid (LA) has poor oxidative stability since it is a polyunsaturated fatty acid. Soybean oil has a high LA content and thus has poor oxidative stability. To identify candidate genes that affect the linolenic acid (LA) content in soybean seeds, a genome-wide association study (GWAS) was performed with 1,060 soybean cultivars collected in China between 2019–2021 and which LA content was measured using matrix-assisted laser desorption/ionization time-of-flight imaging mass spectrometry (MALDI-TOF IMS). A candidate gene, GmWRI14, encoding an APETALA2 (AP2)-type transcription factor, was detected by GWAS in cultivars from all three study years. Multiple sequence alignments showed that GmWRI14 belongs to the plant WRI1 family. The fatty acid contents of different soybean lines were evaluated in transgenic lines with a copy of GmWRI14, control lines without GmWRI14, and the gmwri14 mutant. MALDI-TOF IMS revealed that GmWRI14 transgenic soybeans had a lower LA content with a significant effect on seed size and shape, whereas gmwri14 mutants had a higher LA content. compared to control. The RNA-seq results showed that GmWRI14 suppresses GmFAD3s (GmFAD3B and GmFAD3C) and GmbZIP54 expression in soybean seeds, leading to decreased LA content. Based on the RNA-seq data, yeast one-hybrid (Y1H) and qRT-PCR were performed to confirm the transcriptional regulation of FAD3s by GmWRI14. Our results suggest that FAD3 is indirectly regulated by GmWRI14, representing a new molecular mechanism of fatty acid biosynthesis, in which GmWRI14 regulates LA content in soybean seeds.
Introduction
Soybeans (Glycine max (L.) Merrill) are one of the most important oil crops grown worldwide, particularly in China, and soybean oil now accounts for more than 50% of the total vegetable oil consumption (Liu et al., 2020; Zhao et al., 2019). Approximately 5–11% of soybean oil contains linolenic acid (LA) (Zhang et al., 2018; Clemente & Cahoon, 2009). LA is an essential polyunsaturated fatty acid that acts at multiple levels to alter membrane fluidity and cause inflammation (De Groot et al., 2004; Chan, Bruce & McDonald, 1991). Soybean oil with a high LA concentration degrades easily at high temperatures during the frying cycle, causing a rancid odor (Tompkins, Maskan & Karataş, 1998; Yang et al., 2009; Marangoni et al., 2020). The n-3 double bonds in LA facilitate its combination with oxygen molecules, resulting in poor oxidative stability (Thomsen et al., 1999; Do et al., 2019). Decreasing LA content is important for soybean seed aging and for improving the oxidative stability of soybean oil (Singh et al., 2001). Therefore, investigating the genetic basis of LA content in soybeans has become an important goal for breeding soybeans with a low LA content that are resistant to high temperatures (Sivaraman et al., 2004). Some potential candidate genes associated with LA content in soybeans have been identified in previous genome-wide association analyses (GWAS) (Singh et al., 2001; Li et al., 2019), such as fatty acid desaturase-3 genes (FAD3). Previous studies showed that ω3-fatty acid desaturase increases LA levels (Lakhssassi et al., 2017; Wen et al., 2018; Yang et al., 2018). In addition, many transcription factors (TFs) regulate FAD3 expression and affect LA accumulation, such as bZIP transcription factor bZIP67, ABA INSENSITIVE3 (ABI3) and leafy cotyledon1 (LEC1). These genes are highly correlated with LA synthesis and may affect FAD3 expression or may indirectly influence linolenic acid content by cooperating with other TFs that regulate fatty acid synthesis. An important example is WRINKLED1 (WRI1), which is correlated with fatty acid content and lipid synthesis regulation in different plant seeds (Kong, Yuan & Ma, 2019; Kong et al., 2020). Previous studies have also found that LA levels were higher in wri1 mutant seeds, but with lower total oil volume (Tang et al., 2019; Kuczynski et al., 2020; Guo et al., 2020). However, these studies do not completely explain the function of WRI1, and additional potential target genes and cooperative TFs need to be explored. For instance, fatty acid desaturase (FAD2), Leafy Cotyledon1 (LEC1) and stearoy-acyl-carrier-protein desaturase (SAD) expression have closely related to WRI1, potentially implicating WRI1 in the regulation of different fatty acids in soybean seeds (Zhang et al., 2019; Kong & Ma, 2018; Chen et al., 2018).
In our study, we used GWAS to screen candidate genes associated with LA content in soybeans from 2019–2021 to, and discovered a new candidate gene, the APETALA2 (AP2)-type transcription factor GmWRI14. RNA-seq and qRT-PCR results indicated that GmWRI14 suppressed the expression of both GmFAD3s and bZIP54 in soybean seeds, with the inhibition of GmFAD3s leading to lower LA content. This study not only lays a theoretical foundation for improving the seed oil quality of soybeans but also provides effective strategies and genetic resources to break the mutual constraints between oil content and quality.
