Disease diagnosis and prediction using deep learning: a review
- Published
- Accepted
- Received
- Academic Editor
- Nicole Nogoy
- Subject Areas
- Artificial Intelligence, Data Mining and Machine Learning, Data Science
- Keywords
- Machine learning, Deep learning, Disease diagnosis, Prediction, Healthcare
- Copyright
- © 2026 Krishnan and Thandava Meganathan
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ Computer Science) and either DOI or URL of the article must be cited.
- Cite this article
- 2026. Disease diagnosis and prediction using deep learning: a review. PeerJ Computer Science 12:e3484 https://doi.org/10.7717/peerj-cs.3484
Abstract
Deep learning (DL) is a machine learning technique that processes data in a manner influenced by the functioning of the human brain. It is an effective tool for deciphering complicated data and may be applied to many other processes, such as decision-making, image recognition, and natural language processing. The requirement to process large amounts of data rapidly and precisely drives the demand for deep learning technologies in the healthcare industry. Deep learning can find patterns in medical data, including genomic data, patient records, and medical imaging. Additionally, it can be utilized to create prediction models that can aid clinicians in selecting the course of treatment for patients. This article employed deep learning models to examine medical data for better diagnoses. DL models efficiently improve accuracy, handle complicated medical data, and detect subtle trends. A comparative analysis of deep learning architectures revealed that DL helps boost diagnostic accuracy and recognize subtle disease patterns. However, issues like the need for vast training data, overfitting, model interpretability, and high computational resources exist. Also, we presented the applications in diagnosing heart disease, cancer, Alzheimer’s, and other specific diseases, demonstrating the potential of deep learning in predictive modeling for clinical decision support. This article comprehensively reviews deep learning architectures and comparative research for disease identification and prediction, and explores emerging solutions such as federated learning and explainable artificial intelligence (AI). The study also tackles research obstacles and potential advantages by presenting the current status and probable future directions of deep learning in disease diagnosis and prognosis.
Introduction
Machine learning (ML) is a subgroup of artificial intelligence (AI) that allows machines to learn from complex data to perform tasks without being explicitly programmed. Several industries, including healthcare and disease diagnosis, are adopting machine learning. There are several machine learning applications in healthcare, including administration, treatment, and diagnosis. Numerous researchers and practitioners have shown that Machine Learning-Based Disease Diagnostics (MLBDD) hold the potential to be quick and affordable (Ahsan & Siddique, 2022). However, ML-based algorithms are limitless and do not experience human weariness. Traditional diagnostic approaches are often expensive, time-consuming, and require human participation. Additionally, diagnosis methods are bound by the patient’s capabilities. As an outcome, developing a technique for identifying diseases with surprisingly high patient populations in a medical context may be feasible. X-ray and magnetic resonance imaging (MRI) images, as well as tabular information about the diseases, age, and gender of patients, are used to create MLBDD systems (Ahsan et al., 2020). ML can be used to diagnose various diseases, including pneumonia, breast cancer, heart failure, and Alzheimer’s disease. ML algorithms’ emergence and the use of technology in disease diagnosis sectors demonstrate their value in the medical industry.
Deep learning (DL) has completely transformed the science of disease prediction and medical diagnosis. Medical personnel may increase the precision and speed of diagnosis, lower the cost of medical tests, and ultimately raise the standard of patient care by utilizing the potential of DL algorithms. The application of deep learning algorithms for disease diagnosis and prediction has significantly increased in recent years (Yu et al., 2023). The rationale behind investigating deep learning in disease detection and prediction is to provide researchers with a potent instrument to enhance healthcare. This study intends to serve as a beneficial reference point for anyone working in this sector and its associated domains. Deep learning has notable benefits compared to conventional approaches, such as the possibility of improved diagnosis accuracy by analyzing intricate medical data. Deep learning algorithms may be used to examine data and predict the disease risk, enabling early detection, which is crucial for effective treatment. This allows the implementation of proactive measures. Moreover, deep learning can enhance healthcare accessibility in places with limited resources or remote locations, providing significant advantages to underserved groups.
