Investigation and molecular identification of Eimeria sp. sampled from captive forest musk deer
- Published
- Accepted
- Received
- Academic Editor
- Erika Braga
- Subject Areas
- Biotechnology, Parasitology
- Keywords
- Captive forest musk deer, Eimeria sp., Prevalence, Molecular methods, 18S rRNA
- Copyright
- © 2021 Ren et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2021. Investigation and molecular identification of Eimeria sp. sampled from captive forest musk deer. PeerJ 9:e11751 https://doi.org/10.7717/peerj.11751
Abstract
Forest musk deer (Moschus berezovskii) is an endangered, protected species in China. Intestinal coccidiosis is a significant problem for captive forest musk deer. However, there are few reports on the prevalence and molecular characteristics of Eimeria sp. in forest musk deer. We sought to investigate the prevalence of Eimeria sp. in forest musk deer in the Sichuan and Shaanxi provinces in China. We also investigated the molecular characteristics of Eimeria sp. by analyzing the 18S rRNA gene. We collected a total of 328 fecal samples from forest musk deer on seven farms throughout the Sichuan and Shaanxi provinces. We extracted this parasite’s DNA and used this as a template for nested PCR amplification. The 18S rRNA gene fragment was associated with the plasmid vector, and these products were introduced into Escherichia coli (DH5α). The cultured bacterial solution was used as a PCR reaction template for identification purposes. We collected 328 fecal samples from forest musk deer in Lixian (n = 54), Maoxian (n = 52), Ma’erkang (n = 49), Dujiangyan (n = 55), Hanyuan (n = 41), Luding (n = 36) and Weinan (n = 41). One hundred ninety-eight (60.37%) fecal samples tested positive for Eimeria sp. . In our analysis of the 18S rRNA gene we found 34 types of Eimeria sp. with a similarity of 90.5–100%. We constructed a phylogenetic tree based on the parasite’s 18S rRNA gene sequence. Our findings indicated that the Eimeria sp. that parasitized the intestinal tract of forest musk deer was closely related to Eimeria alabamensis from Bos taurus and Eimeria ahsata from Ovis aries. To the best of our knowledge, ours was the first investigation and molecular identification of Eimeria sp. sampled from captive forest musk deer in China. Our results provide epidemiological data for the monitoring and prevention of Eimeria sp. in captive forest musk deer.
Introduction
The forest musk deer (Moschus berezovskii) is a medium-sized mammal found in alpine forests (Zhao et al., 2011; Zhao et al., 2020a). This deer (Moschus spp.) is an endangered species in China and is listed as a class I protected species nation wide (Wemmer, 1988; Xiuxiang et al., 2006; Yang et al., 2003). Musk is secreted from a musk gland in the groin of the adult male forest musk deer; this musk is of economic and medicinal value (Tian et al., 2017; Zhao et al., 2020b). The forest musk deer population has decreased sharply in recent years due to habitat destruction and other unfavorable factors (Yang et al., 2003), leading China to conduct research on artificial breeding programs in Sichuan, Shaanxi, and other regions (Li et al., 2018; Li et al., 2020). Forest musk deer, however, have strict breeding requirements (Liu et al., 2019; Qi et al., 2011) and there is little in the way of disease prevention and control for herd diseases in this species. The population of captive forest musk deer is very low and large-scale breeding of ruminants similar to cattle and sheep has not been successful to date (Xi et al., 2014).
Coccidia have been described in a wide range of vertebrate hosts (Fehlberg et al., 2018; Skirnisson & Cuyler, 2016). Infections by several species can cause severe gastrointestinal symptoms in certain hosts, which is of concern in some animal production industries (Bawm et al., 2020; Lee et al., 2018; Yabsley & Gibbs, 2006). The genus Eimeria sp. is commonly referred to as coccidia in ruminants. Eimeria sp. species are common gastrointestinal parasites and most species of this genus colonize exclusively in the intestine (Bangoura & Bardsley, 2020). Coccidiosis is widespread among ruminants (Jolley & Bardsley, 2006). Intestinal coccidiosis in captive forest musk deer is a prevalent intestinal disease and can endanger young animals. Coccidiosis can decrease the feed utilization rate, growth rate, production performance, and capacity for musk production (Andrews, 2013). Forest musk deer that are parasitized by Eimeria sp. are emaciated, anemic, and have intestinal inflammation (Bangoura & Bardsley, 2020; Hu et al., 2016; Tomczuk et al., 2015). Sha & Cai (1994) identified two Eimeria sp. species in forest musk deer coccidia and named them Eimeriamoschus and Eimeriajinfengshanenisis. They reported a high prevalence of Eimeria sp. in captive forest musk deer farms in the Sichuan and Shaanxi provinces in China (Yonghua et al., 2016; Zhao et al., 2013). Zhao et al. (2013) detected seven Eimeria sp. species in 50 fresh fecal samples collected from captive forest musk deer from the Chongqing Institute of Drug Cultivation. The species of coccidia eggs were identified as Eimeria stiedai, Eimeria perforans, Eimeria magna, Eimeria media, Eimeria irresidua, Eimeria piriformis, and Eimeria coecicola. Lu et al. (2010) investigated parasitic infections in the intestines of wild musk deer collected in Qinghai Province. In the feces they sampled, the Eimeria sp. coccidian infection rate was 43.66% (31 of 71 samples).
