Characterization of endoplasmic reticulum-associated degradation in the human fungal pathogen Candida albicans
- Published
- Accepted
- Received
- Academic Editor
- Howard Young
- Subject Areas
- Biochemistry, Cell Biology, Genetics, Molecular Biology, Mycology
- Keywords
- Candida albicans, Endoplasmic reticulum-associated degradation, Ubiquitin ligase, Ubiquitin-conjugating enzyme, Mass spectrometry, Hrd1, Doa10, Ubc7, Protein quality control, Pathogenic fungi
- Copyright
- © 2023 Doss et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2023. Characterization of endoplasmic reticulum-associated degradation in the human fungal pathogen Candida albicans. PeerJ 11:e15897 https://doi.org/10.7717/peerj.15897
Abstract
Background
Candida albicans is the most prevalent human fungal pathogen. In immunocompromised individuals, C. albicans can cause serious systemic disease, and patients infected with drug-resistant isolates have few treatment options. The ubiquitin-proteasome system has not been thoroughly characterized in C. albicans. Research from other organisms has shown ubiquitination is important for protein quality control and regulated protein degradation at the endoplasmic reticulum (ER) via ER-associated protein degradation (ERAD).
Methods
Here we perform the first characterization, to our knowledge, of ERAD in a human fungal pathogen. We generated functional knockouts of C. albicans genes encoding three proteins predicted to play roles in ERAD, the ubiquitin ligases Hrd1 and Doa10 and the ubiquitin-conjugating enzyme Ubc7. We assessed the fitness of each mutant in the presence of proteotoxic stress, and we used quantitative tandem mass tag mass spectrometry to characterize proteomic alterations in yeast lacking each gene.
Results
Consistent with a role in protein quality control, yeast lacking proteins thought to contribute to ERAD displayed hypersensitivity to proteotoxic stress. Furthermore, each mutant displayed distinct proteomic profiles, revealing potential physiological ERAD substrates, co-factors, and compensatory stress response factors. Among candidate ERAD substrates are enzymes contributing to ergosterol synthesis, a known therapeutic vulnerability of C. albicans. Together, our results provide the first description of ERAD function in C. albicans, and, to our knowledge, any pathogenic fungus.
Introduction
Candida albicans is a commensal fungus residing in the gastrointestinal tract of humans (Kumamoto, Gresnigt & Hube, 2020). C. albicans is found in or on most humans and can cause superficial infections, such as oral thrush or vaginosis (Goncalves et al., 2016; Millsop & Fazel, 2016). However, in immunocompromised individuals, C. albicans can cause more serious infections such as candidemia and candidiasis, infections of blood or hard organs, respectively (Pappas et al., 2018). These infections can be life-threatening when left untreated. There are a number of antifungal drugs that can be used to treat C. albicans infection; however, drug-resistant isolates have been and continue to be isolated, limiting treatment options (Lee et al., 2021). As such, it is critical that we develop novel antifungal therapeutics to treat these resistant infections. To do this, we must better understand C. albicans function at the molecular level.
One area of C. albicans biology that has yet to be thoroughly characterized is the ubiquitin-proteasome system (UPS). The UPS plays numerous important roles in eukaryotic cells, degrading proteins that are either no longer needed or have been compromised (e.g., by misfolding) (Berner, Reutter & Wolf, 2018). In C. albicans, ubiquitin is important for stress adaptation, and deletion of the gene that encodes ubiquitin, Ca_UBI4, attenuates virulence in a mouse model (Leach et al., 2011). Furthermore, mutation of Ca_UBI4 leads to hypersensitivity to oxidative stress, the antifungal drug caspofungin, and the ER stress inducer tunicamycin. Ubiquitylation plays important roles in protein quality control and regulated protein degradation at the endoplasmic reticulum (ER) via ER-associated protein degradation (ERAD) (Berner, Reutter & Wolf, 2018; Mehrtash & Hochstrasser, 2019); as such, ERAD may be a vulnerability that can be leveraged during antifungal therapeutic discovery.
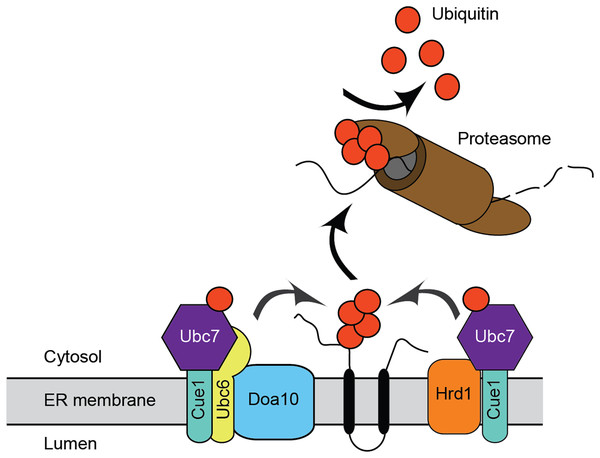
ERAD has been extensively characterized in the model unicellular fungal eukaryote, Saccharomyces cerevisiae, and in mammals. The major ERAD ubiquitin ligases in S. cerevisiae are the multipass transmembrane proteins ScHrd1 and ScDoa10 (Fig. 1). ScHrd1 primarily functions with the ubiquitin-conjugating enzyme ScUbc7 (which is anchored at the ER membrane by the transmembrane protein ScCue1), while ScDoa10 works with two ubiquitin-conjugating enzymes, ScUbc7 and the transmembrane enzyme ScUbc6 (Bays et al., 2001; Lips et al., 2020; Plemper et al., 1999; Swanson, Locher & Hochstrasser, 2001). ScHrd1 and ScDoa10 target distinct, but partially overlapping, subsets of protein quality control substrates for degradation. ScHrd1 mediates turnover of ER soluble luminal proteins, transmembrane proteins, and proteins that persistently engage (i.e., clog) the ER translocon, while ScDoa10 promotes destruction of soluble cytosolic and transmembrane proteins (Carvalho, Goder & Rapoport, 2006; Gauss, Sommer & Jarosch, 2006; Habeck et al., 2015; Huyer et al., 2004; Metzger et al., 2008; Ravid, Kreft & Hochstrasser, 2006; Rubenstein et al., 2012; Runnebohm et al., 2020b; Sato et al., 2009). In mammals, ERAD is mediated by an expanded cadre of ubiquitin ligases and conjugating enzymes, including homologs of ScHrd1 (HRD1/SYVN1 and gp78) and ScDoa10 (MARCHF6/TEB4) (Hassink et al., 2005; Kikkert et al., 2004; Liang et al., 2003). In addition to their well-characterized roles in protein quality control, ScHrd1 and ScDoa10 also contribute to regulated turnover of otherwise normal proteins. For example, S. cerevisiae and mammalian homologs of Hrd1 and Doa10 promote regulated turnover of sterol biosynthetic enzymes (Foresti et al., 2013; Garza, Tran & Hampton, 2009; Hampton, Gardner & Rine, 1996; Huang & Chen, 2023; Jo et al., 2011; Stevenson, Huang & Olzmann, 2016).