Materials and Methods
Determining fatty acid content of different soybean cultivars
A total of 1,060 soybean lines from China were collected to determine their fatty acid content. To obtain more accurate phenotypic data for the GWAS, soybean lines were planted in three regions of China representing typical climates (subtropical, tropical, and temperate zones). The seeds were planted at the following locations: 2019–2020 on 124.65°E, 43.14°N; in 2020–2021 on 114.03°E, 38.29°N; in 2021–2022 on 113.01°E, 24.17°N. After harvesting, the seeds of different soybean lines were dried and the fatty acid content was measured. Five fatty acids (linolenic acid, stearic acid, oleic acid, palmitic acid, and linoleic acid) in soybean cultivar seeds were determined using matrix-assisted laser desorption/ionization time-of-flight imaging mass spectrometry (MALDI-TOF IMS). The correlation coefficients of fatty acids were calculated using SPSS software (SPSS Inc., Chicago, IL, USA). Each calculation was repeated thrice to ensure accuracy.
Genotyping and GWAS analysis
We used the CTAB method to extract genomic DNA from fresh leaves of each soybean line (Porebski, Bailey & Baum, 1997). The Beijing Biomarker Biotechnology Co. Ltd. (Beijing, China) provided the sequencing services. A total of 2,935,226 SNP markers with a minor allele frequency (MAF) > 0.06, obtained from genotyping, were screened for GWAS using the fastlmmc model. GWAS was used to analyze the association between the genotype and phenotype datasets. The Haploview 4.2 software was used to construct a Manhattan map. The threshold value was set to –log(p) > 4.20. The plink2 software was used to calculate the LD decay distances. The NR database and Swiss-Prot were used to search for candidate genes in the 650 kb genomic region of the important SNPs.
Plant transformation
The GmWRI14 gene from the soybean cultivar 010a (approval number 2012010) was cloned into the BamHI-SacI site of plasmid pTF101 named pTF101- GmWRI14-Flag, which was induced by the CaMV35S promoter, and the target gene was terminated by the NOS terminator. The recombinant plasmid was transformed into calli using Agrobacterium tumefaciens strain LBA4404 (Holsters et al., 1978). Individual T0 plant lines were established in a greenhouse, and three independent transgenic soybean lines with the same plasmid were harvested: GmWRI14-1, GmWRI14-2, and GmWRI14-3.
The generation and identification of GmWRI14 gene-edited lines
Following a previous protocol, the novel CRISPR/Cas9 Vector pGES201 was used for soybean genome editing (Do et al., 2019). CRISPRdirect (http://crispr.dbcls.jp) was used to design the target site sequence based on the exon sequence of the GmWRI14 gene in the soybean genome. The GmWRI14 gene from soybean cultivar 010a (approval number 2012010) was cloned into the BamHI-SacI site of plasmid pTF101 named pTF101-GmWRI14-3×Flag, which was induced by the pM4 promoter, and the target gene was terminated by the NOS terminator. The CRISPR/Cas9 expression vector was constructed using the method described by Bai et al. (2020). The CRISPR/Cas9 expression vector was transformed into Agrobacterium tumefaciens LBA4404 using Agrobacterium-mediated soybean cotyledon node genetic transformation method (Holsters et al., 1978). The EasyPure® Plant Genome DNA Kit (TransGen, Beijing, China) was used according to the manufacturer’s instructions to extract total genomic DNA from the seed samples of the putative mutants. The T0–T2 generation seeds and control seeds of the transgenic plants were planted in a greenhouse under a 16/8-h light/darkness cycle. When the T1–T2 generation plants grew the first triple-compound leaf, they were screened for glyphosate resistance. The genomic DNA of genetically modified soybeans and controls was extracted using Kangwei Century Company’s new plant genome DNA extraction kit, and PCR amplification was performed using DNA as a template to detect positive plants. Positive plants were selected and the PCR products were sent to a sequencing company for sequencing. DSDecodeM was used to analyze the sequences of the T3 generation plants to characterize CRISPR/Cas9-induced mutations. Successfully edited strains were identified using sequence peaks and were aligned to the reference sequence. Heterozygous mutants exhibited overlapping peaks near the target site, whereas homozygous mutants exhibited a single peak at the target site. Three independent gmwri14 soybean mutant lines containing the same plasmid were harvested. They are gmwri14-1, gmwri14-2, and gmwri14-3.