The devised approach should handle more data, better algorithms, and improved interpretability to attain high precision, a maximal Receiver Operating Characteristic (ROC), and a low false positive rate. The discovery enabled health systems to easily manage their conventional clinical diagnostics and create deep learning strategies for improving patient outcomes and lowering healthcare costs. There are many areas where standards coverage is lacking when analyzing machine learning methods for predicting medical facts for disease diagnosis. Ahmed & Husien (2024) demonstrated how ensemble learning can enhance the accuracy and robustness of heart disease prediction. However, the lack of interpretability, data biases, and little real-world validation make clinical adoption difficult. Tejaswi, Srinivasu & Gottumukkala (2025) thoroughly analyzed preprocessing techniques, dataset utilization, machine learning, and deep learning approaches for predicting lung cancer. Furthermore, practical clinical issues, including interpretability, data privacy, and healthcare integration, are unresolved. Singh & Gulati (2025) used Convolutional Neural Networks (CNNs) for feature extraction and K-Nearest Neighbors (KNN) for classification to improve accuracy. The research investigates a machine learning-based method for diagnosing and predicting chronic diseases. Combining data compression and noise reduction approaches, the model surpasses traditional methods and reaches over 93% accuracy. However, issues such as model interpretability, real-world validation, and possible biases in dataset representation need to be resolved for broader clinical use. Sia et al. (2025) integrated Support Vector Machine (SVM), random forest, KNN, and Artificial Neural Network (ANN) for symptom-based diagnosis to develop a machine learning-based disease prediction chatbot. SVM’s efficacy in disease categorization was shown by its accuracy of 92.24%. The chatbot improves user engagement by using Long Short-Term Memory (LSTM) and Natural Language Toolkit (NLT). Practical implementation is still hampered by issues in handling equivocal symptom inputs, generalizability across various populations, and real-world validation.
Savitha, Kannan & Logeswaran (2025) combined Harris Hawks Optimization (HHO) with Deep Belief Networks (DBNs) through better feature selection and hyperparameter tuning to improve the prediction of cardiovascular disease. The HHO process is optimized using the Correlation-based Weighted Compound Feature Generation (CWCFG) approach, which outperforms other metaheuristic algorithms, such as RBFO and GWO, and conventional machine learning models. Despite the model’s increased accuracy and better interpretability, issues with clinical validation, practical application, and computational complexity for widespread use still exist. Loganathan et al. (2025) introduced a MATLAB-based automated cancer detection and classification system for brain tumors, skin cancer, and lung cancer that uses CNNs. The system reduces human labor while improving diagnostic accuracy by integrating image processing, feature extraction, and segmentation. However, drawbacks include the need for high-quality medical imaging, the possibility of model bias, and difficulties with practical clinical validation. For broader application, further enhancements may concentrate on dataset variety, real-time adaptation, and interface with medical imaging systems. The main obstacles that AI-driven disease prediction models face include data biases, clinical integration problems, insufficient real-world validation, and a lack of interpretability. High-accuracy models have limited practical use due to their explainability, fairness, and scalability issues. Explainable AI (XAI), bias reduction, privacy-preserving strategies, and extensive clinical validation are needed to close these gaps. Increasing healthcare integration and optimizing computing efficiency are essential for practical implementation, as shown in Table 1. Closing these gaps is crucial to ensuring the practical implementation, transparency, and generalizability of healthcare. This study distinguishes itself from previous evaluations that concentrate on individual diseases or general deep learning methods by offering a cross-disease, modality-aware comparative examination of deep learning techniques in chronic illness diagnosis. It classifies deep learning applications by disease type and data modalities (e.g., imaging, signal, clinical text). It correlates them with suitable deep learning architectures, providing a decision-support matrix for researchers and clinicians. This review highlights significant deficiencies in the literature, including inadequate multimodal fusion strategies, poor generalizability across diverse populations, and a deficiency in explainability. It proposes a comprehensive framework that integrates transfer learning, federated learning, and explainable AI for future deep learning systems in healthcare. This comprehensive viewpoint establishes our evaluation as a strategic framework for the progression of next-generation AI-driven clinical diagnostic tools.