The majority of studies on ruminant coccidiosis have focused on cattle, sheep, and goats, with only a few concentrating on forest musk deer. Meanwhile, studies of forest musk deer parasites have focused on case reports or prevalence surveys utilizing microscopy to study fecal samples rather than identifying the parasite using molecular tools and phylogenetic analysis. Therefore, our study sought to provide scientific experimental data for the molecular epidemiological investigation and population genetic analysis of Eimeria sp. in captive forest musk deer in China.
Materials & methods
Fecal sample collection
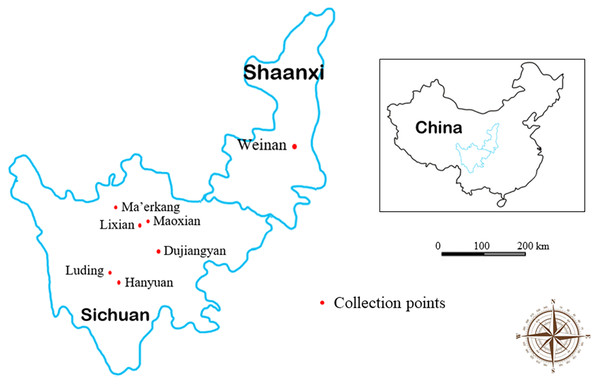
We collected fecal samples in the spring and autumn of 2018 from Lixian, Maoxian, Ma’erkang, Dujiangyan, Hanyuan, Luding, and Weinan in the Sichuan and Shaanxi provinces of China. Specific fecal sampling information is shown in Table 1 and Fig. 1. Fresh fecal samples were collected into sterile individual plastic bags using sterile disposable PE gloves and then packed in ice. The sampling place, time, longitude, latitude, and altitude were recorded at the time of sampling. Collected samples were transferred directly to the Animal Quarantine Laboratory of Sichuan Agricultural University for inspection.
Figure 1: Map of collection area of forest musk deer fecal samples in Sichuan and Shaanxi provinces, China.
| No. of sample | ||||||
|---|---|---|---|---|---|---|
| Province | Location | Longitude/E(°) | Latitude/N(°) | Altitude | Spring | Autumn |
| Sichuan | Hanyuan | 102.5800 | 29.3103 | 1068 ± 7 | 12 | 29 |
| Luding | 102.2269 | 29.7800 | 1179 ± 9 | 27 | 9 | |
| Maoxian | 103.6794 | 31.7413 | 2114 ± 11 | 25 | 27 | |
| Lixian | 103.2336 | 31.4103 | 2570 ± 10 | 13 | 41 | |
| Dujiangyan | 103.6039 | 31.0092 | 810 ± 5 | 30 | 25 | |
| Ma’erkang | 102.1211 | 31.8992 | 2601 ± 3 | 23 | 26 | |
| Shaanxi | Weinan | 109.7125 | 34.3950 | 792 ± 3 | 21 | 20 |
We collected specimens in compliance with the Animal Protection Law of the People’s Republic of China. We followed the institutional and national guidelines for the care and use of laboratory animals. Animal experiments were approved by the National Institute of Animal Health Care and Use Committee at Sichuan Agricultural University (approval number SYXK2019-187). The Sichuan Province Forestry Approval Number is (2019)264.
