Detection of a Yersinia pestis gene homologue in rodent samples
- Published
- Accepted
- Received
- Academic Editor
- Marie Lipoldova
- Subject Areas
- Genetics, Microbiology, Molecular Biology
- Keywords
- Yersinia pestis, Rodents, Screening, Microarray
- Copyright
- © 2016 Giles et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2016. Detection of a Yersinia pestis gene homologue in rodent samples. PeerJ 4:e2216 https://doi.org/10.7717/peerj.2216
Abstract
A homologue to a widely used genetic marker, pla, for Yersinia pestis has been identified in tissue samples of two species of rat (Rattus rattus and Rattus norvegicus) and of mice (Mus musculus and Apodemus sylvaticus) using a microarray based platform to screen for zoonotic pathogens of interest. Samples were from urban locations in the UK (Liverpool) and Canada (Vancouver). The results indicate the presence of an unknown bacterium that shares a homologue for the pla gene of Yersinia pestis, so caution should be taken when using this gene as a diagnostic marker.
Introduction
Yersinia pestis is the causative agent of plague in humans and, in the absence of antimicrobial therapy, the mortality rate can approach 100%. In large parts of the world the threat from Y. pestis has declined substantially over time as a result of improvements in living conditions and in public health, including improved rodent control and antibiotics. However, a plague outbreak following the release of a biological weapon is a potential risk. The presence of Y. pestis in small rodent populations in which it is endemic (Ziwa et al., 2013; Eppinger et al., 2009; Biggins & Kosoy, 2001) can cause human fatalities as a result of zoonotic transmission (International Society for Infectious Diseases, 2014).
The black rat (Rattus rattus) has been a major host of Y. pestis for centuries and can be a reservoir for numerous other pathogens. Although most mammalian species can be infected experimentally with Y. pestis, many species fail to develop the high bacteraemia that is necessary to infect the flea vectors. The majority of mammalian species are therefore likely to be dead end hosts (Eisen & Gage, 2009).
Molecular methods, and in particular PCR, have been widely used to identify Y. pestis in tissue samples and the plasminogen activator/coagulase (pla) gene, located on the pPCP1 plasmid has been used as a target in many studies (Loïez et al., 2003; Raoult et al., 2000; Zhang et al., 2013; Ziwa et al., 2013). The pla gene is commonly used because it has a high copy number in Y. pestis (186 per bacterium) and can be detected relatively easily (Parkhill et al., 2001). The PcP plasmid, and the pla gene in particular, is involved in transmission of Y. pestis (Broekhuijsen et al., 2003). The protein encoded by the pla gene induces fibrinolysis and degrades the extracellular matrix and basement membranes, these activities are thought to disrupt the host’s ability to contain the bacteria (Sebbane et al., 2006). The acquisition by Y. pestis of PcP, and of another virulence plasmid, the pMT, is thought to have contributed to the evolutionary transformation of Y. pestis from the mainly gut-associated Yersinia enterocolitica, into a highly host-adapted mammalian blood-borne pathogen (Eppinger et al., 2010).
Primer Name
Primer sequence
Target
Reference
F1
CAGTTCCGTTATCGCCATTGC
caf1
Norkina et al. (1994)
F2
TATTGGTTAGATACGGTTACGGT
Ypfur1
GAAGTGTTGCAAAATCCTGCG
fur
Hinnebusch, Fischer & Schwan (1998)
Ypfur2
AGTGACCGTATAAATACAGGC
YPtoxU
AGGACCTAATATGGAGCATGAC
Ymt
Riehm et al. (2011)
YPtoxUR
CGTGATTACCAGGTGCAACA
Results and Discussion
Probes specific to Y. pestis hybridised with samples from a subset of each of the rodent species tested (12/33 R. rattus, 48/834 R. norvegicus, 3/163 A. sylvaticus, 2/35 M. musculus) giving a total of 65/1065 samples (6.1%) which tested positive on the array. However, none of the generic Yersinia probes hybridised in those samples for which a positive signal was recorded for the Y. pestis specific probes.
Further testing was then carried out at the University of Nottingham, including real-time PCR which targeted another region of the Y. pestis genome, the caf1 gene, for which primers used were identified from the literature (Janse et al., 2010). A subset (23 samples) of the array-positive samples was tested further with primers for pla and caf1. Of these samples, 12 were positive for the pla gene and all were negative for the caf1 gene. A total of 30 array-positive samples were also sent to colleagues in Berlin for further analysis by in solution-based sequence hybridisation, as described previously (Tsangaras et al., 2014). Briefly, a DNA extract from the samples was fragmented and an aliquot was used to produce illumina libraries following a custom protocol (Meyer & Kircher, 2010). PCR amplicons from Y. pestis genes were used to enrich specific target DNA sequences in the rodent samples, the genes and primers used to make the baits are shown in Table 1. The enriched samples were then sequenced using an illumina Miseq. The results indicated that the majority of reads aligned with the rat genome, and only a small number of reads aligned with the pla gene. No other reads mapped to any other gene from Y. pestis.
