spa diversity of methicillin-resistant and -susceptible Staphylococcus aureus in clinical strains from Malaysia: a high prevalence of invasive European spa-type t032
- Published
- Accepted
- Received
- Academic Editor
- Sharif Aly
- Subject Areas
- Microbiology, Molecular Biology, Infectious Diseases
- Keywords
- Methicillin-resistant Staphylococcus aureus, Methicillin-susceptible Staphylococcus aureus, spa typing, t032
- Copyright
- © 2021 Jones et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2021. spa diversity of methicillin-resistant and -susceptible Staphylococcus aureus in clinical strains from Malaysia: a high prevalence of invasive European spa-type t032. PeerJ 9:e11195 https://doi.org/10.7717/peerj.11195
Abstract
Background
Staphylococcus aureus is one of the important pathogens causing nosocomial infection. spa typing allows identification of S. aureus clones in hospital isolates and is useful for epidemiological studies and nosocomial infection control. This study aims to investigate the spa types in Malaysian S. aureus isolates obtained from various clinical specimens.
Method
A total of 89 methicillin-resistant S. aureus (MRSA) [pus (n = 55), blood (n = 27), respiratory (n = 5), eye (n = 2)] isolates and 109 methicillin-susceptible S. aureus (MSSA) [pus (n = 79), blood (n = 24), respiratory (n = 3), eye (n = 2) and urine (n = 1)] isolates were subjected to spa typing with sequences analysed using BioNumerics version 7.
Results
The spa sequence was successfully amplified from 77.8% of the strains (154/198) and 47 known spa types were detected. The distribution of known spa types in MRSA (36.2%, 17/47) was less diverse than in MSSA (70.2%, 33/47). The most predominant spa types were t032 (50%) in MRSA, and t127 (19%) and t091 (16.7%) in MSSA, respectively. spa type t091 in MSSA was significantly associated with skin and soft tissue infections (p = 0.0199).
Conclusion
The previously uncommon spa type t032 was detected in the Malaysian MRSA strains, which also corresponded to the most common spa type in Europe and Australia, and has replaced the dominant spa type t037 which was reported in Malaysia in 2010.
Introduction
Staphylococcus aureus is both a commensal bacterium and human pathogen that has potential to cause a wide variety of infections, ranging from bacteraemia, infective endocarditis, osteoarticular, skin and soft tissue, pleuropulmonary and device-related infections (Jokinen et al., 2018). The first emergence of methicillin-resistant S. aureus (MRSA) was associated with hospital-acquired infections (HA-MRSA) in the early 1960s, but it has spread to the community and was referred to as community-acquired MRSA (CA-MRSA) (Turner et al., 2019). Generally, HA-MRSA and CA-MRSA are differentiated by functional genomic traits (Otto, 2013).
Methicillin-resistant S. aureus has been known to evolve from methicillin-susceptible Staphylococcus aureus (MSSA) due to the acquisition of mecA gene, which encodes the low-affinity penicillin-binding protein 2a (PBP2a) via horizontal transfer located in the mobile genetic element known as staphylococcal cassette chromosome mec (SCCmec). This mechanism allows the bacteria to become resistant to a wide range of β-lactam antibiotics (Kirmusaoglu, 2018; Wu et al., 2019). However, in 2007, a novel gene showed 69% sequence identity to the original mecA gene, which is now termed as mecC. The mecC gene has also shown resistance towards oxacillin and cefoxitin (Lakhundi & Zhang, 2018). Overall, the number of antibiotics that are effective against MRSA is declining as the years passed which inclined to a future where antibiotics are less effective (Mulani et al., 2019). Hence, MRSA is listed as one of the World Health Organization’s high priority pathogens for which new and effective antibiotics are urgently needed.
MRSA infections has caused persistently high mortality across the world. The prevalence rates of MRSA in hospitals in Asian countries such as China, South Korea, Japan and Taiwan ranged from 70 to 80% (Wu et al., 2019; Ghaznavi-Rad et al., 2010). The frequency of MRSA resistance toward the older line of antibiotics such as erythromycin, gentamicin and ciprofloxacin showed a fluctuating trend from 1990 to 2017 in Malaysia but the overall prevalence of antibiotic resistance among MRSA remained above 70%. On the contrary, the prevalence of antibiotic resistance among MSSA strains were much lower below 18% for all seven antibiotics tested—-erythromycin, gentamicin, ciprofloxacin, co-trimoxazole, clindamycin, rifampin and fusidic acid (McNeil et al., 2016; Che Hamzah et al, 2019a).