Parasitological examination
All fecal samples of captive forest musk deer were examined and tested for parasite eggs or oocysts using the saturated saline floating method. Positive samples showed evidence of typical eggs or oocysts. The mean eggs per gram (EPG) or oocysts per gram (OPG) were counted using the McMaster technique (Cringoli et al., 2004; Hu et al., 2017). The eggs and oocysts were examined and photographed under a microscope at 400×. We then recorded the number of coccidian eggs or oocysts (Hu et al., 2016). We identified eggs or oocysts based on their shape, size, color, and egg shell, using the national standard GB/T 18647-2002 animal coccidiosis diagnosis terminology.
DNA extraction
Fecal samples were washed with double steaming water in a vortex for 2 min at 12,000 rpm. This process was repeated three times until the supernatant was clear. Genomic DNA was then extracted from approximately 200 mg of each semi-purified product using the commercial stool DNA Kit (TD601; Tianmo, Beijing, China). DNA samples were stored in 200 μL of the kit Solution Buffer at −20 °C until ready for use (Liu et al., 2020).
Nested PCR amplification and amplicon sequencing
The extracted DNA samples were used as the templates and primers were designed according to the E. tenella sequence U67121 to amplify a region (approximately 1,500 bp) of the 18S rRNA gene. The forward primer EF1 5′-GAAACTGCGAATGGCTCATT-3′ and the reverse primer ER1 5′-CTTGCGCCTACTAGGCATTC-3′ were used during the first-round PCR reaction (Kvicerová, Pakandl & Hypsa, 2008; Youkang, Hangzhou, China). PCR was performed in a 20 μL solution containing 10 μL 2× Taq PCR Master Mix (Qingke, Beijing, China), 6 μL deionized water (Qingke, Beijing, China), 2 μL DNA, and 1 μL each of forward and reverse primers. Reaction conditions consisted of an initial denaturation phase at 95 °C for 5 min, 36 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s, extension at 72 °C for 1 min 30 s, and a final extension at 72 °C for 10 min. The second PCR was conducted with the first PCR amplification mixture as the template. The target gene (830 bp) was amplified with the forward primer EF2 5′-TTTGATGGTCATTTTTAC-3′ and the reverse primer ER2 5′-AATCCTTCTTATGTCTGG-3′. The second reaction system was the same as the first and thermocycling for the target gene was conducted with an initial denaturation step at 95 °C for 5 min, 32 cycles of denaturation at 95 °C for 30 s, annealing at 57 °C for 30 s, extension at 72 °C for 45 s, and a final extension at 72 °C for 10 min. PCR products were subjected to 1% agarose gel electrophoresis (Liu et al., 2019). The 132 18S rRNA target gene fragment was purified and placed in a T100 thermal cycler (Bio-Rad, Hercules, CA, USA) with plasmid pMD19-T vector (Katara, Beijing, China) overnight at 16 °C. The products were introduced into Escherichia coli (DH5α) (Tiangen, Beijing, China). Positive colonies were selected and inoculated with the Amp-containing liquid LB medium. The culture bacterial solution was used as a PCR reaction template for identification purposes (Youkang, Hangzhou, China).
Data analysis
The content of each base (A, T, G, C) in the sequence was analyzed using MEGA 6.0 software. The 18S rRNA sequences from different types of Eimeria sp. were compared with 18S rRNA sequences of other protozoa retrieved from GenBank in NCBI (Table 2) (https://www.ncbi.nlm.nih.gov/guide/). We used DNAMAN 6.0 software to align the sequences before counting the mutation sites in the gene sequences. Sequence similarity analysis was performed using DNAMAN 6.0 and Megalign in DNASTAR.Lasergene.v7.1. We took the 18S rRNA gene sequence of Isospora ohioensis as the outer group in the phylogenetic tree. We used the MEGA6.0 neighbor-joining method to construct the phylogenetic tree, and tree reliability was determined using bootstrap analyses of 1,000 replicates.