Although a homologue to the pla gene has previously been reported in bacteria found in R. rattus and R. norvegicus from the Netherlands (Janse, Hamidjaja & Reusken, 2013), this is the first time, to the author’s knowledge, that the bacterial homologue has been reported in M. musculus and A. sylvaticus. The potential discovery of a pla gene homologue in other rodent species, and on another continent than the species and locations in which it has previously been reported, suggests that the homologue could be more widely distributed than previously thought and may cause difficulties in accurate Y. pestis detection. The results found here supports other work which suggests that markers other than the pla gene should be included to help avoid false positive results when screening for Y. pestis, as has been stressed by Janse, Hamidjaja & Reusken (2013). This was recently confirmed by Hänsch et al., as they found evidence that the pla gene is present in some strains of Escherichia coli and Citrobacter koseri (Hänsch et al., 2015). It is not clear why the homologue was present in a larger percentage of R. rattus samples than in the other species tested; perhaps R. rattus carries more E. coli or C. koseri, but this is something that needs to be investigated further.
Materials and Methods
This work was part of a EU project (FP7 WildTech) to develop and use a microarray to detect zoonotic pathogens in rodent tissues. A sequence of the pPCP1 plasmid of the Y. pestis genome (CP000310.1) was obtained from the NCBI database for microarray probe design. Probes were designed using two publicly available software packages: OligoWiz (http://www.cbs.dtu.dk/services/OligoWiz/) and Unique Probe Selector (http://array.iis.sinica.edu.tw/ups/). All probes were checked for suitability using an in silico BLAST analysis. The results of the in silico analysis at the time indicated that the probe sequences were specific to Y. pestis and no cross-hybridisation should occur with eukaryotic or prokaryotic species. Primers were designed using the software Primer3 (http://bioinfo.ut.ee/primer3-0.4.0/). The sequence of each oligonucleotide probe specific to Y. pestis is given in Table 2. During the confirmatory testing, both real-time PCR and end-point PCR were used. The primers used in standard end-point PCR and real-time PCR are shown in Table 3. These probes were evaluated thoroughly for specificity using reference samples of genomic DNA from Y. pestis NCTC5923 Type strain and non-related pathogens before screening took place. The microarray platform used was the ArrayStrip from Alere Technologies GmbH (Jena, Germany).
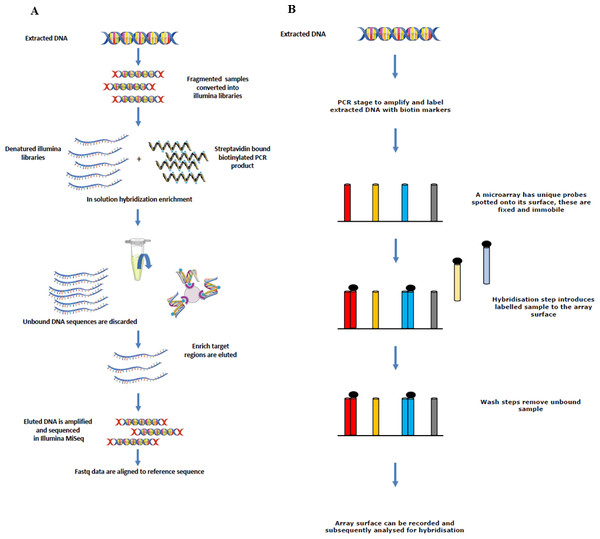
Four different rodent species (R. rattus, R. norvegicus, Mus musculus and Apodemus sylvaticus) were screened for a number of zoonotic pathogens. Tissue samples were obtained from Vancouver (Canada), Liverpool (UK), and Lyon (France) as part of other studies. Automated nucleic acid extraction was performed on the samples using the QIAcube (Qiagen, Hilden, Germany) and the kit (Cador Pathogen Mini Kit; Qiagen, Hilden, Germany). Liver, kidney and lung tissues were available from each rodent sampled from Vancouver and Lyon, and extracted nucleic acid from each tissue was pooled to make a single sample per individual animal which was tested on the array. Only liver and kidney samples were available from the rodents sampled from Liverpool, and again, extracted nucleic acid was pooled to make a single sample. Figure 1 depicts the sequence enrichment and microarray hybridisation used in this study.