According to the Malaysian National Antibiotic Resistance Surveillance Report from 2004 to 2017, the prevalence rate of MRSA among S. aureus clinical isolates in the country ranged from 17.2% to 28.1% (Ministry of Health Malaysia, 2017). Most published studies of isolates from Malaysia focused on the spread of MRSA in hospitals and the community rather than MSSA, thus data concerning Malaysian MSSA infections is limited (Che Hamzah et al., 2019a).
Prior to the emergence of MRSA, MSSA was the prominent cause of both serious and uncomplicated S. aureus infections. Some cases describing outbreaks and global spread of MSSA in hospitals affecting neonatal units and hospital staff in UK, USA and Canada were reported, and today this organism still remains as a prime species in hospital infections (DeLeo et al., 2011; Uhlemann et al., 2014; Monaco et al., 2017). Therefore, the understanding of population structure of MRSA and MSSA is important and this can be achieved by identifying the dominant strains circulating in the hospital settings associated with disease outbreaks. The result also helps to monitor bacterial resistance, transmission chain and provide insight into clinical infection controls.
There are various molecular typing tools to characterize the population structure of S. aureus such as pulsed field gel electrophoresis, multilocus sequence typing, SCCmec typing and spa typing (Lakhundi & Zhang, 2018). spa typing is a reliable and discriminative method based on the sequencing analysis of variable number tandem repeats in the highly polymorphic region X of the spa gene encoding for staphylococcal protein A (SpA). The SpA is a surface protein contributing to S. aureus pathogenesis by binding to immunoglobulin which allows the bacteria to be inaccessible to opsonins hence impairing phagocytosis (Asadollahi et al., 2018). The spa region consists of variable number of 21- to 27-bp repeats (Mun & Hwang, 2019). The spa type distribution of S. aureus isolates exhibits different patterns in different geographic location around the world. Constant emergence of new strains that are continuously evolving over time often results in sustained epidemics (Lakhundi & Zhang, 2018). Some studies have suggested that certain S. aureus lineages exhibited a significant trend toward hematogenous complications (Fowler et al., 2017; Rasmussen et al., 2013).
The distribution of spa types in Asia is well studied; however, detailed report on the distribution of spa type of MRSA and MSSA strains from clinical isolates in Malaysia remains scanty. Most studies reported spa type t037 being the predominant type in Malaysia collected from clinical sources, though some studies also reported from the community (Wu et al., 2019; Ghasemzadeh-Moghaddam et al., 2011; Zarizal et al., 2018). The aims of the present study were: (i) to investigate the distribution of spa types in MRSA and MSSA strains collected from clinical sources from a tertiary hospital in the state of Terengganu, Malaysia and (ii) to assess the potential association between the spa type and clinical presentation.
Material and Methods
Bacterial strains
A total of 198 bacterial strains which consisted of 89 clinical MRSA [pus (n = 55), blood (n = 27), respiratory (n = 5), eye (n = 2)] and 109 clinical MSSA [pus (n = 79), blood (n = 24), respiratory (n = 3), eye (n = 2) and urine (n = 1)] isolates that were obtained from Hospital Sultanah Nur Zahirah, Terengganu from July 2016 to June 2017 in a previous study (Che Hamzah et al., 2019b) and that had been cryopreserved in 20% glycerol at −80 °C were used. S. aureus isolates were grown in Luria-Bertani (LB) broth overnight at 37 °C.
Ethics statement
The samples obtained are mainly used for diagnostic laboratory testing in the Hospital Sultanah Nur Zahirah, Terengganu and sample collection were approved by the Malaysia Ministry of Health, Medical Research and Ethics Committee with National Medical Research Registry no: NMRR-15-2369-28130 (IIR).