Figure 1: Endoplasmic reticulum-associated degradation.
Mechanism for ER protein degradation in S. cerevisiae. See text for details.Virtually nothing is known about ERAD mechanisms in pathogenic fungi, including C. albicans. Candida glabrata UBC7 mRNA abundance exhibits a ~3-fold increase following azole treatment, suggesting a role for ERAD in pathogenic fungal stress response (Li, Skinner & Bennett, 2012). In this study, we performed initial characterization of Ca_HRD1, Ca_DOA10, and Ca_UBC7, genes predicted to encode enzymes that function in ERAD. We generated homozygous C. albicans mutants lacking Ca_HRD1, Ca_DOA10, and Ca_UBC7. Consistent with a role in protein quality control, yeast lacking putative ERAD components exhibited hypersensitivity to proteotoxic stress. Further, mass spectrometric analyses of wild type and mutant yeast revealed distinct proteomic profiles for each mutant, illuminating candidate physiological ERAD substrates, co-factors, and compensatory stress response factors. Together, these results provide the first description of ERAD function in C. albicans, and, to our knowledge, any pathogenic fungus.
Materials and Methods
Generation of mutants
Mutations to Ca_HRD1, Ca_DOA10, and Ca_UBC7 were made using CRISPR-mediated genome editing. Guide RNAs to the 5′ region of Ca_HRD1, Ca_DOA10, and Ca_UBC7 were cloned into plasmid pV1093 (Vyas, Barrasa & Fink, 2015), which also contains Cas9 as well as nourseothricin (Nat) resistance marker (NatR). Repair templates that introduce a stop codon and restriction site upon homologous recombination with the cut chromosome were generated. C. albicans wild type strain SC5314 was transformed with Cas9 guide constructs and repair templates using lithium acetate. Transformants were selected on 200 µg/ml Nat. Correct homozygous transformants were identified by colony PCR followed by restriction digestion. Sanger sequencing was used to confirm the mutants. Homozygous mutants will be henceforth referred to as Ca_hrd1, Ca_doa10, and Ca_ubc7.
Growth analyses
Spot assays: Yeast growth assays were performed using a modified version of a previously published protocol (Watts et al., 2015). Yeast were grown to saturation overnight at 30 °C in yeast extract-peptone-dextrose media (YPD). Cells were diluted to an OD600 of 1.0. Fivefold dilutions were prepared in a 96-well plate in YPD. Cells were plated using a pin replicator onto YPD agar lacking or containing increasing concentrations of hygromycin B (Brodersen et al., 2000; Ganoza & Kiel, 2001). Cells were grown at 30 and 37 °C for 5 days and at 25 and 40 °C for 10 days.
Inhibition assays: A modified drug susceptibility assay (Bauer et al., 1966) was performed. Yeast were grown to saturation overnight at 30 °C in YPD. 1 × 107 cells were spread on 20 ml of synthetic complete media in 150 mM plates and were allowed to dry. A sterile filter paper disc was placed on the cells, and 15 µl of 1 M dithiothreitol (DTT) suspended in water, 33% (v/v) β-mercaptoethanol (βME) in water (Jia et al., 2019), or sterile water were directly pipetted onto the paper disc. Plates were incubated for 3 days at 37 °C before visualization.
Mass spectrometry and data analysis
Sample preparation, mass spectrometry analysis, bioinformatics, and data evaluation for quantitative proteomics experiments were performed in collaboration with the Indiana University Proteomics Center for Proteome Analysis at the Indiana University School of Medicine (IUSM) similarly to previously published protocols (Kumar et al., 2022; Morris et al., 2022; Soundararajan et al., 2022; Stanhope et al., 2023).
Sample preparation: 12 samples (n = 3 of wild type, Ca_hrd1/Ca_hrd1, Ca_doa10/Ca_doa10, and Ca_ubc7/Ca_ubc7 yeast) were submitted to the IUSM Center for proteome analysis, where proteins were denatured in 8 M urea, 100 mM Tris-HCl, pH 8.5 with sonication using a Bioruptor® sonication system (Diagenode Inc, Denville, NJ, USA) with 30 s/30 s on/off cycles for 15 min in a water bath at 4 °C. After subsequent centrifugation at 14,000 rcf for 20 min, protein concentrations were determined by Bradford protein assay (BioRad Cat No: 5000006). A total of 100 µg equivalent of protein from each sample were reduced with 5 mM tris (2-carboxyethyl) phosphine hydrochloride (TCEP, Sigma-Aldrich Cat No: C4706) for 30 min at room temperature and alkylated with 10 mM chloroacetamide (CAA, Sigma Aldrich Cat No: C0267) for 30 min at room temperature in the dark. Samples were diluted with 50 mM Tris.HCl, pH 8.5 to a final urea concentration of 2 M for Trypsin/Lys-C based overnight protein digestion at 37 °C (1:70 protease:substrate ratio, Mass Spectrometry grade, Promega Corporation, Cat No: V5072.).