The RNA-seq data analysis
To analyze the transduction pathways induced by GmWRI14, the total RNA of each sample (GmWRI14 transgenic soybeans, CK, and gmwri14 mutant) was isolated from the seeds for RNA sequencing, with three biological and three technical replicates, using the Super total RNA® extraction kit (TaKaRa, San Jose, CA, USA). A total of 2 μg RNA was obtained from each plant sample for the RNA sequencing. RNA quality was assessed using Nanodrop 2000c (Thermo Fisher Scientific, Waltham, MA, USA). All transcriptome data were analyzed and calculated using methods outlined in previous studies (Zhong et al., 2011; Sultan et al., 2012; Kumar et al., 2012). The RNA-sequencing depth was 10×, and the 2−ΔΔCt method was used for the fold change of gene expression (Livak & Schmittgen, 2001). The original data were deposited in the China National GenBank database (accession number PRJNA435633). Each sample had an average of 5.68 GB of data for a total of 102.11 GB of sequencing data. We used bioinformatics software (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/) to improve the data quality, resulting in 41.66 GB of clean data. The clean data of all samples reached 6 GB, and the base percentage of Q30 was more than 93%. The statistical power of this experimental design was calculated in cluster Profiler Power with a q value 0.01. A KEGG statistical analysis (Kyoto Encyclopedia of Genes and Genome) and Gene Ontology analysis (http://www.geneontology.org/) were performed to identify the functions of the differentially expressed genes (DEGs).
Gene expression analysis by qRT-PCR
After 25 days of seedling growth, RNA samples were extracted from the seeds of soybean lines with low LA content and high LA content using an RNA extraction kit (TaKaRa, San Jose, CA, USA). Using overexpressed GmWRI14 soybeans, gmwri14 mutants, and control soybean seeds as materials, total RNA was extracted and reverse transcribed into cDNA at 10, 11, 12, 13, and 14 days after podding, using 24-h sampling (T0, T4, T8, T12, T16, T20, and T24 soybean seeds, respectively). qRT-PCR primers for bZIP54, GmFAD3C and GmFAD3B, were used to perform a 24-h expression regulation analysis of the candidate target genes of GmWRI14. qRT-PCR was performed with three biological replicates, according to the MIQE guidelines (Bustin et al., 2009). Real-time fluorescence analysis was performed using a LightCycler (Roche, Basel, Switzerland). The lectin gene (Gene ID: 100775957) was used as an internal control (Qin et al., 2022). All primers used are listed in the additional files (Table S1).
Yeast one-hybrid (Y1H) assay
To test whether GmWRI14 could affect the expression of FAD3s by binding to the promoter region of FAD3s, both the FAD3s (GmFAD3B and GmFAD3C) and the GmbZIP54 promoter regions were amplified from soybean cultivar 010a (Approval number 2012010). The Y1H assay was performed according to the manufacturer’s instructions (Zhao et al., 2019).
Results
Identification of candidate gene GmWRI14 by GWAS
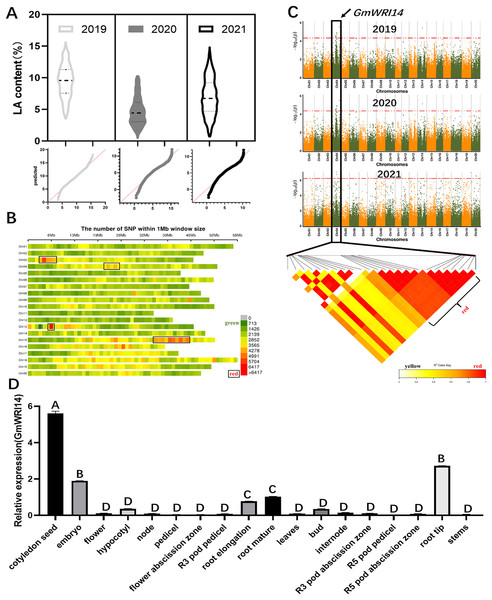
The LA content in 1,060 soybean lines collected from different regions of China between 2019 and 2021 ranged from 1.01% to 15.92%. These 3-year measurements suggest that there is a wide variation in LA content in the soybean population, the distribution is continuous, and the LA content conforms to a normal distribution (Fig. 1A). A total of 2,923,211 SNP markers were obtained from the dataset (Fig. 1B), of which 899,218 SNPs had an MAF > 6%. To identify potential candidate genes associated with LA, a GWAS was used to screen candidate genes using the fastlmmc model. These SNP markers were evaluated for their association with LA content. We identified 17 candidate genes associated with LA content from to 2019–2020, ten candidate genes from to 2020–2021, and ten candidate genes from to 2021–2022. Ten SNPs were associated with LA, all of which were located on chromosome 4. Glyma.04G127300.1, a candidate gene for LA, was detected by GWAS in each of the 3 years (Fig. 1C), indicating that Glyma.04G127300.1 is a candidate gene for LA. The results of the bioinformatics analysis showed that Glyma.04G127300.1 belongs to the plant WRI1 family containing two AP2/EREB domains; therefore, we named it GmWRI14. The expression of the GmWRI14 gene in the flowers, internodes, stems, and roots of the soybean plants was investigated using qRT-PCR. GmWRI14 was expressed in different organs of soybean plants but was mainly expressed in soybean seeds, with low levels of GmWRI14 expression detected in the other organs (Fig. 1D). The qRT-PCR results showed that the GmWRI14 gene expression was highly tissue-specific in soybeans. The raw qRT-PCR data are available in File S1.
Figure 1: Linolenic acid (LA) content and GWAS analysis from soybean germplasms collected from 2019–2021.