DNA extraction
Total genomic DNA was extracted from overnight cultures using in-house boiling method (Puah et al., 2018). Briefly, one mL of bacterial culture was harvested by centrifugation at 13,000 rpm for 10 min and was washed twice with distilled water. A total of 100 µL sterile distilled water was added to resuspend the pellet and followed by boiling at 95 °C for 10 min. The cell lysate was incubated on ice for 10 min and pelleted down, followed by transferring the supernatant to a new microcentrifuge tube. The quantity and quality of the total genomic DNA was measured spectrophotometrically. The extracted DNA was stored at −20 °C for further investigation.
spa typing
The spa locus was PCR-amplified using the spa primer pair, spa-1113F (5′-TAAAGACGATCCTTCGGTGAGC-3) and spa-1618R (5′-TTAGCATCTGCATGGTTTGC-3), as described previously (Larsen, Stegger & Sorum, 2008). DNA amplification was carried out in 25 µL PCR mixture consists of 2.5 µL of 10× DreamTaq PCR buffer (20 mM Tris HCl, pH 8.0, 1 mM DTT, 0.1 mM EDTA, 100 mM KCl, 0.5% Nonidet P40, 0.5% Tween 20 and 50% glycerol), 0.4 µM of each 1113F and 1618R primers, 1 U of DreamTaq DNA polymerase (Thermo Fisher, USA), 0.2 mM of each dNTPs and 25 ng of DNA template. The thermal cycling condition was set at initial denaturation at 95 °C for 3 min, followed by 30 cycles of 95 °C for 1 min, 52 °C for 45 s and 72 °C for 45 s, with a final extension at 72 °C for 10 min. The amplified products were visualised on 1% (w/v) agarose gel, followed by purification and subjected to DNA sequencing in a commercial sequencing facility (Apical Laboratories, Malaysia). spa types were subsequently assigned using BioNumerics version 7 (Applied Math, Belgium).
Data analysis
Analysis of spa sequences was performed using the spa typing plug-in tool of the BioNumerics version 7 (Applied Maths, Belgium) via DSI (duplication, substitutions and indels) model for pairwise alignment of repeats. With this plug-in, a similarity matrix was generated based on the DSI model and used to construct a minimum spanning tree (MST). MST was constructed with the node distance between spa types using Dice correction, the node size of spa frequency and a bin unit distance that was set to 1.0. Therefore, the distance between spa types of 99% to 100% similarity was 0 bin distance whereas 98% to 99% similarity was assigned a distance of 1, on MST. In the analysis of association of spa types and clinical characteristics, data was analysed using Fisher’s exact test with a 2 × 2 contingency table and differences at p < 0.05 calculated with two tails was considered statistically significant (McDonald, 2014).
Results
Distribution of spa type
Overall, the spa locus was successfully amplified from 77.8% (154/198) of the strains. Of the forty-four non-typeable strains (i.e., in which no spa amplified product was obtained), 19 were MRSA and 25 were MSSA. The nucleotide sequence analysis of the 154 typeable strains revealed 47 distinct spa types. Distribution of the most common (n ≥ 2) spa types in MRSA and MSSA strains according to clinical source is presented in Table 1. The distribution of spa types in MRSA was less diverse than in MSSA as 36.2% (17/47) were observed among MRSA whereas 70.2% (33/47) were found in MSSA strains. Of the 33 spa types in the MSSA population, 90.9% (30/33) were only detected once in the MSSA population. Three spa types were detected in both MRSA and MSSA, i.e., spa type t127 (3 MRSA, 16 MSSA), spa type t315 (1 MRSA, 5 MSSA) and spa type t548 (2 MRSA, 2 MSSA), while all the other spa types were found only either in MRSA or in MSSA strains. The majority of the MRSA strains belonged to t032 (50%, 35/70), followed by t304 (11.4%, 8/70), and t022 (7.1%, 5/70). The majority of MSSA strains belonged to t127 (19%, 16/84), followed by t091 (16.7%, 14/84), t189 (8.3%, 7/84) and t084 (8.3%, 7/84).