Peptide purification and labeling: Digestions were acidified with trifluoroacetic acid (TFA, 0.5% v/v) and desalted on Sep-Pak® Vac cartridges (WatersTM Cat No: WAT054955) with a wash of 1 ml 0.1% TFA followed by elution in 70% acetonitrile 0.1% formic acid (FA). Peptide concentrations were checked by Pierce Quantitative colorimetric assay (Cat No: 23275) and confirmed to be consistent across all samples. 50 µg peptides were then labeled with 0.25 mg Tandem Mass Tag pro (TMTpro) reagent (Thermo Fisher Scientific, TMTpro™ Isobaric Label Reagent Set; Cat No: A44520, Lot VL313890; see Table S1) for 2 h at room temperature, quenched with a final concentration v/v of 0.3% hydroxylamine at room temperature for 15 min. Labeled peptides were mixed and dried by speed vacuum.
High pH basic fractionation: For high pH basic fractionation, peptides were reconstituted in 0.1% trifluoroacetic acid and half of the mixture was fractionated on a 50 mg Sep-Pak® Vac cartridge using methodology and reagents from Pierce™ High pH reversed-phase peptide fractionation kit (Thermo Fisher Cat No: 84868).
Nano-LC-MS/MS: Global proteomics were performed on an EASY-nLC 1200 HPLC system (SCR: 014993; Thermo Fisher Scientific, Waltham, MA, USA) coupled to Lumos Orbitrap™ mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA). A total of 1/5 of each fraction was loaded onto a 25-cm EasySpray column (Thermo Fisher Scientific ES902A, Waltham, MA, USA) at 400 nl/min. Peptides were eluted from 4–30% with mobile phase B (Mobile phases A: 0.1% FA, water; B: 0.1% FA, 80% Acetonitrile (Thermo Fisher Scientific Cat No: LS122500, Waltham, MA, USA)) over 160 min, 30–80% B over 10 min, and dropping from 80–10% B over the final 10 min. The mass spectrometer was operated in positive ion mode with a 4-s cycle time data-dependent acquisition method with advanced peak determination and Easy-IC (internal calibrant) on. Precursor scans (m/z 375–1,600) were done with an orbitrap resolution of 120,000, RF lens% 30, maximum inject time 50 ms, AGC target of 100% (4e5), MS2 intensity threshold of 2.5e4, MIPS mode to peptide, including charges of 2 to 7 for fragmentation with 30 s dynamic exclusion. MS2 scans were performed with a quadrupole isolation window of 0.7 m/z, 37% HCD CE, 50,000 resolution, 200% normalized AGC target, dynamic maximum IT, fixed first mass of 100 m/z.
Mass spectrometry data analysis: Resulting RAW files were analyzed in Proteome Discover™ 2.5 (Thermo Fisher Scientific, Waltham, MA, USA) with a C. albicans UniProt FASTA (downloaded 03/15/2022, 6,030 entries) plus common contaminants. SEQUEST HT searches were conducted with a maximum number of three missed cleavages, precursor mass tolerance of 10 ppm, and a fragment mass tolerance of 0.02 Da. Static modifications used for the search were carbamidomethylation on cysteine residues and TMTpro label on lysine residues. Dynamic modifications included oxidation of methionine, TMTpro label on peptide N terminus, and acetylation, methionine-loss, or methionine-loss plus acetylation on protein N terminus. Percolator False Discovery Rate (FDR) was set to a strict peptide spectral match FDR setting of 0.01 and a relaxed setting of 0.05. In the consensus workflow, peptides were normalized by total peptide amount with no scaling. Co-isolation thresholds of 50% and average reporter ion S/N cutoffs of five were used for quantification. All peptides were used for normalization and protein roll-up and modified peptides were excluded in the pairwise ratio calculation. Protein abundance-based ratio calculations were done with no imputation. Protein FDR validator node was set to a strict target FDR of 0.01 and relaxed of 0.05. Resulting normalized abundance values for each sample type, abundance ratio and log2 (abundance ratio) values, and respective p-values (protein abundance-based ratio calculation and individual protein ANOVA) from Proteome Discover™ were exported to Microsoft Excel.
Results
Structural analyses of ERAD enzymes
We identified putative C. albicans genes encoding ERAD enzymes CaHrd1, CaDoa10, and CaUbc7. A summary of amino acid sequence identity and similarity between homologous proteins in C. albicans, S. cerevisiae, and H. sapiens is presented in Table S2. Domain organization and AlphaFold-predicted (Jumper et al., 2021) or experimentally determined (Arai et al., 2006; Cook et al., 1997; Wu et al., 2020) structures of these enzymes are presented in Fig. 2.
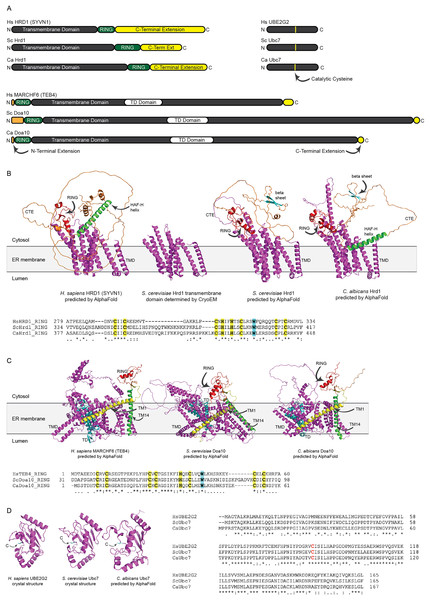
Figure 2: Structural analysis of H. sapiens, S. cerevisiae, and C. albicans Hrd1, Doa10, and Ubc7 homologs.