| Related work | Dataset | Methodology | Contributions | Drawbacks |
|---|---|---|---|---|
| Shatnawi, Abuein & Al-Quraan (2025) | Kaggle CT-scan dataset (four cancer types) | Enhanced CNN (ConvNeXt, VGG16, ResNet50, EfficientNetB0) with preprocessing |
|
|
| Khalfallah et al. (2025) | Large EEG datasets from multiple centers (UNIVERSITY OF SHEFFIELD EEG, CHB-MIT EEG, IBIB PAN EEG, AHEPA GENERAL HOSPITAL OF THESSALONIKI EEG) | CNN, ResNet, ChronoNet + Multi-head Attention |
|
|
| Alzahrani (2025) | Small, imbalanced clinical dataset (309 samples) | Conditional Tabular Generative Adversarial Network(CTGAN) + Synthetic Minority Oversampling Technique (SMOTE) + Random Forest |
|
|
| Saryazdi & Mostafaeipour (2025) | Hospital dataset (Real time -medium-sized) | Decision Tree + Fuzzy Clustering |
|
|
| Abbas et al. (2025) | 2,310 image dataset (augmented; four skin conditions) | VGG16 + LRP for explainability |
|
|
| Lopez Alcaraz et al. (2025) | MIMIC-IV ECG + ECG-View II (large ECG dataset) | XGBoost with SHAP; tree-based classifiers |
|
|
| Stabellini et al. (2025) | Cancer-specific hospital dataset | SVM, KNN, Decision Tree, Random Forest |
|
|
Although deep learning has revolutionized disease prediction and diagnosis, each model class possesses intrinsic constraints that have spurred the creation of succeeding architectures. Initial deep learning models, including fundamental ANNs, were constrained in processing high-dimensional medical data due to their superficial architectures and the vanishing gradient problem. CNNs tackled these issues by implementing local connectivity and weight sharing, facilitating the fast acquisition of hierarchical information directly from medical pictures. Nonetheless, CNNs are hampered by their static inductive biases and narrow receptive fields, which impede their ability to grasp long-range relationships across various parts of an image. Recurrent Neural Networks (RNNs) and LSTM networks were created to address the limits of temporal modeling, rendering them effective for sequential healthcare data like electrocardiogram (ECG) signals or clinical text, however, they encounter difficulties with long-range dependencies and elevated training costs. Vision Transformers (ViTs) were developed to rectify the limitations of CNNs by utilizing self-attention processes that collect global context from initial layers, enabling the modeling of distant relationships and intricate spatial dependencies in medical images. Notwithstanding their benefits, ViTs need substantial data and computing resources, prompting the development of hybrid CNN-Transformer architectures that combine the efficiency of CNNs in local feature extraction with the global representation capabilities of ViTs. Recently, ensemble learning techniques and optimization-driven hybrids, such as Deep Belief Networks (DBNs) augmented with metaheuristics, have been suggested to improve resilience and accuracy; nevertheless, these methods present issues related to interpretability and computational complexity. The progression of deep learning models in healthcare demonstrates an ongoing cycle of overcoming previous restrictions while introducing new challenges, highlighting the necessity for explainable, efficient, and clinically validated AI systems.
The main contributions of this study are as follows:
-
1.
Reviewed the most recent machine learning and deep learning architectures and methodologies that can be applied to predicting and diagnosing diseases.
-
2.
Highlighting the algorithmic steps from data collection to model deployment and continuous learning to design a DL model.
-
3.
Identified deep learning’s potential for predicting medical outcomes and disease diagnosis.
-
4.
A review of the comprehensive comparative analysis of deep learning approaches is presented.
-
5.
Opportunities, research challenges, and recent advancements related to the use of DL for disease diagnosis and prediction are discussed.