| Spa type | No. of strains (n, %) | ST | ||||||
|---|---|---|---|---|---|---|---|---|
| Blood | Pus | Respiratory | Eye | Urine | Total | |||
| MRSA | t032 | 12 (13.4) | 18 (20.2) | 4 (4.4) | 1 (1.1) | – | 35 (39.3) | ST-22 |
| t304 | 1 (1.1) | 7 (7.8) | – | – | – | 8 (9.0) | ST-6 | |
| t022 | – | 4 (4.4) | 1 (1.1) | – | – | 5 (5.6) | ST-22 | |
| t127 | 1 (1.1) | 2 (2.2) | – | – | – | 3 (3.4) | ST-1 | |
| t458 | 3 (3.3) | – | – | – | – | 3 (3.4) | ST-225 | |
| t037 | 2 (2.2) | 1 (1.1) | – | – | – | 3 (3.4) | ST-239 | |
| t548 | – | 1 (1.1) | – | 1 (1.1) | – | 2 (2.2) | ST-5/97 | |
| t3841 | 1 (1.1) | 1 (1.1) | – | – | – | 2 (2.2) | ST-672 | |
| MSSA | t127 | 4 (3.6) | 11 (10.1) | 1 (0.9) | – | – | 16 (14.7) | ST-1 |
| t091 | 1 (0.9) | 12 (11.0) | – | – | 1 (0.9) | 14 (12.8) | ST-7 | |
| t189 | 1 (0.9) | 6 (5.5) | – | – | – | 7 (6.4) | ST-188 | |
| t084 | 3 (2.7) | 3 (2.7) | – | 1 (0.9) | – | 7 (6.4) | ST-15/18 | |
| t315 | 2 (1.8) | 3 (2.7) | – | – | – | 5 (4.6) | ST-361 | |
| t003 | – | 3 (2.7) | – | – | – | 3 (2.8) | ST-5/225 | |
| t2078 | 1 (0.9) | 2 (1.8) | – | – | – | 3 (2.8) | ST-101 | |
| t548 | 1 (0.9) | 1 (0.9) | – | – | – | 2 (1.8) | ST-5/97 | |
| t2526 | – | 2 (1.8) | – | – | – | 2 (1.8) | ST-88 | |
| t4407 | 1 (0.9) | 1 (0.9) | – | – | – | 2 (1.8) | NA | |
Notes:
- MRSA
-
Methicillin-resistant Staphylococcus aureus
- MSSA
-
Methicillin-sensitive Staphylococcus aureus
- ST
-
Sequence type
- NA
-
Not available
Genetic diversity of spa types
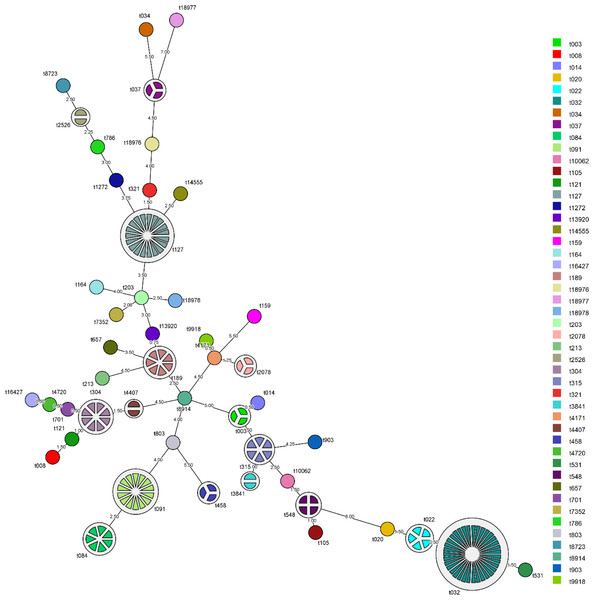
The MST showing the relationship among spa types detected in the Malaysian S. aureus strains was shown in Fig. 1. spa type t022 which accounts for 7.1% (5/70) of MRSA strains was closely related to the predominant spa type t032 in MRSA (50%, 35/70) with 98.5% similarity. Repeats among the known spa types varied from one repeat (t458) to sixteen repeats (t032). The distribution of known spa types with repeat succession above 95% similarity is presented in Table 2. spa type t022 and t032 shared similar sequences (98.5%) except for a single variation due to deletion of one repeat unit in a repeat sequence (t022: 26-23-13-23-31-29-17-31-29-17-25-17-25-16-28 and t032: 26-23-23-13-23-31-29-17-31-29-17-25-17-25-16-28). Another spa type, t084 (07-23-12-34-34-12-12-23-02-12-23) which accounts for 8.3% (7/84) of MSSA strains, was closely related genetically to the most common spa type in MSSA strains, spa type t091 (07-23-21-17-34-12-23-02-12-23) (16.7%, 14/84) with 96.5% similarity.