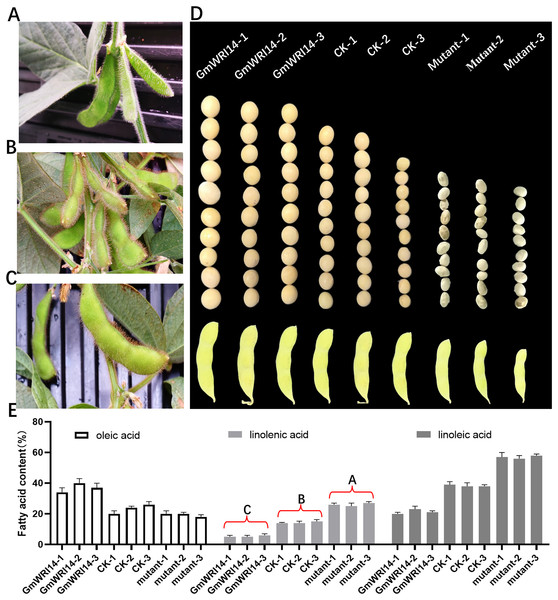
(A) The floating range of LA content of 1,060 soybean germplasms collected from 2019–2021. LA content conforms to normal distribution. In 2019, LA content ranged from 2.62–15.75%; in 2020, LA content ranged from 0.25–10.32%; and in 2021, LA ranged from 0.31–16.12%. (B) Density distribution map of SNP on 20 chromosomes of soybean. The green area indicates that the number of SNPs is less than 1,426. (C) GWAS was used to screen candidate genes related to LA from 2019–2021, using the fastlmmc model. Each point represents an SNP on each chromosome. The threshold line is the P-value considered for significance. (D) Analysis of expression of GmWRI14 in different tissues. GmWRI14 was mainly expressed in soybean seeds, with an expression of 5.8 ± 0.2. The uppercase letters in (D) represent significant differences (P < 0.01).The GmWRI14 gene from the soybean cultivar 010a (Approval number 2012010) was cloned into the BamHI-SacI site of the plasmid pTF101, named pTF101- GmWRI14-Flag (Fig. S1). GmWRI14 transgenic soybean and gmwri14 mutant were then produced (Figs. 2A–2D), and Southern blotting was used to detect GmWRI14 expression (Fig. S2), and MALDI-TOF MS was used to analyze the LA distribution in soybean seeds (Fig. S3). MALDI-TOF IMS results showed that the total fatty acid content of GmWRI14 transgenic soybean increased from 41.32% to 56.02% (Fig. 2E); however, the LA content of GmWRI14 transgenic soybean decreased from 12.24% to 6.12%. MALDI-TOF IMS results showed that, compared to soybean receptors (CK), the linolenic acid content in the gmwri14 mutant significantly increased from 21.01% to 23.92% (Fig. 2E). These results indicate that the expression of GmWRI14 and its ability to regulate specific genes are important for determining LA content in soybean seeds.
Figure 2: Phenotyping of CK, GmWRI14 transgenic lines, and gmwri14 mutant.
(A) Phenotype of gmwri14 mutant. (B) Phenotype of CK. (C) Phenotype of GmWRI14 transgenic soybean plants. (D) Phenotype of soybean seed from three varieties (CK, GmWRI14 transgenic lines, gmwri14 mutant). (E) LA content of different soybean lines. GmWRI14-1, GmWRI14-2 and GmWRI14-3 are three independent transgenic soybean lines with the same plasmid. gmwri14-1 mutant, gmwri14-2 mutant and gmwri14-3 mutant are three independent transgenic soybean lines with the same CRISPR/Cas9 expression vector. The uppercase letters in (E) represent significant differences (P < 0.01).Overexpression of GmWRI14 suppresses GmFAD3s and bZIP54 expression in soybean seeds
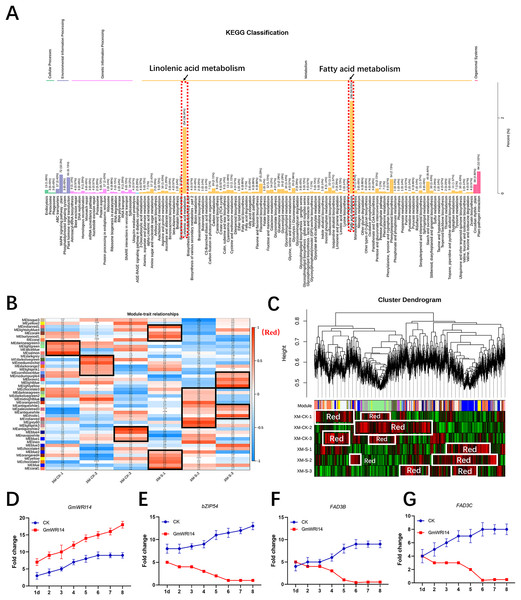
RNA-seq was used to verify the genes related to fatty acid synthesis that were induced by GmWRI14 in transgenic soybean seeds and control plants (CK, GmWRI14 transgenic lines, and gmwri14 mutant). RNA extracted from the seeds was used for RNA-seq to analyze the differentially expressed genes (DEGs) between the three varieties (CK, GmWRI14 transgenic lines, and gmwri14 mutant). The RNA-seq results showed that the expression of more than 50 genes related to fatty acid synthesis changed significantly. The KEGG and GO analyses revealed that the 891 identified DEGs were mainly involved in fatty acid and linolenic acid metabolism (Fig. 3A). Weighted correlation network analysis (WGCNA) was performed to identify GmWRI14 co-expressed genes using the WGCNA R package. The correlation coefficient between any two genes was calculated using WGCNA to analyze the expression patterns of different genes. The square of the threshold of the correlation coefficient was 0.8, and 15,134 potential genes were transferred to the co-expression network to explore the relationship between phenotypes and candidate genes. A gene co-expression correlation matrix