The structure of this study is as follows: In ‘Survey Methodology’ and ‘Disease Diagnosis Using Machine Learning’, survey methodology and recent research works related to disease diagnosis and prediction using ML models are discussed. DL models for disease diagnosis and prediction are discussed in ‘Deep Learning Models’. ‘Deep Learning Techniques for Disease Prediction and Diagnosis’ provides Deep Learning Techniques for Prediction and Disease Diagnosis. A comprehensive comparative analysis of the various DL approaches for disease diagnosis and prediction is provided in ‘Comprehensive Comparative Analysis of Deep Learning Approaches’. Challenges and open issues are discussed in ‘Research Challenges and Open Issues in Disease Diagnosis and Prediction ’. Recent advancements in disease diagnosis and prediction are discussed in ‘Recent Advancements in Deep Learning for Disease Diagnosis and Prediction’. The conclusion and future works are provided in ‘Conclusion and Future Work’, respectively.
Survey methodology
This review employs a systematic survey methodology to investigate the role of deep learning in disease diagnosis and prediction. Literature pertinent to the years 2018 to 2025 was obtained from sources such as IEEE Xplore, PubMed, and Scopus using specific keywords associated with deep learning in healthcare. Studies were chosen based on their use of deep learning models, clinical datasets, and documented performance measures. The search approach included phrases such as “deep learning,” “disease prediction,” “medical imaging,” “clinical diagnosis,” “healthcare AI,” “CNN,” “Recurrent Neural Network,” “transformer,” and “health informatics.” Works were classified according to disease category, data modality, and deep learning architecture. A comparative examination elucidates the strengths, limits, and new developments, including explainable AI, federated learning, and transformer-based models. This technique guarantees a thorough and analytical evaluation of deep learning’s influence on clinical decision-making and predictive healthcare.
To enhance the rigor of this survey, a systematic technique was implemented. This entails the precise delineation of the databases consulted (e.g., PubMed, IEEE Xplore, Scopus), the keywords employed, and the temporal parameters established. Explicit inclusion and exclusion criteria must be defined to guarantee impartial selection.
The study selection process followed the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines. The detailed flow of records through each phase is described as follows:
Identification
-
•
A total of 105 records were identified through database searches:
IEEE Xplore
PubMed
Scopus
-
•
No additional records were identified through manual searches or other sources.
Screening
After the removal of duplicates, 100 unique records remained.
Titles and abstracts of all 100 records were screened.
Five studies were excluded at this stage for not meeting the core objectives (e.g., absence of deep learning, irrelevant disease domain, or insufficient methodological description).
Eligibility
Ninety-five full-text articles were assessed for eligibility based on the predefined inclusion and exclusion criteria.
No additional full-text articles were excluded at this stage because the initial screening had already filtered non-relevant articles.
Included
Finally, 100 studies met all inclusion criteria and were incorporated into the qualitative synthesis (survey analysis).
Disease diagnosis using machine learning
The process of evaluating which disease or condition best explains a person’s symptoms and indicators is known as disease diagnosis. The intricate procedure includes a physical examination, information collection from the patient, and ordering the necessary tests. Several methods of machine learning for disease diagnosis exist. Algorithms based on machine learning may be used to, among other things, personalize therapy, identify trends in medical data, and predict disease risk.
Overview of machine learning
ML, a branch of AI, allows computers to “self-learn” from training data and improve over time without explicit programming. Detecting patterns in data and learning from them permits machine learning algorithms to develop their predictions. Medical experts can apply machine learning in healthcare to create better diagnostic tools for examining medical images. For instance, medical imaging (such as X-rays or MRI scans) can utilize a machine-learning algorithm for pattern recognition. Machines learn on their own without being programmed by humans. Discovering patterns and learning from ML are useful in healthcare research for better diagnostic tools and for examining images. Machine learning can address multiple disease diagnoses and prevention. ML is used in the healthcare industry in many ways, such as diagnostics, treatment, research, drug discovery, and healthcare administration.