| Parasite type | Region | Host | GenBank accession number |
|---|---|---|---|
| Eimeria sp. | Colombia | Antillean Manatee | MG652359 |
| Eimeria faurei | Turkey | Ovis aries | AF345998 |
| Eimeria ahsata | Canada | Ovis aries | KT184334 |
| Eimeria bovis | Canada | Bos taurus | KT184336 |
| Eimeria auburnensis | Turkey | Bos taurus | KU052235 |
| Eimeria alabamensis | Canada | Bos taurus | KT184335 |
| Eimeria cylindrica | Japan | Bos taurus | AB769618 |
| Isospora ohioensis | Canada | Collie | AF029303 |
| Eimeria arloingi | Portugal | Capra aegagrus hircus | MF356556 |
| Eimeria tenella | India | Chicken | JX312808 |
| Eimeria magna | China | Rabbit | MK590202 |
Results
Microscopic examination and identification of parasites
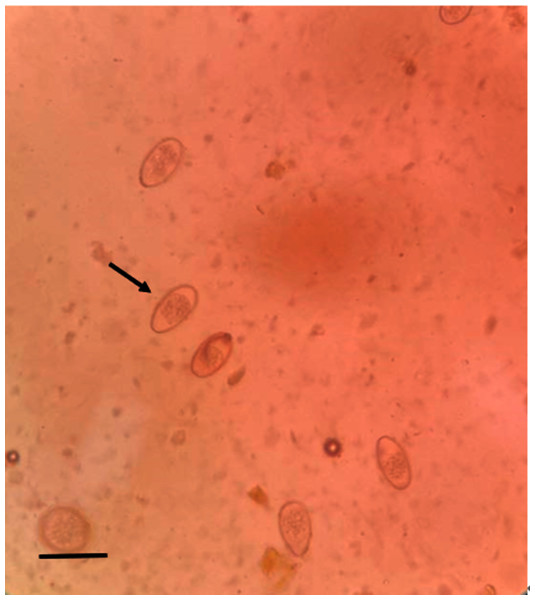
We screened 328 fecal samples using the saturated salt water floatation method. One hundred ninety-eight samples (60.37%) were found to be Eimeria sp. positive (Fig. 2). Eimeria sp. oocysts are large and oval, with yellow or brown walls. There are clear egg membrane pores, which are distributed outside the wall of the oocyst. The sporangium and sporangium remnant are observable.
Figure 2: Microscopic examination of Eimeria sp. (400×) (scale bar = 10µm).
The prevalence of Eimeria sp. in forest musk deer
Eimeria sp. was found in 60.37% of fecal samples and the OPG ranged from 200 to 98,600. Eimeria sp. in forest musk deer from seven forest musk deer farms in Lixian, Maoxian, Ma’erkang, Dujiangyan, Hanyuan and Luding and Shaanxi differed greatly. The highest prevalence was found in Ma’erkang at 91.84% (45/49), followed by Luding at 63.89% (23/36) and Maoxian at 57.69% (30/52). Eimeria sp. was similarly prevalent in Lixian and Dujiangyan at 57.14% (31/54) and 54.55% (30/55), respectively. We found 20 (48.78%) positive samples in Weinan, and the lowest prevalence was identified in Hanyuan (46.34%, 19/41).
The prevalence of Eimeria sp. is variable at different altitudes. The seven forest musk deer farms are distributed at altitudes of 500–3,000 m. Weinan and Dujiangyan are at altitudes of 0–1,000 m, Hanyuan and Luding are at an altitude of 1,000–2,000 m, and Maoxian, Lixian and Ma’erkang are at an altitude of 2,000–3,000 m. Positive rates of Eimeria sp. were 52.08% (50/96), 54.55% (42/77), and 68.38% (106/155) at altitudes of 0–1,000 m, 1,000–2,000 m, and 2,000–3,000 m, respectively. The OPG values were 100–32,600, 200–57,200, and 800–98,600, respectively. The positive rate of intestinal Eimeria sp. infection (68.38%) in captive forest musk deer at 2,000–3,000 m above sea level was significantly higher than that at 0–1,000 m (52.08%) and 1,000–2,000 m (54.55%) above sea level.
Fecal Eimeria sp. infection rates varied in the spring and autumn, and infections were more common in spring. The prevalence of Eimeria sp. in spring and autumn were 66.23% (100/328) and 55.37% (98/328), respectively. The OPG values were 200–98,600 and 100–12,600, respectively.
Nested PCR amplification of 18S rRNA
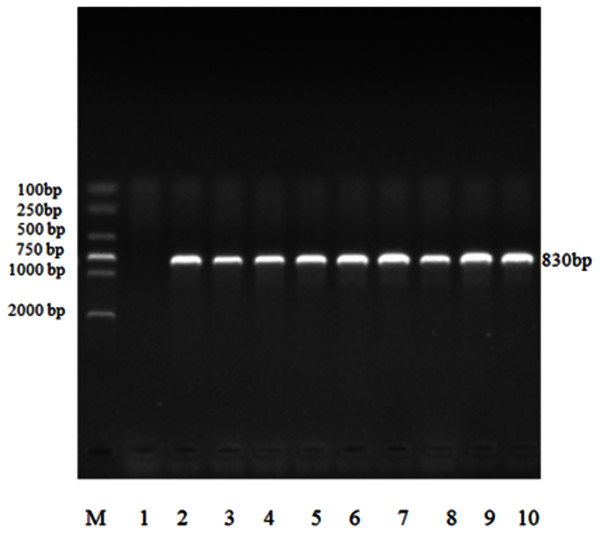
All 132 positive samples of Eimeria sp. were able to amplify 18S rRNA target gene fragment, which was approximately 830 bp, by nested PCR (Fig. 3).