was defined by calculating the dissimilarity coefficients of different nodes, and a hierarchical clustering tree was constructed. Abnormal samples were removed based on clustering tree results. Different colors were used to distinguish the 13,211 genes, with gray representing the genes that could not be classified into any module (Fig. 3B). The 13,211 genes were divided into 45 distinct modules (Fig. 3B), and the turquoise, yellow, and light-yellow modules had the highest correlations with LA content in the module–trait correlation analysis (Fig. 3C). RNA-seq showed that DEGs related to linolenic acid metabolism were differentially expressed (more than 2.5 times), namely GmFAD3C and GmFAD3B (Fig. S4). qRT-PCR showed that the expression of GmWRI14 was significantly increased in the seeds of GmWRI14 transgenic soybean lines (Fig. 3D), and the expression of GmbZIP54 in GmWRI14 transgenic soybean seeds on different days was also measured by qRT-PCR (Fig. 3E), with GmbZIP54 expression ranging from 2.45 ± 0.52 to 8.44 ± 0.22. The expression of GmFAD3C and GmFAD3B was significantly decreased in the seeds of GmWRI14 transgenic soybean lines (Table S2), with GmFAD3C expression ranging from 22.75 ± 0.26 to 26.12 ± 0.26 and GmFAD3B expression ranging from 4.12 ± 0.15 to 9.23 ± 0.13 in GmWRI14 transgenic seeds (Figs. 3F and 3G). The expression levels of GmFAD3B and GmFAD3C were 24% and 31% higher, respectively, in control seeds than in the GmWRI14 transgenic soybean lines (Table S2). Notably, GmFAD3B and GmFAD3C expression remained virtually unchanged in the stems and leaves of GmWRI14 transgenic soybeans, with GmFAD3B and GmFAD3C expression in seeds ranging from 11.25 ± 0.41 29.22 ± 0.25. qRT-PCR results suggested that the GmWRI14 gene inhibited the expression of GmFAD3s (GmFAD3C and GmFAD3B) in soybean seeds (Table S2).
Figure 3: KEGG enrichment analysis and weighted gene co-expression network analysis (WGCNA).
(A) The KEGG enrichment analysis shows the differentially-expressed genes are mainly concentrated in fatty acid metabolism and linolenic acid metabolism (red area). (B) The module-sample correlation heatmap. The color scale on the right shows correlations from −1 (red) to 1 (blue). (C) Dendrogram of co-expression modules identified by WGCNA. (D–G) the expression of GmFAD3B, GmFAD3C and GmbZIP54 genes in GmWRI14 transgenic soybean. 1d = 1 day.Disruption of GmWRI14 altered total fatty acid content, and expression of GmFAD3s increased
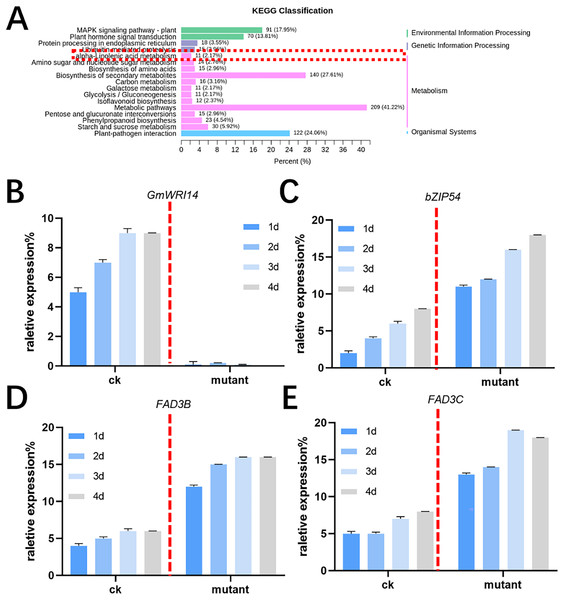
MALDI-TOF MS was used to profile fatty acid species in the mature seeds of gmwri114 mutant and control soybean lines. There was a significant difference between the gmwri114 and wild-type soybeans (P < 0.01). The LA content increased from 11.31% to 20.59% in the gmwri14 mutant, and owing to the reduction in total fatty acids, the size and quality of the gmwri14 mutant seeds were also reduced. The RNA-seq data showed that in gmwri14 mutant soybean seeds, the DEGs related to fatty acid synthesis from the two varieties (CK and GmWRI14 mutant) changed significantly (Fig. 4A), with a threshold of >2.5-fold change indicating significance. In the gmwri14 mutant, qRT-PCR results indicated that the lack of GmWRI14 induced the transcriptional activity of GmFAD3s or GmbZIP54 genes (Figs. 4B–4D). GmWRI14 is a transcriptional inhibitor that negatively regulates the expression of FAD3s. The abundance of FAD3s genes is closely related to linolenic acid content in different plant seeds (Flores et al., 2008; Hu et al., 2006). Therefore, the GmWRI14 gene likely regulates linolenic acid synthesis by regulating FAD3s genes. The qRT-PCR results suggested that disruption of GmWRI14 enhanced the expression of GmFAD3s and GmbZIP54 in soybean seeds.