(A) To-scale domain architecture of human and fungal homologs of Hrd1, Doa10, and Ubc7. See text for details. RING, Really Interesting New Gene domain. TD, TEB4-Doa10 domain. (B) Top, AlphaFold-predicted structures of H. sapiens HRD1, S. cerevisiae Hrd1, and C. albicans Hrd1 and cryo-electron microscopy structure of transmembrane portion of S. cerevisiae Hrd1 (PDB 6VJZ) (Wu et al., 2020). Magenta, transmembrane domain (TMD). Red, RING domain. Orange, C-terminal extension (CTE). Green, predicted HAF-H helix shared by HsHRD1 and CaHrd1. Cyan, predicted beta sheet shared by ScHrd1 and CaHrd1. Bottom, Amino acid alignment of catalytic RING domains of Hrd1 homologs. Bold yellow, Zinc-coordinating Cys and His residues. Bold cyan, Trp residue commonly found in RING domains. (C) Top, AlphaFold-predicted structures of H. sapiens MARCHF6, S. cerevisiae Doa10, and C. albicans Doa10. Magenta, transmembrane domain (TMD). Cyan, TD domain. Red, RING domain. Orange, C-terminal extension. Yellow and Green, first and fourteenth transmembrane segments, respectively (TM1 and TM14). Bottom, Amino acid alignment of catalytic RING domains of Doa10 homologs. Bold yellow, Zinc-coordinating Cys and His residues. Bold cyan, Trp residue commonly found in RING domains. (D) Left, Crystal structures of H. sapiens UBE2G2 (Arai et al., 2006) and S. cerevisiae Ubc7 (PDB 2UCZ) (Cook et al., 1997) and AlphaFold-predicted structure of C. albicans Ubc7. Cyan, catalytic cysteine (C). Right, Amino acid alignment of Ubc7 homologs. Bold red, catalytic cysteine.All three (H. sapiens, S. cerevisiae, and C. albicans) Hrd1 homologs possess an N-terminal transmembrane domain with eight predicted membrane-spanning segments, a catalytic Really Interesting New Gene (RING) domain, and a largely unstructured C-terminal extension (CTE), which likely contributes to substrate and cofactor interactions (Figs. 2A and 2B) (Omura et al., 2006; Schulz et al., 2017). The predicted CaHrd1 CTE possesses H. sapiens-like and S. cerevisiae-like features. The CaHrd1 CTE includes an alpha helix with amphipathic character (green helix in Fig. 2B) resembling the human C-terminal HAF-H domain implicated in HsHRD1 complex formation (Schulz et al., 2017). Like the S. cerevisiae enzyme, the CaHrd1 CTE includes a predicted two-stranded beta-sheet (cyan strands in Fig. 2B) with uncharacterized function.
AlphaFold-predicted structures of H. sapiens, S. cerevisiae, and C. albicans Doa10/MARCHF6 proteins include N-terminal RING domains, large C-terminal transmembrane domains, and short N- and C-terminal extensions (Figs. 2A and 2C). The transmembrane portions of these proteins include a conserved three-transmembrane segment TEB4-Doa10 (TD) domain (cyan in Fig. 2C) (Kreft & Hochstrasser, 2011). The large transmembrane portion of Doa10 homologs has been proposed to function as a retrotranslocation channel for ER export of transmembrane substrates, with the first (yellow) and fourteenth (green) transmembrane segments forming a lateral gate for substrate entry (Mehrtash & Hochstrasser, 2022; Schmidt, Vasic & Stein, 2020). The ScDoa10 and HsMARCHF6 CTEs promote substrate ubiquitylation (Mehrtash & Hochstrasser, 2022; Zattas et al., 2016); AlphaFold-guided mutational analyses indicate interactions between the CTE and N-terminal RING domain are essential for optimal ScDoa10 function (Mehrtash & Hochstrasser, 2022). AlphaFold structural predictions indicate this interaction is likely also present in CaDoa10 and HsMARCHF6. Alignments of Hrd1 and Doa10 homolog RING domains demonstrate conservation of zinc-coordinating cysteine and histidine residues as well as a tryptophan residue commonly found in ubiquitin ligase catalytic domains (Figs. 2B and 2C). Complete alignments of Hrd1 and Doa10 homologs are presented in Supplemental Files 1 and 2.
H. sapiens, S. cerevisiae, and C. albicans Ubc7 homologs exhibit strong conservation of primary and predicted tertiary structure (Figs. 2A and 2D). Comparison of crystal structures of ScUbc7 and HsUBE2G2 with AlphaFold-predicted CaUbc7 reveals all three adopt a characteristic ubiquitin-conjugating enzyme fold; the position of the invariant catalytic cysteine is indicated.
Generation of mutants
We generated functional deletions of Ca_HRD1, Ca_DOA10, or Ca_UBC7 using CRISPR-mediated genome editing. Stop codons were inserted at the 5′ ends of each of these genes, effectively creating knockouts of Ca_HRD1, Ca_DOA10, and Ca_UBC7. PCR and Sanger sequencing were used to confirm the correct mutations were made and that mutations were homozygous, preventing translation of both alleles’ transcripts. Heterozygous Ca_HRD1/Ca_hrd1 mutants were also isolated. Introduction of these mutations did not impact growth on YPD (Fig. 3).