Figure 3: Electrophoresis map of nested PCR of 18S rRNA gene of Eimeria sp. in forest musk deer.
PCR identification of 18S rRNA gene clone
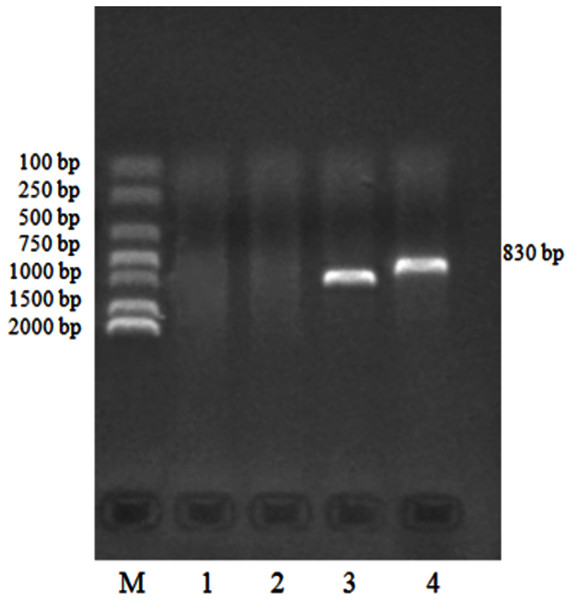
We identified 132 templates of the Eimeria sp. 18S rRNA gene clone using routine PCR. PCR products detected by agarose electrophoresis are shown in Fig. 4.
Figure 4: PCR identification of pMD19-T/18S rRNA recombinant cloning vector.
M: DNA Marker DL2000, 1:18S rRNA negative control; 2: M13 negative control; 3: M13 positive control; 4: 18S rRNA gene positive clone.Molecular identification results
We found that the 18S rRNA gene sequence results of some samples were identical after comparing and editing the sequencing results of Eimeria sp. The 132 18S rRNA gene sequences obtained by sequencing were divided into 34 species, including Maoxian (MX1-7), Dujiangyan (DJY1-5), Ma’erkang (MEK1-8), Lixian (LX1-3), Luding (LD1-2), Hanyuan (HY1-3), and Weinan (SX1-6). GenBank accession numbers are shown in Table 3. We compared and analyzed the 18S rRNA gene sequences of 34 types of Eimeria and our results showed a high intraspecific similarity (Fig. 5). There was a 90.5–100% sequence similarity of the 34 types of 18S rRNA genes. The similarity rates were as follows: Maoxian (MX1-7) was 96.8–99.0%, Dujiangyan (DJY1-5) 95.0–96.8%, Ma’erkang (MEK1-8) 96.1–99.5%, Lixian (LX1-3) 95.2–97.0%, Luding (LD1-2) 92.6%, Hanyuan (HY1-3) 95.1–100%, and Weinan (SX1-6) 98.8–99.0%. The 18S rRNA gene sequences of the 34 types of Eimeria sp. were 91.0–98.8%, which is similar to Eimeria cylindrica from Japan Bos taurus. The similarity with Eimeria faurei from Turkey Ovis aries was 91.3–99.3%; the similarity with Eimeria ahsata from Canada Ovis aries was 91–99.3%; and the similarity with Eimeria bovis and Eimeria alabamensis of Canada Bos taurus was 90.6–99.0% and 91.6–98.0%, respectively. The similarity with Eimeriidae auburnensis from Turkey Bos taurus was 91.0%--99.6%, and the similarity with Isospora ohioensis was 80.8–87.3%.