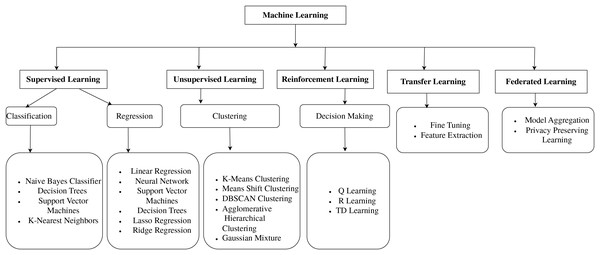
Figure 1 shows the ML models, which are classified into supervised learning, unsupervised learning, reinforcement learning, transfer learning, and federated learning. This organized overview contextualizes the methodological underpinnings on which the study’s proposed model is built, guiding readers through the reasoning behind the selection of various methodologies. In machine learning, supervised learning involves using labeled datasets to train algorithms to recognize patterns and predict outcomes (Jiang, Gradus & Rosellini, 2020). In unsupervised learning, the training datasets are not used to supervise the models throughout the machine-learning process. Instead, the models extract the insights and hidden patterns from the provided data (DataRobot, 2021).
Figure 1: Classification of machine learning models.
Machine learning techniques for disease diagnosis
New methods of disease diagnosis in medicine are being developed using ML. Figure 1 shows the number of techniques that can be used to diagnose the disease accurately, and depending on the disease, the best algorithm can be used. Here are a few related works. Heart disease is a significant health problem that affects millions of people worldwide. Early and accurate diagnosis of heart disease is essential for adequate treatment and prevention. Parkinson’s disease is a neurodegenerative disease brought on by the death of brain cells that produce dopamine. One interesting strategy being developed by researchers for the early diagnosis of Parkinson’s disease is using machine learning. A machine learning-based method created by Senturk (2020) successfully classified Parkinson’s patients with an accuracy of 93.84%. It showed that machine learning might be a valuable technique for the early diagnosis of Parkinson’s disease. Based on ML theory, notably Support Vector Machines (SVM) and Random Forests (RF), Huang, Gao & Ye (2021) created an intelligent data-driven model. The performance and accuracy of Cough Variant Asthma (CVA) diagnosis can be enhanced. Additionally, it was demonstrated that the suggested methodology was a simple way to increase the effectiveness of disease diagnosis. ML is effective in predicting several diseases, including diabetes, cancer, and heart disease, and ML algorithms are as accurate as or even more accurate than human doctors. Early diagnosis is critical for the successful treatment of many diseases. ML has the potential to diagnose diseases early when they are more curable. This could lead to improved patient outcomes and reduced healthcare costs.
Using spirometry data, Bhattacharjee et al. (2022) created machine-learning models to categorize lung disorders into obstructive and non-obstructive categories. Using 5-fold Cross-Validation (CV), models were trained using spirometry data from 1,163 patients. An additional blind dataset of 151 patients was used for external validation. With an accuracy of 83.7%, the Multi-Layer Perceptrons (MLP) model operated at peak efficiency. In summary, despite the many challenges that need to be solved, recent advances in ML have led to new challenges in the medical field, like unbalanced data, ML interpretation, and ML ethics (Ahsan et al., 2020). Deep Learning is a potential approach for disease detection and prediction, with ANN serving as the fundamental notion of machine learning. DL models can recognize intricate links and patterns in data. Advanced DL architectures like CNN and RNN can efficiently process large volumes of data in real-time.
The comparative analysis in Table 1 indicates that while each research work significantly adds to its field, there is evident fragmentation in methodological integration. It emphasizes the essential need to include interpretability, multimodal data, and empirical validation to enhance the therapeutic relevance of deep learning models. Few research studies have concurrently examined essential AI components, including explainability, privacy, generalizability, and real-world scalability. This suggests that contemporary research often prioritizes performance criteria above configuration with the broader requirements of clinical implementation. Notwithstanding sophisticated models, no comprehensive research integrates large-scale data utilization, explainable AI, Transformer models, class balancing, and federated learning elements essential for ethically responsible and transparent AI in healthcare. This indicates a considerable deficiency in cohesive, comprehensive, intelligent diagnostic systems. Consequently, there is an urgent need for multidisciplinary and multimodal AI pipelines that exhibit high performance while being interpretable, secure, and relevant to various patient demographics.