Figure 4: KEGG classification in gmwri14 mutant soybean plants and qRT-PCR for DEGs.
(A) The functional characterization of GmWRI14 mutant soybean for a KEGG enrichment analysis. (B) The relative expression of GmWRI14 in gmwri14 mutant soybean seeds. (C) The relative expression of GmbZIP54 in GmWRI14 mutant soybean plants. (D) The relative expression of GmFAD3B in gmwri14 mutant soybean plants. (E) The relative expression of GmFAD3C in gmwri14 mutant soybean plants.GmWRI14 can inhibit GmFAD3s in the presence of GmbZIP54
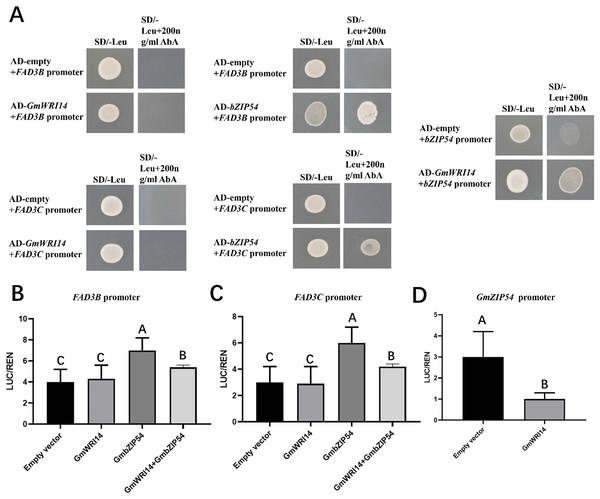
To investigate whether the GmWRI14 gene binds to the promoter regions of GmFAD3B, GmFAD3C, and GmbZIP54 genes in soybean, a yeast one-hybrid assay (Y1H) was performed (Fig. 5A). The results showed that the GmWRI14 gene activated the GmbZIP54 promoter region, but not the promoter regions of the GmFAD3s genes. GmbZIP54 directly bound to the promoter region of GmFAD3s genes (Fig. 5A). When GmFAD3s were co-infiltrated with GmbZIP54, the promoter regions of the GmFAD3s (GmFAD3B and GmFAD3C) were significantly activated (Figs. 5B and 5C), with the promoter regions of the GmbZIP54 were significantly suppressed. The raw data for the dual-luciferase assay are shown in Table S3. Together, these data suggest that GmWRI14 inhibits the expression of GmFAD3s genes by interacting with GmbZIP54, leading to decreased linolenic acid production, and that strong inhibition of FAD3s can potentially be achieved by the combined action of GmbZIP54 and GmWRI14.
Figure 5: GmWRI14 inhibited GmFAD3s (GmFAD3B and GmFAD3C) in the presence of GmbZIP54.
(A) The yeast one-hybrid assay indicates there is no interaction between GmWRI14 and GmFAD3s (GmFAD3B and GmFAD3C) promoter. It also indicates an interaction between GmWRI14 and the GmbZIP54 promoter, and between GmbZIP54 and GmFAD3s (GmFAD3B and GmFAD3C) promoter. (B and C) Transient dual-luciferase detections of GmFAD3B, GmFAD3C, and GmbZIP54 promoters in Nicotiana benthamiana leaves. The relative LUC/REN activity was lower than that of the empty vector, indicating that GmbZIP54 directly binds to the GmFAD3s (GmFAD3B and GmFAD3C) promoter region as a transcription activator of GmbZIP54. (D) The relative LUC/REN activity was lower than that of the empty vector, indicating that GmWRI14 directly binds to the GmbZIP54 promoter region as a transcription inhibitor of GmWRI14. Values are means ± SDs, n = 3. The uppercase letters in (B)–(D) represent significant differences (P < 0.01).Discussion
An estimated 62–77% of the fatty acid content in soybean seeds is polyunsaturated, with linoleic acid comprising 50–58% and linolenic acid comprising 10–12% of the fatty acid content. Linolenic acid (LA, 18:3) is an important component of soybean seeds. It is an essential fatty acid necessary for both oil quality and seed aging (Clemente & Cahoon, 2009). LA also results in a low oil oxidation stability (Tompkins, Maskan & Karataş, 1998). Breeders have screened several genes associated with LA using GWAS (Zeng et al., 2017; Wang et al., 2020; Yu et al., 2019; Zhang et al., 2019). Soybean oil with a low linolenic acid content has great market potential because it is more stable and resistant to high temperatures (Hammond et al., 2018; Perkins, 2000). FAD3 encodes the key enzymes involved in linolenic acid synthesis in different plants, and FAD3 expression levels directly affect linolenic acid and linoleic acid content in seeds (Thapa et al., 2018). During fatty acid metabolism in plants, FAD3 genes are under the control of some TFs such as FUS3 and ABI3, and the cooperation of different types of TFs plays a key role in LA anabolism.