Figure 5: Sequence similarity analysis of Eimeria sp. 18S rRNA gene.
| Types | GenBank ID | Nucleotide at position | ||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 7 | 38 | 57 | 67 | 166 | 203 | 263 | 363 | 411 | 413 | 421 | 427 | 431 | 435 | 451 | 456 | 463 | 471 | 481 | 488 | 507 | 525 | 554 | 568 | 583 | 668 | 695 | ||
| MX1 | MT801007 | T | A | G | A | A | A | A | T | C | T | C | C | T | C | T | C | T | G | C | A | C | G | C | C | A | T | A |
| MX2 | MT801008 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | . | . | . | . | . | . |
| MX3 | MT801009 | . | . | A | . | . | . | . | . | T | . | . | . | C | . | . | T | . | T | . | . | G | . | . | . | G | . | . |
| MX4 | MT801010 | . | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| MX5 | MT801011 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | . | . | . | . | . | . |
| MX6 | MT801012 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | . | . | . | . | . | . |
| MX7 | MT801013 | . | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | . | . | . | . | . | . |
| DJY1 | MT801014 | . | G | . | . | . | . | T | . | . | C | . | T | C | T | . | . | . | . | . | . | . | A | T | T | . | . | . |
| DJY2 | MT801015 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | . | T | . | T | . | . | G | . | . | . | G | . | . |
| DJY3 | MT801016 | C | . | A | . | . | . | . | . | T | . | . | . | C | T | . | T | . | T | . | . | A | . | . | . | G | . | . |
| DJY4 | MT801017 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | . | T | . | T | . | . | G | . | . | . | G | G | . |
| DJY5 | MT801018 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | . | T | . | T | . | . | G | . | . | . | G | . | . |
| MRK1 | MT801019 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | . | T | . | T | . | . | G | . | . | . | G | . | . |
| MRK2 | MT801020 | . | G | . | . | . | . | T | . | . | C | . | T | C | T | . | . | . | . | . | . | . | . | . | T | . | . | . |
| MRK3 | MT801021 | . | G | . | . | . | . | . | . | . | . | . | . | . | T | . | . | . | . | . | . | . | . | . | . | . | . | . |
| MRK4 | MT801022 | . | G | . | . | . | . | . | . | . | . | . | . | . | T | . | . | . | . | . | . | . | . | . | . | . | . | . |
| MRK5 | MT801023 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | . | T | . | T | . | . | G | . | . | . | G | . | . |
| MRK6 | MT801024 | . | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| MRK7 | MT801025 | . | G | . | . | . | . | T | . | . | C | . | T | C | T | . | . | . | . | . | . | . | . | . | T | . | . | . |
| MEK8 | MT801026 | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| LX1 | MT801027 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| LX2 | MT801028 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | A | T | A | T | A | . | G | . | . | . | G | . | . |
| LX3 | MT801029 | C | . | A | . | . | . | . | . | T | . | . | . | C | A | A | T | A | T | A | . | G | . | . | . | G | . | . |
| LD1 | MT801030 | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| LD2 | MT801031 | . | C | . | G | . | . | . | . | . | . | . | . | . | . | . | . | . | T | . | . | . | A | . | . | . | . | . |
| HY1 | MT801032 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | A | T | A | T | A | . | G | . | . | . | G | . | . |
| HY2 | MT801033 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . |
| HY3 | MT801034 | C | . | A | . | . | . | . | . | T | . | . | . | C | . | A | T | A | T | A | . | G | . | . | . | G | . | . |
| SX1 | MT801035 | C | T | . | . | G | G | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | |
| SX2 | MT801036 | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | |
| SX3 | MT801037 | . | . | . | . | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | |
| SX4 | MT801038 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | |
| SX5 | MT801039 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | |
| SX6 | MT801040 | C | T | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | |
We found nucleotide variations in the 18S rRNA gene sequences belonging to 34 types of Eimeria sp. (Table 3). There was only one type of Eimeria sp. 18S rRNA gene sequence base variation in sequence base positions 67, 166, 203, 363, 554, 668, and 695. The bases at the variation positions were (A-G), (A-G), (A-G), (T-C), (C-T), (T-G), and (A-G), respectively. Only the Eimeria sp. 18S rRNA gene sequence in Weinan had no variation at positions: 57, 411, 431, 456, 507, and 583. Dujiangyan and Ma’erkang (DJY1, MEK2, MRK7) changed from base C to T at base positions 427 and 568. There was fewer difference between Ma’erkang and Weinan than that in other regions.