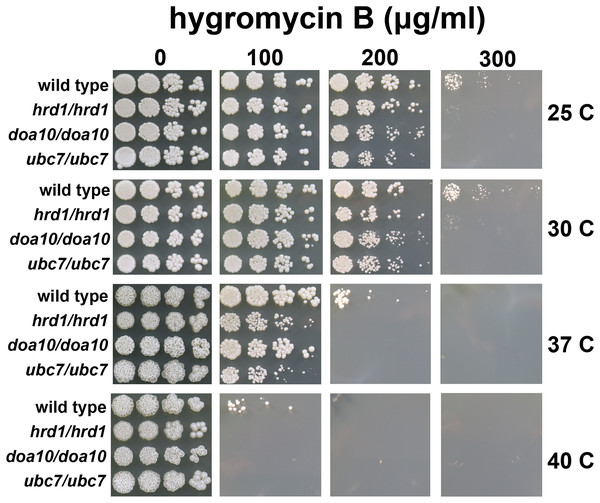
Figure 3: C. albicans ERAD enzymes confer resistance to hygromycin B.
Wild type or indicated homozygous mutant C. albicans were serially diluted and spotted onto YPD growth medium in the presence of increasing concentrations of hygromycin B and incubated at the indicated temperatures.C. albicans lacking HRD1, DOA10, or UBC7 exhibit sensitivity to proteotoxic stress
A quality control function for C. albicans genes encoding ERAD factors has not previously been reported. S. cerevisiae with mutations in protein quality control genes (including those encoding ERAD enzymes) exhibit elevated sensitivity to hygromycin B (Crowder et al., 2015; Runnebohm et al., 2020a; Verma et al., 2013; Woodruff et al., 2021). Hygromycin B distorts the ribosomal A site, thereby reducing translational fidelity, which is predicted to increase the abundance of aberrant proteins (Brodersen et al., 2000; Ganoza & Kiel, 2001). We compared the growth of wild type yeast and yeast with homozygous disruptions of Ca_HRD1, Ca_DOA10, or Ca_UBC7 in the presence of increasing concentrations of hygromycin B across a range of temperatures (Fig. 3). Homozygous deletion of Ca_HRD1, Ca_DOA10, or Ca_UBC7 caused a pronounced growth defect in the presence of hygromycin B. Sensitivity to hygromycin B was markedly enhanced at elevated temperatures.
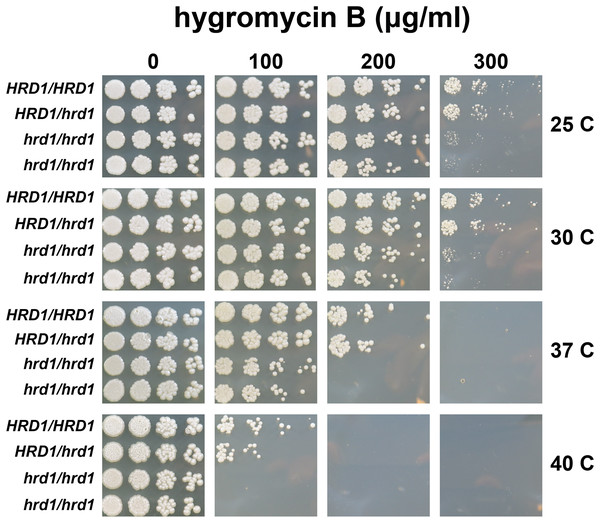
We also compared hygromycin B sensitivity of yeast possessing homozygous and heterozygous mutations in Ca_HRD1 (Fig. 4). Heterozygous Ca_HRD1/Ca_hrd1 yeast exhibited wild type resistance to hygromycin B at 25, 30, and 37 °C, indicating a single copy of Ca_HRD1 is sufficient to confer protection from hygromycin B at these temperatures. We observed a subtle growth defect of Ca_HRD1/Ca_hrd1 yeast in the presence of hygromycin B at 40 °C.
Figure 4: A single copy of C. albicans HRD1 is sufficient to confer resistance to hygromycin B.
Yeast of the indicated genotypes were serially diluted and spotted onto YPD growth medium in the presence of increasing concentrations of hygromycin B and incubated at the indicated temperatures.We analyzed susceptibility of wild type yeast and ERAD mutants to drugs that cause ER stress (disulfide-reducing agents beta-mercaptoethanol (βME) and dithiothreitol (DTT)) (Jia et al., 2019; Wu, Ng & Thibault, 2014). Homozygous mutation of Ca_HRD1, Ca_DOA10, or Ca_UBC7 enhanced sensitivity to βME (Fig. 5). By contrast, wild type and ERAD mutant strains exhibited similar susceptibility to DTT. Together, our results suggest Ca_HRD1, Ca_DOA10, or Ca_UBC7 are important for C. albicans growth under proteotoxic stress.
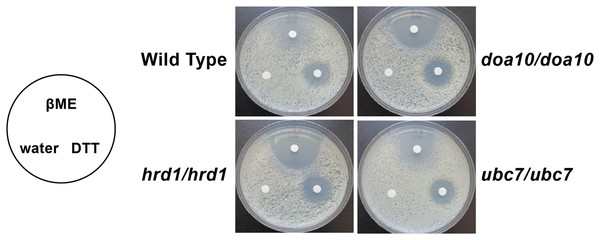
Figure 5: C. albicans ERAD enzymes confer resistance to β-mercaptoethanol.