Deep learning models
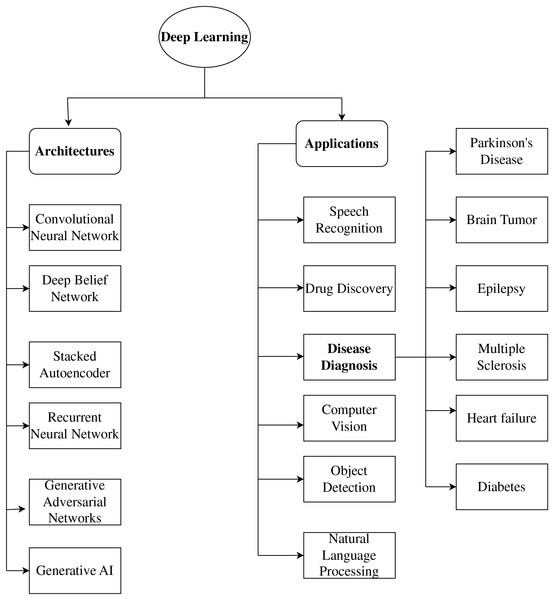
Recently, various healthcare applications, including disease diagnosis and prognosis, have begun to use deep learning. Deep learning algorithms can extract significant features helpful for diagnosis and prediction from vast volumes of data. CNNs, Recurrent Neural Networks (RNNs), and Deep Neural Networks (DNNs) are some of the deep learning techniques that have been employed for disease diagnosis and prediction. Figure 2 shows the representation of Deep learning models. A unique synthesis of deep learning model categories closely correlated with certain neurological diseases, connecting architectural design with clinical significance. Figure 2 highlights functional alignment rather than providing a generic list of models; for instance, it illustrates how generative models, such as Generative Adversarial Network (GANs), are appropriate for data augmentation in rare disease contexts, while convolutional networks are superior for spatial biomarker extraction, thus presenting a disease-specific framework for model selection that is seldom discussed in existing literature.
Figure 2: Representation of deep learning models.
Numerous deep learning architectures and methods are frequently employed to diagnose and predict diseases. CNN architectures are among the most commonly utilized architectures. CNNs are very helpful for image-based data, like that from medical imaging. CNNs can use different levels of abstraction to extract information from images that can be used for disease diagnosis and prediction (Mohades Deilami, Sadr & Tarkhan, 2022). RNN is another often-utilized model for disease diagnosis. For time-series data, like Electrocardiogram (ECG) data, RNNs are especially helpful. The temporal dependencies in the data can be modeled by RNNs and used for disease diagnosis and prediction. Various alternative deep learning architectures and methods are frequently employed for disease detection and prediction, in addition to CNNs and RNNs. These include Deep Belief Networks (DBNs), GANs, Generative Artificial Intelligence (GAI), and autoencoders, as shown in Fig. 2.
Convolutional neural networks
Convolutional Neural Networks are deep learning models used in image recognition, video analysis, Natural Language Processing (NLP), medical imaging, remote sensing, generative tasks, and speech recognition, and they need a large set of input parameters. CNNs are used for automatic feature extraction and operate well on large data sizes. CNNs are used in medical image analysis for disease diagnosis and prediction and mathematical notation is outlined in Eq. (1). Compared to MultiLayer Perceptron (MLP)s, CNNs require less memory and time to train on the data. CNN architecture contains layers such as convolutional, pooling, fully connected, and output.
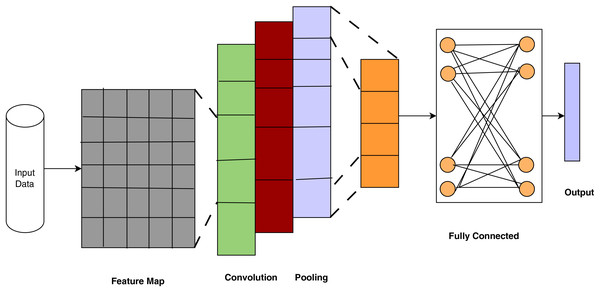
Figure 3 depicts the architecture of CNNs, which have proven essential for disease detection via deep learning. It demonstrates how CNNs systematically extract and abstract spatial elements from medical images, emulating clinical pattern recognition for disease diagnosis. Each layer represents physiologically significant modifications that facilitate precise categorization. The stratified architecture of CNNs facilitates a gradual and autonomous learning process from unprocessed medical images to precise diagnostic predictions. Wang (2024) used a CNN to diagnose breast cancer from X-ray images and achieved an accuracy of 92.1%. The result highlights the capability of deep learning to aid radiologists by offering a second opinion or automating preliminary screening, thereby alleviating the burden and enhancing early detection rates.