The A9PETALA2 (AP2)-type transcription factor WRINKLED1 (WRI1) was the first positive regulator of fatty acid biosynthesis in plants (Chen et al., 2018; Cerna & Benning, 2004; An & Suh, 2015; Baud et al., 2009). The high sequence similarity observed between the AP2 DNA-binding domains of WRI1, WRI2, WRI3, and WRI4 suggests that these four factors may recognize and bind to similar cis-regulatory elements (To et al., 2012). However, WRI1 is the only member of the WRI clade that triggers high rates of acyl chain production in seeds (To et al., 2012). A previous study found that WRI1 was markedly induced in developing seeds, whereas WRI2, WRI3, and WRI4 mRNAs was barely detectable. Another study showed that WRI1 is the only member of the WRI family that participates in the activation of fatty acid biosynthesis in maturing embryos, providing acyl chains for triacylglycerol (TAG) production (To et al., 2012). WRI1 can also bind to the AW box to activate several target genes involved in fatty acid synthesis at the onset of seed maturation. The WRI1 gene can both directly regulate the synthesis rate of fatty acids and works with a variety of transcription factors to indirectly regulate the synthesis and metabolism of fatty acids (Baud et al., 2009). Many target genes upregulated or downregulated by WRI1 have been discovered by RNA-seq and qRT-PCR, such as pPK, TAL, E1, E2, BS, MCMT, SAD, BLISTER (BLI) and Arabidopsis hemoglobin 1 (AtGLB1) (Ye et al., 2020; To et al., 2012).
Compared with the promotional effect of WRI1, the inhibitory effect of WR1 has not been extensively studied. More attention is usually paid to the impact of WRI on the total oil content of plants, rather than its possible function in regulating the biosynthesis of different fatty acids, or whether the regulation of individual fatty acids explains WRI’s impact of WRI on total oil content. In this study, we used GWAS and RNA-seq to identify potential candidate genes associated with LA content in soybean seeds. We identified a previously unidentified candidate gene, GmWRI14, that is related to LA content. The amino acid sequence encoded by GmWRI14 was compared to AtWRI1 (Gene ID:824599). High sequence similarity was observed between the AP2 DNA-binding domains of AtWRI1 and GmWRI14 (homology of the amino acid sequences between GmWRI14 and AtWRI1 was 62.34%). suggested that GmWRI14 may recognize and bind to a similar cis-regulatory element, leading to a significant increase in seed oil content. In addition, WrI1 seeds have an altered fatty acid composition, with higher levels of linolenic acid (C18:3) and erucic acid (C22:1), but lower levels of oleic acid (C18:1) and linoleic acid (C18:2) (Deng et al., 2019), indicating that the relationship between WRI1 and fatty acid metabolism is complex. Compared to the control group, the seeds of GmWRI14 transgenic soybean lines had a higher total oil content but a 40% lower LA content, resulting in a high oleic acid/linolenic acid ratio. WRI1 has been shown to be necessary for the regulation of carbon partitioning for fatty acid synthesis in plant seeds (Deng et al., 2019; Chen et al., 2018), but WRI1 is unable to directly regulate genes associated with fatty acid modification and triacylglycerol (TAG) assembly in the endoplasmic reticulum (Deng et al., 2019), such as Desaturase2 (FAD2), FAD3, Fatty acid elongase1 (FAE1), and Diacylglycerol acyltransferase1 (Dgat1), because WRI1 specifically recognizes the binding cis element AW-box [CnTnG(n)7CG] (Baud et al., 2009). The RNA-seq and qRT-PCR results in this study demonstrated that GmWRI14 represses FAD3, but the promoter region of FAD3 does not contain an AW-box conserved sequence. Therefore, another transcription factor likely combines with GmWRI14 to convert it into a repressor. GmWRI14 may indirectly mediate the regulation of FAD3s genes. For example, in Arabidopsis seeds, bZIP regulates the omega-3 fatty acid content of seed oil by activating FAD3.