Figure 1: Minimum spanning tree (MST) of the clinical MRSA and MSSA strains.
MST was performed using the spa typing plug-in tool of the BioNumerics software version 7. The node distance between spa type was computed using Dice correction. Each spa type is depicted by a single node, node size is proportional to spa frequency and line length is proportional to the number of mutational steps between the types. A bin distance of 1.0 was used for the MST parameter.| Spa type | Repeat succession | Strains from clinical source, n | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Pus | Blood | Respiratory | Urine | Eye | Total | |||||||||||||||||
| t022 | 26 | 23 | 13 | 23 | 31 | 29 | 17 | 31 | 29 | 17 | 25 | 17 | 25 | 16 | 28 | 4 | 1 | 5 | ||||
| t032 | 26 | 23 | 23 | 13 | 23 | 31 | 29 | 17 | 31 | 29 | 17 | 25 | 17 | 25 | 16 | 28 | 18 | 12 | 4 | 1 | 35 | |
| t084 | 07 | 23 | 12 | 34 | 34 | 12 | 12 | 23 | 02 | 12 | 23 | 3 | 3 | 1 | 7 | |||||||
| t091 | 07 | 23 | 21 | 17 | 34 | 12 | 23 | 02 | 12 | 23 | 12 | 1 | 1 | 14 | ||||||||
| t127 | 07 | 23 | 21 | 16 | 34 | 33 | 13 | 11 | 4 | 1 | 16 | |||||||||||
| t321 | 07 | 23 | 16 | 34 | 33 | 13 | 1 | 1 | ||||||||||||||
Association of spa types and clinical manifestations caused by MRSA and MSSA
Analysis on the association of spa type and the clinical manifestations caused by MRSA and MSSA strains is shown in Table 2. The predominant spa type t032 and t091 in this study were more commonly associated with skin and soft tissue infections (SSTIs) with 36% (9/25) and 25% (12/48) respectively. However, only spa type t091 from MSSA strains was significantly associated with SSTI (p = 0.0199) (Table 3).
| Spa type | SSTI, n (%) | Non-SSTI, n (%) | P value | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | B | P | PD | OA | IOA | IE | OM | Others | |||||
| MRSA | t032 | 9 (36) | 26 (58) | 5 (11) | 5 (11) | 2 (4) | 1 (2) | 3 (7) | 1 (2) | 9 (20) | 0.1338 | ||
| t304 | 4 (16) | 4 (9) | 1 (2) | 1 (2) | 2 (4) | 0.4435 | |||||||
| t022 | 3 (12) | 2 (4) | 2 (4) | 0.3405 | |||||||||
| t127 | 0 (0) | 3 (7) | 1 (2) | 1 (2) | 1 (2) | 0.5473 | |||||||
| t458 | 1 (4) | 2 (4) | 2 (4) | 1.0000 | |||||||||
| t037 | 1 (4) | 2 (4) | 1 (2) | 1 (2) | 1.0000 | ||||||||
| t548 | 1 (4) | 1 (2) | 1 (2) | 1.0000 | |||||||||
| t3841 | 2 (8) | 0 (0) | 0.1242 | ||||||||||
| Others | 4 (16) | 5 (11) | 0.7119 | ||||||||||
| Total | 25 | 45 | |||||||||||
| MSSA | t127 | 7 (15) | 9 (25) | 2 (6) | 1 (3) | 1 (3) | 1 (3) | 4 (11) | 0.2692 | ||||
| t091 | 12 (25) | 2 (6) | 1 (3) | 1 (3) | 0.0199* | ||||||||
| t189 | 4 (8) | 3 (8) | 1 (3) | 1 (3) | 1 (3) | 1.0000 | |||||||
| t084 | 3 (6) | 4 (11) | 3 (8) | 1 (3) | 0.4551 | ||||||||
| t315 | 2 (4) | 3 (8) | 2 (6) | 1 (3) | 0.6470 | ||||||||
| t003 | 2 (4) | 1 (3) | 1 (3) | 1.0000 | |||||||||
| t2078 | 1 (2) | 2 (6) | 2 (6) | 0.5738 | |||||||||
| t548 | 1 (2) | 1 (3) | 1 (3) | 1.0000 | |||||||||
| t2526 | 2 (4) | 0 (0) | 0.5043 | ||||||||||
| t4407 | 2 (4) | 0 (0) | 0.5043 | ||||||||||
| Others | 12 (25) | 11 (31) | 0.0634 | ||||||||||
| Total | 48 | 36 | |||||||||||
Notes:
Others in MRSA; t315, t203, t657, t020, t8723, t803, t531, t10062, t18978.
