PeerJ Section
Plant Biology
Welcome to your community’s home at PeerJ. Sections are community led and exemplify a research community’s shared values, norms and interests.
The citation average is 6.5 (view impact metrics).
42,108 Followers
Section Highlights
View all Plant Biology articles 
4 April 2025
Root fragment weight and carbohydrate dynamics of two weedy thistles Cirsium arvense (L.) Scop. and Sonchus arvensis L. during sprouting
"The dynamics of carbohydrates in perennial weed species play a significant role in managing these weeds in various crop fields. I believe the data obtained from this study will greatly aid in controlling these weeds, which cause problems in spring-planted field crops."
 Ahmet Tansel Serim, Handling Editor
Ahmet Tansel Serim, Handling Editor
 Ahmet Tansel Serim, Handling Editor
Ahmet Tansel Serim, Handling Editor
4 April 2025
Advancing medicinal plant agriculture: integrating technology and precision agriculture for sustainability
"An informative , timely, and interesting review on a wide range of technological advances in agriculture. Although the focus is on application to medicinal plants, this review will be of interest to readers in a broader array of crops."
 Julin Maloof, Section Editor
Julin Maloof, Section Editor
 Julin Maloof, Section Editor
Julin Maloof, Section Editor
31 March 2025
Comparative transcriptome analysis identified candidate genes associated with kernel row number in maize
"The authors identified genes potentially involved in increasing corn yield."
 Curtis Daehler, Section Editor
Curtis Daehler, Section Editor
 Curtis Daehler, Section Editor
Curtis Daehler, Section Editor
24 March 2025
Identification and biological characterization of pathogen causing sooty blotch of Ardisia crispa (Thunb.) A.DC.
"This article identifies the causing agents of a disease in Ardisia crispa"
 Fernando Mata, Handling Editor
Fernando Mata, Handling Editor
 Fernando Mata, Handling Editor
Fernando Mata, Handling Editor
21 February 2025
Genetic structure and designing a preliminary core collection of Zizania latifolia in China based on 12 microsatellites markers
"Maintaining and using the biodiversity of wild crop species such as Zizania latifolia can help to mitigate challenges in current agrotechnology and food production."
 Robert Winkler, Section Editor
Robert Winkler, Section Editor
 Robert Winkler, Section Editor
Robert Winkler, Section Editor
16 January 2025
The effectiveness of arbuscular mycorrhizal fungal species (Funneliformis mosseae, Rhizophagus intraradices, and Claroideoglomus etunicatum) in the biocontrol of root and crown rot pathogens, Fusarium solani and Fusarium mixture in pepper
"Arbuscular mycorrhiza have a crucial role in sustainable crop production."
 Nasim Yasin, Handling Editor
Nasim Yasin, Handling Editor
 Nasim Yasin, Handling Editor
Nasim Yasin, Handling Editor
13 January 2025
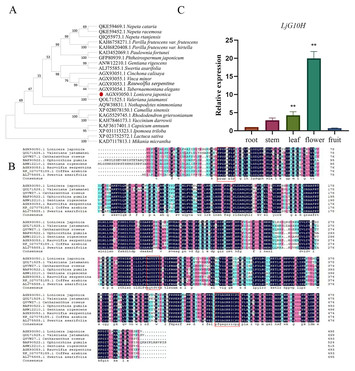
Cloning and functional verification of Geraniol-10-Hydroxylase gene in Lonicera japonica
"An interesting finding has emerged regarding the role of Geraniol 10-hydroxylase in the iridoid biosynthetic pathway."
 Paripok Phitsuwan, Handling Editor
Paripok Phitsuwan, Handling Editor
 Paripok Phitsuwan, Handling Editor
Paripok Phitsuwan, Handling Editor
6 January 2025
Research progress on the impact of climate change on wheat production in China
"An in depth review and analysis of how climate change is affecting wheat production in China and what steps might be taken to mitigate the impacts."
 Julin Maloof, Section Editor
Julin Maloof, Section Editor
 Julin Maloof, Section Editor
Julin Maloof, Section Editor
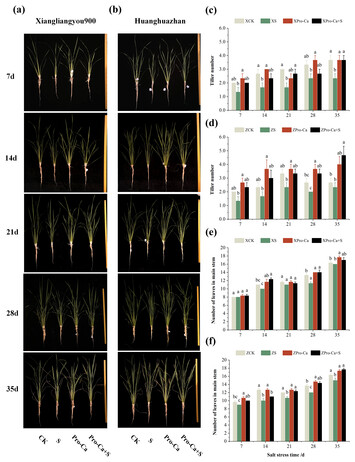
27 December 2024
Effect of salt stress on different tiller positions in rice and the regulatory effect of prohexadione calcium
"The focus on traits in regards to stress conditions is important and this work also keys on a possible treatment which may beneficially influence the crops performance and addresses the possible pathways affected.."
 Gerard Lazo, Section Editor
Gerard Lazo, Section Editor
 Gerard Lazo, Section Editor
Gerard Lazo, Section Editor

10 December 2024
Comparative leaf anatomy of two species of Ipomoea L. (Convolvulaceae): taxonomic importance and adaptations to xeric conditions of the cangas
"The work reports significant results on stomatal anatomy of a specific species for which such information has not been available previously."
 Janendra De Costa, Handling Editor
Janendra De Costa, Handling Editor
 Janendra De Costa, Handling Editor
Janendra De Costa, Handling Editor
42,108 Followers