Figure 3: Convolutional neural networks (ul Haq et al., 2022).
In the study of medical images for disease diagnosis and prognosis, CNNs have produced encouraging results. To extract information from images and learn a hierarchical representation of the image, CNNs employ convolutional layers. CNNs have been used to diagnose several disorders, including pulmonary, cardiovascular, and cancer. Sudhish, Nair & Shailesh (2024) incorporated CNNs into the Content-Based Medical Image Retrieval (CBMIR) framework, providing a revolutionary method for disease identification and facilitating the automated, efficient, and precise retrieval of pertinent medical images. The suggested pipeline utilizes hierarchical feature extraction, multi-level gain-based selection, and sophisticated indexing algorithms to achieve optimal efficiency in managing sizeable medical image libraries, effectively tackling significant.
Recurrent neural networks
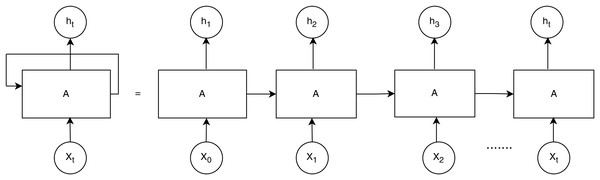
RNNs are useful for tasks requiring sequential data, such as time series or electronic health records. RNN performs better than other DL models when healthcare data includes time series properties, as shown in Fig. 4. Each unit processes the input sequentially while retaining prior context, making it effective for analyzing time-dependent clinical data, such as ECG, Electroencephalogram, or patient history and mathematical notation is outlined in Eqs. (1), (2).
Hidden state update:
(1)
Output:
(2)
Figure 4: Recurrent neural networks (ul Haq et al., 2022).
RNNs are widely used to predict changes in healthcare conditions when the variables exhibit a time series aspect. They have been utilized to predict the course of disease and the effectiveness of treatment. CNNs have extracted the geographical distribution of skin lesions to help identify skin cancer.
By examining the temporal trends in heart rate variability, RNNs have been utilized to detect cardiac disease. The spatial distribution of blood glucose levels has been analyzed using CNNs to identify diabetes. RNNs have been used to identify Alzheimer’s disease by examining the temporal patterns of brain activity. CNNs have studied the geographical distribution of skin lesions to help identify skin cancer. By reviewing the temporal trends in heart rate variability, RNNs have been utilized to detect cardiac disease. CNNs have examined the spatial distribution of blood glucose levels to identify diabetes. Kumar & Ghosh (2024) emphasized the efficacy of BiLSTM-based deep learning models for the early identification of Parkinson’s Disease using online handwriting analysis. The suggested approach integrates sophisticated kinematic feature extraction with sequential modeling, providing a robust, efficient, and accurate solution that surpasses current machine learning methods. This method boosts diagnostic precision and offers a non-invasive, accessible tool for early diagnosis, improving patient outcomes.
Long short-term memory networks
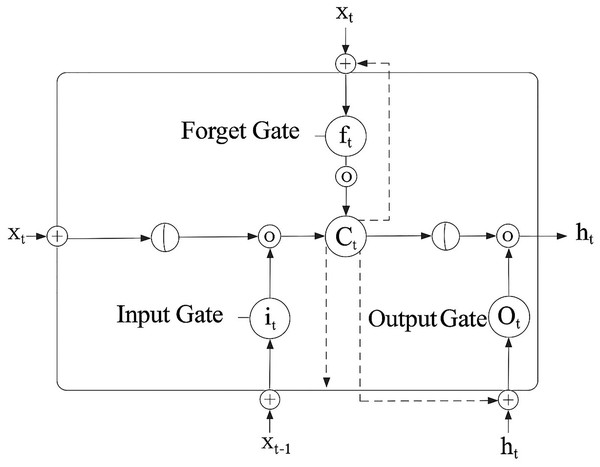
RNNs that can manage long-term data dependencies include LSTM networks. LSTMs incorporate both short- and long-term memory by integrating a gating mechanism, as shown in Fig. 5, which shows how synchronized gating mechanisms in LSTM enable the selective retention and updating of diagnostic patterns across time, essential for understanding learning progression trends in chronic disease modeling. Mathematical notation is outlined in Eqs. (3)–(6).