Others in MSSA; t008, t786, t213, t903, t014, t164, t8914, t4171, t9918, t321, t701, t1272, t034, t121, t7352, t105, t4720, t159, t18977.
- SSTI
-
skin and soft tissue infection
- B
-
bacteraemia
- P
-
pleuropulmonary
- PD
-
prosthetic device
- OA
-
osteoarticular
- IOA
-
internal organ abscess
- IE
-
infective endocarditis
- OM
-
otitis media
others; epidural abscess, meningitis, toxic shock syndrome, urinary tract infection.
Discussion
Overall, the spa gene was detected in 154 (70 MRSA, 84 MSSA) out of 198 clinical strains, but not in the remaining 44 strains (19 MRSA, 25 MSSA), which were thus referred as non-typeable. This may due to either mutations in the spa gene which prevented the spa primers from annealing and hence, unable to be PCR-amplified, or a true deficiency of spa. Baum et al. (2009) were the first to describe the occurrence of natural protein A mutations in two non-spa typeable clinical S. aureus strains. Both strains were isolated from blood cultures of patients with severe S. aureus infections in Denmark, indicating that the strains were still virulent and invasive in spite of being spa-deficient. On the other hand, Brignoli and co-workers (2019) reported that two MSSA and seven MRSA strains did not express spa in a collection of 133 staphylococcal strains due to the presence of mutations in the spa gene. Another possible reason is due to the weakness of commonly used spa typing primers as rearrangements in the IgG-binding region of the spa gene lead to 1–2% of strains to be designated as “non-typeable” (Votintseva et al., 2014).
The most common spa type, t032, was also the most prevalent spa type causing invasive MRSA infections in Europe. This spa type is reported as one of the spa types associated with the epidemic MRSA known as EMRSA-15 which emerged in the 1980s and spread rapidly in the late 1990s to become the dominant clone of hospital acquired-MRSA strains in the United Kingdom (Grundmann et al., 2010). Interestingly, this clone is reportedly rare outside of European countries. However, it was first detected at two different hospitals in Malaysia, i.e., Hospital Kuala Lumpur and University of Malaya Medical Centre, in 2007 (Wu et al., 2019; Lim et al., 2012). This is the first report which described the dominance of spa type t032 (50%, 35/70) in Malaysia. Our findings showed that spa type t032 colonizes and establishes infection in a wide range of body sites suggesting that it does not possess a tropism for specific sites of infections. Nevertheless, this spa type has caused four deaths isolated from blood culture related to sepsis (2), septic shock (1), and hepatic encephalopathy and coagulopathy (1) as well as three deaths isolated from respiratory specimens from one infant and two adults admitted to the intensive care unit.
A single base pair change within the spa gene can produce different spa type but still highly related (Durand et al., 2018). For example, spa type t022 is a shorter form of spa type t032 with the deletion of one repeat and was detected in this study. spa type t022 is one of the most frequent spa types found in Europe and associated with EMRSA-15, however, it has never been reported in Malaysia (Asadollahi et al., 2018; Grundmann et al., 2010). In this study, the frequency of spa type t022 was low, accounting for 7.1%, (5/70) among MRSA strains but it still caused the death of a 41-year old patient from cellulitis and pneumonia.