Lawns of yeast of the indicated genotypes were seeded onto YPD media. Prior to incubation at the indicated temperatures, sterile filter papers were impregnated with 15 µl sterile water, 33% β-mercaptoethanol (βME), or 1 M dithiothreitol (DTT).Proteomic analysis of ERAD mutant yeast strains
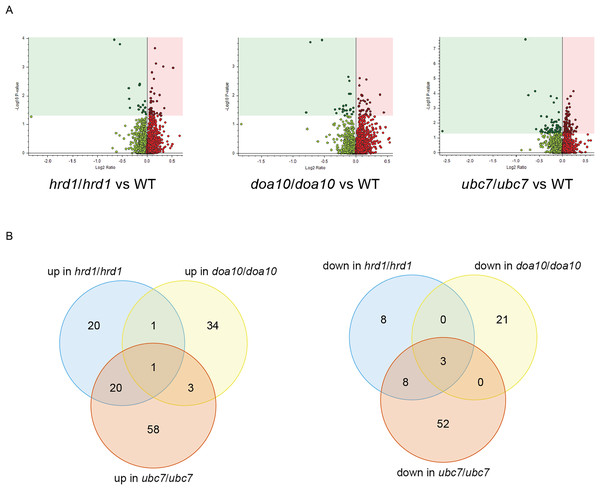
To identify candidate physiological substrates and interactors of ERAD enzymes, we identified proteins with altered abundance in yeast lacking Ca_HRD1, Ca_DOA10, or Ca_UBC7 relative to wild type yeast using quantitative tandem mass tag (TMT) mass spectrometry. The complete results of these experiments are presented in Supplemental File 3 and summarized in Fig. 6, Tables 1–3 and Tables S2–S8. In total, 229 proteins exhibited significant changes in abundance in at least one of the three mutants analyzed (Fig. 6A; Table 1). Compared to wild type yeast, 42 proteins exhibited a statistically significant increase and 19 proteins demonstrated a significant decrease in abundance in Ca_hrd1/Ca_hrd1 yeast. In Ca_doa10/Ca_doa10 yeast, 39 proteins were significantly elevated, and 24 were significantly downregulated relative to wild type yeast. Consistent with a broader role for CaUbc7, a greater number of C. albicans proteins exhibited significant alterations in Ca_ubc7/Ca_ubc7 yeast (82 increased, 63 decreased) compared to wild type yeast. The proteins with the largest significant increases in abundance in each mutant strain are presented in Table 2. 52% of proteins altered in Ca_hrd1/Ca_hrd1 yeast, and 11% of proteins altered in Ca_doa10/Ca_doa10 yeast, exhibited coordinated shifts in abundance in Ca_ubc7/Ca_ubc7 yeast (Fig. 6B; Tables S3 and S4). Seven proteins exhibited significant, opposite-direction changes in abundance in multiple mutants (Table S5). Loss of HRD1 caused a greater enrichment of predicted ER-targeted proteins (i.e., proteins with signal peptides and/or transmembrane segments) than did loss of DOA10 (Table 3).
Figure 6: Proteomic analysis of C. albicans ERAD mutants.
(A) Volcano plots illustrating changes in protein abundance between ERAD mutants and wild type C. albicans vs statistical significance of differences in abundance. Shaded regions in plots indicate data points for which P < 0.05. (B) Venn diagrams illustrating proteins present in increased (left) or decreased (right) abundance in indicated C. albicans mutants relative to wild type yeast.| # of C. albicans proteins with significant differences in abundance | # of C. albicans proteins significantly increased in mutant | # of C. albicans proteins significantly decreased in mutant | |
|---|---|---|---|
| hrd1/hrd1 vs WT | 61 | 42 | 19 |
| doa10/doa10 vs WT | 63 | 39 | 24 |
| ubc7/ubc7 vs WT | 145 | 82 | 63 |
| hrd1/hrd1 | doa10/doa10 | ubc7/ubc7 |
|---|---|---|
| orf19.1186 | Rct1 (Ynl208w) | orf19.1186 |
| Rct1 (Ynl208w) | Ptk2 (Ptk2) | Sfp1 (Sfp1) |
| CR_06510W | Wal1 (Las17) | Sod3 (Sod2) |
| Tlo16 | Hsp70 (Ssa4) | orf19.5825.1 (Yos1) |
| Tlo34 | Cta26 | Tip20 (Tip20) |
| Tlo9 | Cta24 | orf19.252 (Mpc1) |
| Cta2 | Tlo1 | Rfc52 (Rfc5) |
| orf19.3140.1 (Ump1) | Tlo11 | Erg25 (Erg25) |
| Erg3 (Erg3) | Tlo8 | Oct1 (Oct1) |
| Taf145 (Taf1) | Tlo13 | Erg3 (Erg3) |
Note:
S. cerevisiae best hits or orthologs (per Candida Genome Database ortholog finder) are indicated in parentheses.
| Increased abundance in hrd1/hrd1 | Decreased abundance in hrd1/hrd1 | Increased abundance in doa10/doa10 | Decreased abundance in doa10/doa10 | Increased abundance in ubc7/ubc7 | Decreased abundance in ubc7/ubc7 | Candida albicans proteome | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| # of proteins | % of proteins | # of proteins | % of proteins | # of proteins | % of proteins | # of proteins | % of proteins | # of proteins | % of proteins | # of proteins | % of proteins | # of proteins | % of proteins | |
| TM | 6 | 14.3% | 1 | 5.3% | 3 | 7.7% | 2 | 8.3% | 13 | 15.9% | 15 | 23.8% | 973 | 15.6% |
| SP | 2 | 4.8% | 0 | 0.0% | 0 | 0.0% | 1 | 4.2% | 2 | 2.4% | 3 | 4.8% | 325 | 5.2% |
| TM and SP | 1 | 2.4% | 1 | 5.3% | 0 | 0.0% | 0 | 0.0% | 1 | 1.2% | 1 | 1.6% | 112 | 1.8% |
| Globular | 33 | 78.6% | 17 | 89.5% | 36 | 92.3% | 21 | 87.5% | 65 | 79.3% | 44 | 69.8% | 4,808 | 77.3% |
| Beta | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 0 | 0.0% | 1 | 1.2% | 0 | 0.0% | 3 | 0.05% |
| Likely ER-targeted (TM + SP + TM and SP) |
9 | 21.4% | 2 | 10.5% | 3 | 7.7% | 3 | 12.5% | 16 | 19.5% | 19 | 30.2% | 1,410 | 22.7% |
| Total | 42 | 100.0% | 19 | 100.0% | 39 | 100.0% | 24 | 100.0% | 82 | 100.0% | 63 | 100.0% | 6,221 | 100.0% |
Note:
Proteins with altered abundance in C. albicans mutants were analyzed using the DeepTMHMM prediction tool (https://dtu.biolib.com/DeepTMHMM). TM, proteins with at least one predicted alpha-helical transmembrane segment. SP, proteins with predicted ER-targeting signal peptide. TM and SP, proteins with at least one predicted alpha-helical transmembrane segment and a predicted ER-targeting signal peptide. Globular, globular proteins without a predicted ER-targeting signal peptide or TM segment. Beta, protein with predicted transmembrane beta strands (i.e., beta barrel proteins).