The Eimeriidae Isospora ohioensis 18S rRNA gene sequence was used as the outgroup of the phylogenetic tree. We constructed the phylogenetic tree from rRNA-related gene sequences using 18S rRNA gene sequences from 34 Eimeria sp. species and 18S rRNA gene sequences downloaded from NCBI. The Eimeria sp. 18S rRNA gene sequence was located in two branch systems, respectively. The sequence was under the same branch as Eimeria alabamensis from Bos taurus and Eimeria ahsata from Ovis aries, which indicated that they were closely related (Fig. 6). Eimeria tenella from chicken and Eimeria magna from rabbit were not in the same branch with the Eimeria sp. 34 species, indicating that they were not closely related.
Figure 6: Phylogenetic tree of Eimeria sp. based on the 18S rRNA gene sequence.
MX1-7, DJY1-5, MEK1-8, LX1-3, LD1-2, HY1-3, and SX1-6 are the 18S rRNA gene sequences determined by this experiment, and Isospora ohioensis is the evolutionary tree outgroup.Discussion
Artificial breeding is an effective way to sustainably utilize forest musk deer resources. Breeding operations in China are more successful than in other countries (Li et al., 2020; Shrestha, 1998). However, intestinal diseases incidence and mortality rate in captivity are still high (Li et al., 2020), which may due to Eimeria sp. infections in forest musk deer. In general, the pathogenic species tend to inhabit the posterior part of the intestines (Taylor, 2012). Coccidia can invade and destroy the intestinal epithelial cells of the host, resulting in digestive dysfunction, acute or chronic damage, and secondary damage especially in young animals (Burke et al., 2013; Saratsis et al., 2012).
The molecular data on Eimeria sp. infections in forest musk deer have not been well-documented in China. We found that the prevalence of intestinal parasites in captive forest musk deer was relatively high, which may be due to the single captive breeding mode in seven regions. The captive habitat of forest musk deer mainly consists of several individual brick cells and an outdoor activity yard, with two-to-six single brick cells supporting a larger enclosure for activity. Disease transmission is most efficient on farms with high stocking densities (Tomczuk et al., 2015). Parasites spread in forest musk deer in the same outdoor yard and may cause the animals to become chronic carriers. Infected animals without clinical symptoms remain infected throughout the year and contaminate the environment with oocysts (Bawm et al., 2020; García-Sanmartín et al., 2007). Therefore, these infected forest musk deer are the source of reinfection and transmission to other animals. Oocysts can be introduced into a susceptible herd by the fecal-oral route, contaminated clothing, boots, or cleaning tools (Alcala-Canto et al., 2020; Hu et al., 2018). An artificial enclosure does not effectively simulate the wild environment and habitat changes stress the animals. Stress can weaken the immunity of forest musk deer (Wang & Ha, 2018) and may increase parasitic infection rates.
Eimeria sp. prevalence differs greatly in each location. Factors including the climate, altitude, breeding and management mode of farms, and various region’s parasite control measures may affect the prevalence of Eimeria sp. A high breeding density and unsanity enclosures may explain the highest infection rate in Ma’erkang. The dissemination of coccidia is facilitated by high population density feeding (Tomczuk et al., 2015). Lower infection rates were found in Hanyuan and Weinan, which are known to have good deworming procedures and management methods.
It is worth noting that the feeds given to the animals in different regions were not identical, which may alter the composition of the intestinal microbial community of animals and affect the parasitism of coccidia in the intestine (Dheilly, Poulin & Thomas, 2015; Mabbott, 2018).
Environmental and climatic factors have been shown to influence the spatial distribution of parasites. Climate strongly influences the occurrence, transmission and frequency of infectious disease outbreaks. Different climatic conditions may lead to differences in the geographical distribution of parasites. Regional factors seem to have an impact on the epidemiology of parasitic diseases, which can be attributed to geographical location and climatic conditions (Acheson, Plowright & Kerr, 2015; Sun et al., 2018). Sichuan and Shaanxi are located in the southwest and central parts of China respectively. Sichuan has a mainly plateau alpine climate and subtropical monsoon climate, while the climate of Shaanxi is mainly a temperate monsoon climate. The differences of rainfall, vegetation, soil conditions and other factors due to variations in climate will affect the distribution of parasites. The altitudes of Maoxian, Lixian and Ma’erkang were higher than other areas. The moderate heat and moisture in these areas favor the development, survival and transmission of Eimeria sp. (Keeton & Navarre, 2018). Oocysts can survive for weeks to months in this extreme environment (Alcala-Canto et al., 2020).