In our study, we found that the interaction of GmbZIP54 with GmWRI14 resulted in a reduction of GmbZIP54 activity. Our study also suggests that WRI1 not only improves oil accumulation but also fine-tunes fatty acid biosynthesis by indirectly inhibiting the expression of FAD3s in soybean seeds. The LA content of GmWRI14 transgenic soybean lines was five-fold lower compared of the control group. The qRT-PCR results showed that the expression of GmFAD3-1B and GmFAD3-1C was significantly reduced in GmWRI14 transgenic soybeans. When GmWRI14 was knocked out, there was a significant increase in LA content, and FAD3 and bZIP54 expression, which are involved in LA metabolism, was affected. We also showed that bZIP54 could bind to the FAD3 promoter region, GmWRI14 could bind to the AW-box in the GmbZIP54 promoter region and limit GmbZIP54 expression, and that GmWRI14 could inhibit FAD3 expression in the presence of bZIP54. Since GmWRI14 could not bind to the FAD3 promoter region because it lacks an AW-box sequence, bZIP54 could bind to the FAD3 promoter region to improve FAD3 expression, and GmWRI14 could bind to the bZIP54 promoter region to inhibit bZIP54 expression, indirectly decreasing the levels of LA production (Fig. 6).
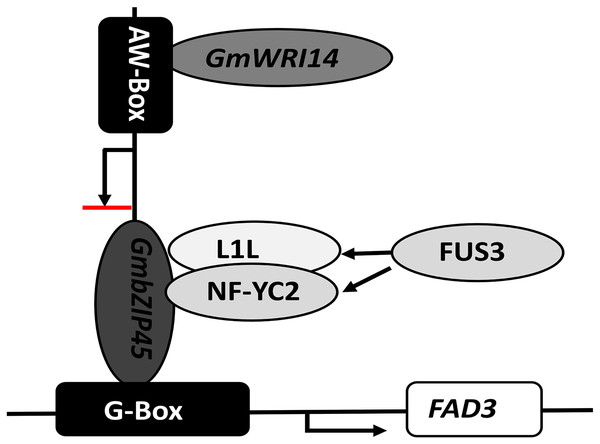
Figure 6: The proposed role of GmWRI14 in the transcriptional regulation of FAD3 in soybean seeds.
GmWRI14 could not bind to the FAD3 promoter region directly, because it lacks an AW-box sequence; bZIP54 could bind to the FAD3 promoter region via G-Box to improve the FAD3 expression, and GmWRI14 could bind to the GmbZIP54 promoter region to inhibit GmbZIP54 expression, indirectly decreasing the levels of LA production. Here, we show that GmWRI14 binds to the GmbZIP54 promoter region, influencing the expression of GmFAD3s.In conclusion, we demonstrated that the WRI1 gene plays an important role in the structure of fatty acids, which helps to maintain the stability of soybean oil. GmWRI13 can directly promote the content of seed oil, but also works with other TFs to indirectly regulate the content of specific fatty acids in soybean oil.
Conclusions
In the present study, a new candidate gene associated with LA content, GmWRI14, was identified in soybean seeds using GWAS and RNA-seq. We generated transgenic soybean plants overexpressing GmWRI14. These GmWRI14 transgenic soybeans had a lower LA content. The RNA-seq results indicated that the transcription factor GmWRI14 inhibited the expression of FAD3s (GmFAD3B and GmFAD3C) genes, and that this inhibition is likely the underlying mechanism of WRI1’s contribution to fatty acid homeostasis in soybean oil.
Supplemental Information
pTF101- GmWRI14-Flag.
The GmWRI14 gene from soybean cultivar 010a (Approval number 2012010) was cloned into the BamHI-SacI site of plasmid pTF101 named pTF101-GmWRI14-Flag, which was induced by the CaMV35S promoter, and the target gene was terminated by the NOS terminator.
The full-length southern blot was used to detect the GmWRI14 expression.
The southern blot was used to detect the GmWRI14 expression. (A) Southern blot analysis of the copy number of the GmWRI14 expression cassette in T0 plants.(B) Southern blot analysis of the copy number of the GmWRI14 expression cassette in T1 plants.
MALDI-TOF IMS was used to analyze LA distribution in the soybean seeds.
The LA content of the GmWRI14 transgenic soybean decreased compared to soybean receptors (CK), the content of linolenic acid in the gmwri14 mutant was significantly increased compared to soybean receptors (CK). (A–C) The LA distribution in different soybean lines, (D–F) Ion peaks of LA in MALDI-TOF-MS of different soybean lines.
RNA-seq showed that DEGs related to linolenic acid metabolism were differentially expressed (more than 2.5 times), namely GmFAD3C and GmFAD3B.
(A) RNA-seq data from transgenic soybean of 8 days. (B) RNA-seq data from transgenic soybean of 10 days. Red represents up-regulated, green represents down-regulated.
Analysis of gene expression in different days.
(A) Analysis of gene expression in different days. (B) Analysis of GmFAD3s and GmbZIP54 expression in different tissues.