We performed Gene Ontology (GO) analysis of genes encoding proteins with statistically significant changes in abundance in ERAD mutants (Tables S6–S8). Among proteins exhibiting increased abundance in Ca_hrd1/Ca_hrd1 yeast, genes with products that function in or are predicted to function in the ER and secretory pathway were enriched, including CaPdi1, CaKar2, CaDpm1, and CaSec61, homologs of proteins that are upregulated by the ER homeostatic unfolded protein response (UPR) in other organisms, including S. cerevisiae (Chapman, Sidrauski & Walter, 1998; Travers et al., 2000). GO analysis indicates a statistically significant enrichment of proteins mediating sterol synthesis in Ca_ubc7/Ca_ubc7 yeast (Table S8). The C-5 sterol desaturase ScErg3 is a bona fide ScHrd1/ScUbc7 substrate in S. cerevisiae (Jaenicke et al., 2011); enrichment in both Ca_hrd1/Ca_hrd1 and Ca_ubc7/Ca_ubc7 mutants indicates CaErg3 is likely to be a physiological target of the C. albicans enzymes as well. Overall, our results suggest C. albicans ERAD pathway homologs play important roles in protein quality control and regulated protein degradation.
Discussion
To our knowledge, in this study, we conducted the first functional analysis of ERAD genes in a pathogenic fungus. We generated homozygous knockouts of Ca_HRD1, Ca_DOA10, and Ca_UBC7 and compared stress resistance phenotypes and proteome composition among mutants and wild type yeast. Our results provide evidence for both quality control and regulatory function for C. albicans Hrd1, Doa10, and Ubc7.
Consistent with conserved roles in protein quality control, yeast with homozygous mutations in genes encoding CaHrd1, CaDoa10, or CaUbc7 exhibited sensitivity to hygromycin B and β-mercaptoethanol, which increase the burden of aberrant and misfolded proteins globally and in the ER, respectively. Enhanced sensitivity of mutants to proteotoxic stress at elevated temperatures (approximating human baseline and febrile temperatures) suggests a potential therapeutic vulnerability of C. albicans. It will be important to assess the impact of loss or inhibition of ERAD enzymes in animal models of infection in future studies.
Our proteomic analysis provides initial characterization of regulatory function of putative ERAD enzymes in C. albicans. Proteins with increased abundance in mutant yeast reflect candidate physiological ERAD substrates, ERAD co-factors, and other stress-response factors whose synthesis is induced as a compensatory response to defective ER protein quality control. We note additional biologically relevant proteins may exist that were not identified in this experiment; such proteins may be revealed by future analyses that include deeper offline fractionation for higher proteome coverage. Based on homology with other species, CaHrd1 and CaDoa10 are both expected to mediate ubiquitin transfer from CaUbc7 to substrate proteins. Consistent with shared substrates, we observed substantial overlap between proteins with altered abundance in Ca_ubc7/Ca_ubc7 yeast and in Ca_hrd1/Ca_hrd1 yeast and, to a lesser extent, between Ca_ubc7/Ca_ubc7 yeast and Ca_doa10/Ca_doa10 yeast (Fig. 6B; Tables S3 and S4).
In S. cerevisiae, a division of labor exists among ER protein quality control machinery such that Hrd1 promotes degradation of soluble ER luminal proteins and ER-targeted proteins that clog the Sec61 translocon, while Doa10 targets soluble cytosolic proteins. Both enzymes target misfolded or short-lived transmembrane proteins (Mehrtash & Hochstrasser, 2019). In our proteomic analysis, HRD1 deletion caused greater enrichment of predicted ER-targeted proteins (i.e., proteins with signal peptides and/or transmembrane segments) than did DOA10 deletion (Table 3), consistent with a more heavily ER-biased clientele for CaHrd1 relative to CaDoa10.
Notably, predicted ERAD-linked proteins (i.e., CaPdi1, CaKar2, CaDpm1, and CaSec61) were upregulated in both Ca_hrd1/Ca_hrd1 and Ca_ubc7/Ca_ubc7 yeast. ERAD disruption induces the UPR in other species, and homologs of these proteins are upregulated by the S. cerevisiae UPR (Chapman, Sidrauski & Walter, 1998; Travers et al., 2000). Previous studies have shown that the UPR effector protein CaHac1 undergoes characteristic non-canonical splicing, regulates gene expression, and alters cell morphology in response to ER stress. Our results provide additional evidence for conservation of ER homeostatic mechanisms in C. albicans (Wimalasena et al., 2008).
Among proteins upregulated in putative ERAD-defective C. albicans, several possess homologs present in increased abundance in comparable S. cerevisiae proteomics experiments (Foresti et al., 2014). Four proteins upregulated in Ca_hrd1/Ca_hrd1 cells have S. cerevisiae homologs exhibiting increased abundance in Sc_hrd1 yeast (CaKar2, CaErg3, CaOrf19.1796/ScYpl113C, and CaOrf19.2346/ScFmp40). Likewise, three proteins upregulated in Ca_ubc7/Ca_ubc7 C. albicans have homologs exhibiting increased abundance in Sc_ubc7 yeast (CaErg5, CaErg3, and CaErg25). One protein was upregulated in Ca_doa10/Ca_doa10 C. albicans for which an S. cerevisiae homolog was also found by mass spectrometry to be present at elevated levels in Sc_doa10 yeast (CaVtc4).