The prevalence of Eimeria sp. is closely related to seasonality (Paoletti et al., 2017). Temperature, humidity and rainfall are the main environmental factors that affect the survival and transmission of gastrointestinal parasites (Hu et al., 2018). Eimeria sp. is more common in temperate humid environments and high temperature and humidity increases oocyte numbers and development, resulting in a high prevalence of parasites. Eimeria sp. can remain viable and infectious for at least one year in vitro and is able to withstand many adverse environmental influences because of a thick oocyst wall (Norton, 1986). However, direct exposure to ultraviolet light for a few hours or extremely dry conditions are detrimental to oocysts (Abo-Shehada & Abo-Farieha, 2003). None of the seven sampled areas had these conditions. The Sichuan and Shaanxi provinces are similar geographically and climatically. The annual average temperature is above 10 °C with seasonal temperature fluctuations. There is significantly more precipitation in spring than winter with 1,500–2,800 annual total hours of sunshine (Yu et al., 2013; Zhang, 2017). Oocysts tend to sporulate faster and disperse better in the spring with increasing precipitation (Balicka-Ramisz et al., 2012), whereas cold and dry weather during winter inhibits development and transmission (Hu et al., 2018). Accordingly, Eimeria sp. infections were more serious in spring. These findings may strengthen the seasonal prevention and control of coccidiosis through improved farm management.
The diagnosis of Eimeria sp. mainly depends on morphological examination. Our findings were confirmed using molecular techniques based on the amplification of parasite 18S rRNA. The 18S rRNA gene is highly conserved and has been widely used in the molecular detection of Eimeria sp. species. In this study, the 18S rRNA gene sequence of different species of Eimeria sp. was highly similar, indicating that the Eimeria sp. sequences parasitized in captive forest musk deer were homologous among species.
The 18S rRNA gene sequences of 34 types of Eimeria sp. in seven regions had some base variations and the similarity between the sequences was 90.5–100%. This may be due to the fact that ribosomal rRNA gene was a highly repetitive tandem sequence unit in eukaryotes and its 18S rRNA gene sequence was highly conserved in the evolutionary process (El-Sherry et al., 2013; Kvicerová, Pakandl & Hypsa, 2008; Power et al., 2009). Eimeria sp. has a variety of species and can colonizes in a wide range of hosts, such as chickens, rabbits, cattle, sheep, goats, and deer. Eimeria sp. is considered to have high host specificity and has strict selectivity to parasitic host animals (Bangoura & Bardsley, 2020). Therefore, each host animal has its own Eimeria origin in the classification of Eimeria sp. and cannot cross-infect each other (Faber et al., 2002; Jolley & Bardsley, 2006; Matjila & Penzhorn, 2002).
The phylogenetic tree of Eimeria sp. was constructed using 18S rRNA sequences. The 18S rRNA gene sequences of the 34 types of Eimeria sp. were located in two major cladistic systems. These were closely related to Eimeria alabamensis from Canada Bos taurus and Eimeria ahsata from Canada Ovis aries, but far from Eimeria tenella from chicken and Eimeria magna from rabbit. According to the phylogenetic tree, the Eimeria sp. parasitized in forest musk deer were in the same branch as most Eimeria sp. parasitized in cattle and sheep. This may be related to the fact that sheep, cattle, and musk deer are ruminants. The closer the relationship of the host animals, the closer the relationship of Eimeria sp. parasitized on the host animals, which was confirmed by the phylogenetic tree (Silva et al., 2017; Zhao & Duszynski, 2001). The sequence data obtained in this study will help to understand the genetic diversity and geographical distribution of Eimeria species that infect ruminants worldwide.
Conclusions
We determined that the prevalence of Eimeria sp.in captive forest musk deer was 60.37% (198/328) in China. To the best of our knowledge, this is the first report on the molecular identification of Eimeria sp. sampled from captive forest musk deer in China. We showed that there were significant differences in the prevalence of Eimeria sp. among different regions, altitudes, and seasons. We confirmed that the Eimeria sp. parasitized in the intestinal tract of forest musk deer was closely related to Eimeria alabamensis from Bos taurus and Eimeria ahsata from Ovis aries using the 18S rRNA gene for identification. Future studies are needed to elucidate the prevalence of Eimeria sp. infection in forest musk deer and its relationship with farm management practices, as well as seasonal and geographical changes.
There is no effective drug to treat or prevent Eimeria sp. infections in forest musk deer and measures should be taken to prevent this species from being infected with Eimeria sp. Our results suggest the use of prevention and control strategies on captive breeding farms to manage the parasite in epidemic areas and may be used to create seasonal prevention plans in different areas.