| Probe | Sequence (5′–3′) | Pathogen | Gene target | Positiona |
|---|---|---|---|---|
| Y.pestis_Owiz_117 | TACAGATCATATCTCTCTTTTCATCCTCCCCTAGCGGGGAGGATGTCTGTGGAAAGGAGG | Y. pestis | pPCP1 | 8781–8840 |
| Y.pestis_Owiz_120 | TGTTGTCCGCTAGGACGATGCGATTTCGGTTATTATTCAGAATGTCTTCGTTCTCTTTC | Y. pestis | pPCP1 | 6626–6684 |
| Y.pestis_Owiz_121 | TGTCCGGGAGTGCTAATGCAGCATCATCTCAGTTAATACCAAATATATCCCCTGACAGC | Y. pestis | pPCP1 | 7878–7936 |
| Y.pestis_Owiz_127 | GTGGAGATTCTGTCTCTATTGGCGGAGATGCTGCCGGTATTTCCAATAAAAATTATACTG | Y. pestis | pPCP1 | 8688–8747 |
| Y.pestis_Owiz_129 | GAATCGCGCCCGGATATGTTTTAACGCGATTTTCAGACTCAGACAAATTCAGCAGAAT | Y. pestis | pPCP1 | 9990–10047 |
| Y.pestis_Owiz_147 | TCGCTGGCTAAAAAGTACCATCCACATGCTCAACCCTATAACCTGTAGCTTACCCCAC | Y. pestis | pPCP1 | 9583–9640 |
| YpestisUPS_785 | AATAGGTTATAACCAGCGCTTTTCTATGCCATATATTGGACTTGCAGGCCAGTATCGCAT | Y. pestis | pPCP1 | 8392–8451 |
| YpestisUPS_786 | AATGATGAGCACTATATGAGAGATCTTACTTTCCGTGAGAAGACATCCGGCTCACGTTAT | Y. pestis | pPCP1 | 8510–8569 |
| YpestisUPS_787 | TAAATTCAGCGACTGGGTTCGGGCACATGATAATGATGAGCACTATATGAGAGATCTTAC | Y. pestis | pPCP1 | 8479–8538 |
| Y.pestisUPS_788 | AGCCCGACCACTGCGCCTTATCCGGTAACTATCGTCTTGAGTCCAACCCGGTAAGACACG | Y. pestis | pPCP1 | 4977–5036 |
| YpestisUPS_789 | TCATCCTCCCCTAGCGGGGAGGATGTCTGTGGAAAGGAGGTTGGTGTTTGACCAACCTTC | Y. pestis | pPCP | 8801–8860 |
| YpestisUPS_790 | AAAGGACAGCATTTGGTATCTGTGCTCCACTTAAGCCAGCTACCACAGGTTAGAAAGCCT | Y. pestis | pPCP | 5129–5188 |
| YpestisUPS_791 | AAGGAGTGCGGGTAATAGGTTATAACCAGCGCTTTTCTATGCCATATATTGGACTTGCAG | Y. pestis | pPCP | 8379–8438 |
| YpestisUPS_792 | TTTGTACCGAGAACCTTTCACGGTATCGGCATATGGCCTGGGTAACTCAGGTCCGTAAAC | Y. pestis | pPCP | 9451–9510 |
Notes:
| Forward Primer | Sequence (5′–3′) | Reverse primer | Sequence (5′–3′) | Probe | Sequence (5′–3′) | Gene | Position |
|---|---|---|---|---|---|---|---|
| Y.pes/pPCP/8374/F | CCCGAAAGGAG TGCGGGTAA | Y.pes/pPCP/8902/R | CGCCCCGTCATT ATGGTGAA | N/A | N/A | pla | 8374–8902a |
| cafpri_f | CCAGCCCGCAT CACT | cafpri_r | ATCTGTAAAGTTAA CAGATGTGCTAGT | Tqpro_caf | JOE-AGCGTACCAA CAAGTAATTCTGTA TCGATG-BHQ1 | caf1 | 109–255b |
| Y.pes_pPCP_F | AGACATCCGG CTCACGTTAT | Y.pes_pPCP_R | GAGTACCTCCT TTGCCCTCA | Y.pes_pPCP_Pr | FAM-CACCTAA TGCCAAAGTCTTT GCGGA-TAMRA | pla | 8550–8669a |
Figure 1: Sequence showing how extracted DNA is used for sequence enrichment capture (A) and microarray hybridisation (B).
Key findings
-
1.
Sequences homologous to the pla gene, which is present in Y. pestis, have been found in samples from several rodent species, in the absence of Y. pestis.
-
2.
PCR, microarray and sequencing data suggest that these sequences may be present in environmental bacteria.
-
3.
Caution is warranted in interpreting screening results for detection of Y. pestis when the pla gene is used as the sole marker for the presence of the pathogen.