The proteins with the greatest increases in abundance in C. albicans ERAD mutants do not possess obvious homologs in H. sapiens. For instance, the protein with the greatest increase in abundance in both Ca_hrd1/Ca_hrd1 and Ca_ubc7/Ca_ubc7 yeast was the product from the uncharacterized open reading frame, Ca_orf19.1186 (Ca_C6_00270W). This protein has domains with predicted function in glycosylphosphatidyl inositol (GPI) anchor attachment to proteins, which occurs at the ER membrane. Orf19.1186 homologs are detectable in several Candida species, but not in S. cerevisiae or metazoans. Systematic analysis in S. cerevisiae has revealed both negative and positive genetic relationships between genes encoding ERAD machinery and those encoding GPI-biosynthetic enzymes (Costanzo et al., 2019; Costanzo et al., 2016). The mechanisms underlying these interactions are unknown; it has been hypothesized that misfolded proteins that accumulate in ERAD mutants impact the GPI pathway (Nakatsukasa, 2021).
The protein with the greatest increase in abundance in Ca_doa10/Ca_doa10 yeast (and the second greatest increase in abundance in Ca_hrd1/Ca_hrd1 yeast) is encoded by Ca_RCT1, an uncharacterized ORF. The molecular function of CaRct1 is unknown, but the poorly characterized S. cerevisiae homolog (ScYnl208w) has been found to associate with ribosomes (Fleischer et al., 2006). Ca_RCT1 homologs are detectable in multiple fungal, archaeal, and bacterial species, but not in metazoans. CaOrf19.1186 and CaRct1 may be ERAD substrates in C. albicans. Alternatively, increased abundance of these proteins may reflect enhanced synthesis of compensatory response factors due to loss of ERAD enzymes.
In S. cerevisiae and humans, ERAD machinery controls sterol abundance to meet cellular demands via feedback-regulated degradation of sterol biosynthetic enzymes (Foresti et al., 2013; Garza, Tran & Hampton, 2009; Hampton, Gardner & Rine, 1996; Huang & Chen, 2023; Jo et al., 2011; Stevenson, Huang & Olzmann, 2016). We observed enrichment of proteins with roles in sterol synthesis in C. albicans ERAD mutants, including CaErg3, the homolog of a bona fide S. cerevisiae ERAD substrate (Jaenicke et al., 2011). These results suggest sterol synthesis is also regulated by ERAD in C. albicans. Two distinct aspects of ergosterol biology are targeted by antifungal therapeutic agents, including amphotericin B and azoles. This link could provide avenues for development of novel antifungal drugs and identification of novel drug targets. Our findings highlight the need for future studies to validate and characterize the relationship between C. albicans ERAD biology and sterol production and to interrogate molecular similarities and differences of ERAD mechanisms in humans and C. albicans.
Conclusions
To our knowledge, we provide the first-ever characterization of genes predicted to encode ERAD enzymes in a pathogenic fungus. Homozygous deletion of Ca_HRD1, Ca_DOA10, or Ca_UBC7 sensitized C. albicans to proteotoxic stress and altered the proteome in distinct, but overlapping, ways. Among other perturbations, ERAD disruption increased the abundance of sterol-biosynthetic enzymes. These results strongly suggest ERAD machinery performs both protein quality control and regulatory functions. Future work will be conducted to characterize putative physiological substrates, co-factors, and compensatory stress response factors that exhibited altered abundance in C. albicans ERAD mutants. Heightened sensitivity of C. albicans ERAD mutants to proteotoxic stress at physiologically relevant temperatures suggests a potential therapeutic vulnerability. The role of ER homeostatic mechanisms in moderating virulence will be investigated in subsequent studies.
Supplemental Information
CLUSTAL multiple sequence alignment of amino acid sequence of Hrd1 homologs in H. sapiens, S. cerevisiae, and C. albicans.
CLUSTAL multiple sequence alignment of amino acid sequence of Doa10 homologs in H. sapiens, S. cerevisiae, and C. albicans.
Sequence identity and similarity of C. albicans, S. cerevisiae, and H. sapiens homologs of ERAD enzymes Hrd1, Doa10, and Ubc7.
Full alignments are available in Supplemental File 1 (Hrd1), Supplemental File 2 (Doa10), and Figure 2D (Ubc7).
Proteins with significant increases in abundance in multiple C. albicans mutants.
Proteins with significant decreases in abundance in multiple C. albicans mutants.
Proteins with significant, opposite-direction changes in abundance in multiple C. albicans mutants.
Gene Ontology Term analysis for proteins present in increased abundance in C. albicans hrd1/hrd1 mutants.
Genes encoding proteins with altered abundance in hrd1/hrd1 C. albicans mutants were analyzed using the Gene Ontology Term Finder at the Candida Genome Database ( http://www.candidagenome.org/cgi-bin/GO/goTermFinder ). No significant Process, Function, or Component GO Terms were found for proteins present in decreased abundance in hrd1/hrd1 mutants.
Gene Ontology Term analysis for proteins present in decreased abundance in C. albicans doa10/doa10 mutants.
Genes encoding proteins with altered abundance in doa10/doa10 C. albicans mutants were analyzed using the Gene Ontology Term Finder at the Candida Genome Database ( http://www.candidagenome.org/cgi-bin/GO/goTermFinder ). No significant Process, Function, or Component GO Terms were found for proteins present in increased abundance in doa10/doa10 mutants. No significant Function or Component GO Terms were found for proteins present in decreased abundance in doa10/doa10 mutants.
Gene Ontology Term analysis for proteins present in increased abundance in C. albicans ubc7/ubc7mutants.
Genes encoding proteins with altered abundance in ubc7/ubc7 C. albicans mutants were analyzed using the Gene Ontology Term Finder at the Candida Genome Database ( http://www.candidagenome.org/cgi-bin/GO/goTermFinder ). No significant Process, Function, or Component GO Terms were found for proteins present in decreased abundance in ubc7/ubc7 mutants.