In most Asian countries, spa type t037 was the dominant MRSA strain found in clinical settings except for South Korea and Japan (Wu et al., 2019). In Malaysia, spa type t037 was first detected in 2003 (Lim et al., 2012). Over 90% of MRSA strains isolated from 2007 and 2008 belonged to spa type t037 (Lim et al., 2013; Wu et al., 2019). However, the spa type t037 in the present study accounted for only 4.3% (3/70) of all MRSA strains. The spa type t037, also known as the Hungarian/Brazilian clone, was one of the most successful MRSA lineages that is now being replaced by the spa type t032 (Holden et al., 2013; Knight et al., 2012). The second most common spa type in this study, spa type t304, which accounted for 11.4% of our MRSA strains was first detected in 2007 by Lim et al. (2012). spa type t304 was associated with a continuous neonatal ward outbreak in Denmark mainly in 2011 and 2012 (Bartels et al., 2015). Apart from that, it was reported as the dominant strain (39.2%, 31/79) circulating in a tertiary hospital in Oman in 2014 as well as the predominant clone (49.1%, 27/55) in convenience samples collected in Martinique (Uhlemann et al., 2012; Udo et al., 2014). Hence, the rise of spa type t304 clone among MRSA in our study may reflect the on-going expansion of this clone and suggest the need for on-going epidemiological surveillance.
MSSA strains with spa types t127, t091 and t189 were introduced between the year 2009 and 2010 (Wu et al., 2019; Ghasemzadeh-Moghaddam et al., 2011), and they have been reported as the top 20 most frequent spa types among MSSA strains in 25 Europe countries (Grundmann et al., 2014). In this study, spa type t127 was present in both MRSA and MSSA strains, however it was more frequently found in MSSA strains (13.7%, 15/84). Despite the clinical origin, spa type t127 was recently reported in processed food in China and animals, indicating the risk of transfer of food-associated and animal-associated MRSA to humans or vice versa (Franco et al., 2011; Mulani et al., 2019). spa type t091 has been shown to be the major genotype of MSSA with 16.1% (20/124) isolated from bacteraemia patients in China (Chen et al., 2014; He et al., 2013). However, our study reported that only one strain belonged to spa type t091 and this strain was isolated from blood. spa type t189 has been reported as the predominant MSSA strain in China and was mostly isolated from wounds (Chen et al., 2014). This is in accordance to their findings as in this study, spa type t189 (6.4%, 7/84) was mainly isolated from the pus of MSSA patients.
One MSSA strain belonging to spa type t008 was isolated from a 50-year old patient with an abscess. spa type t008 is the predominant MSSA strain in United States, and it was reported to have over 80% similarity to the well-known community-acquired MRSA USA300 (Miko et al., 2013). MRSA USA300 is a pandemic hypervirulent, community-acquired, MRSA clone that spread and caused an outbreak in the United States in the year 2004 and 2008 (Chadwick et al., 2013). Fontanilla et al. (2010) indicated that USA300 strains of CA-MSSA can cause outbreaks comparable to USA300 strains of CA-MRSA. Other studies also reported that CA-MRSA particularly USA300 was the most frequent cause of all infections such as severe skin and soft tissue, necrotizing fasciitis and pneumonia reporting to emergency departments in United States (Moran et al., 2006; Otto, 2009).
Despite spa type t032 being the most prevalent type in this study, there was no association with any specific clinical manifestation. Similar phenomenon had been reported by a previous study whereby molecular genotypes may be independently associated with clinical characteristics of MRSA (Mccarthy et al., 2015). In contrast to MRSA, spa type t091 from MSSA strains were statistically associated with SSTI (p = 0.0199) albeit with a small sample size (n = 25). Therefore, further validation using larger sample cohort collected from different hospitals in Malaysia is required.
Understanding the pathophysiology of S. aureus infections associated with S. aureus clonal lineage is a major challenge in the field of infectious disease. Larger sample size is needed to investigate the association of clinical manifestations with a specific spa type. Furthermore, the observation of untypable strains which might partially due to the limitation of the study where only a single set of primers and an in-house DNA extraction method used. An alternative set of primers and DNA extraction method using lysostaphin should be included to improve the typing sensitivity in future study (Brignoli et al., 2019; Van-tongeren, Degener & Harmsen, 2011). The reason for the serial rises and fall of specific strain types remains poorly understood and the ability to replace a well-established clone is indeed a concern.
Conclusion
In summary, spa type t032 has replaced the previously predominant spa type t037 in 2010 in Malaysia. The results of this study have shown that MRSA and MSSA strains have diverse genetic backgrounds with few dominant clones circulating in the studied hospital.