Forget gate:
(3) Input gate and candidate gate:
(4) Cell state update:
(5) Output gate:
(6)
Figure 5: Long short-term memory (ul Haq et al., 2022).
Oktay & Kocer (2020) proposed a convolutional LSTM approach to distinguish between Parkinsonian Tremor (PT) and Essential Tremor (ET). They used a jump motion controller with a 4D camera to record tremors and tried to extract the key characteristics using a CNN. They classified the ET and PT using the LSTM network. However, the normalization method and strategy to prevent over-fitting were not disclosed. RNNs with LSTM are particularly effective at handling jobs involving sequential data. For instance, by investigating the temporal patterns of brain activity, LSTMs are used to detect cardiac disease.
A collection of heart rate variability measurements from heart disease patients and healthy controls was used to train an LSTM model. An accuracy of 85% was achieved in the model’s ability to recognize patients with heart disease. Goyal, Rani & Singh (2024) presented a multilayered framework based on deep learning that revolutionizes Alzheimer’s Disease diagnosis, attaining enhanced accuracy via transfer learning, LSTM-based temporal modeling, and GAN-facilitated data augmentation. The approach addresses critical issues in early detection, data scarcity, and overfitting, surpassing current methodologies and paving the way for future progress in personalized medicine and AI-driven diagnostics. Its prospective applications in early detection, multimodal integration, and clinical implementation underscore its importance as a crucial instrument in combating Alzheimer’s Disease.
Generative adversarial networks
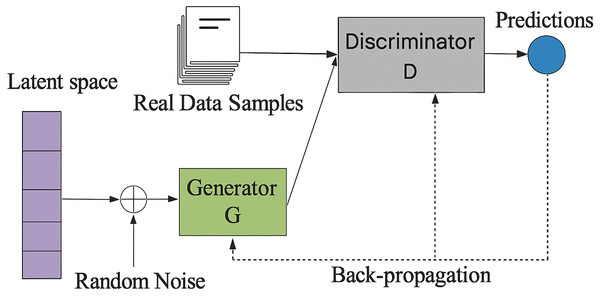
GANs are a kind of deep learning model that produces artificial data. They are frequently employed in the healthcare industry to produce artificial medical images and supplement data. Figure 6 presents a GAN that generates synthetic patient data to supplement constrained clinical datasets, thereby improving the efficacy of deep learning in diagnosing uncommon diseases. They train antagonistic neural networks side by side. There are two networks: the Discriminator and the Generator and mathematical notation is outlined in Eqs. (7)–(9).
Generator:
(7) Discriminator:
(8) Minimax Objective:
(9)
Figure 6: Framework of generative adversarial network (ul Haq et al., 2022).
The Generator creates bogus data samples to trick the Discriminator by increasing its chances of making a mistake. The Discriminator tries to discriminate between fake and actual samples. The two neural networks that build GANs are the generator and the discriminator networks. The discriminator network separates actual and artificial data, while the generator network creates artificial data.
Tufail et al. (2024) indicated substantial improvements in classification accuracy, sensitivity, and overall performance, underscoring the effectiveness of the suggested method. Using GANs to supplement minority classes improves the model’s generalizability and establishes a benchmark for using data augmentation in medical imaging. This study highlights the efficacy of integrating transfer learning with GAN-based augmentation to enhance the early identification of Alzheimer’s disease (AD), facilitating improved disease management and patient care.
Deep neural networks
An ANN with numerous layers of neurons is known as a DNN. The connections between the neurons in each layer and the subsequent layer are weighted. The connections’ weights are changed throughout the training phase so the DNN can learn how to do a particular job and mathematical notation is outlined in Eqs. (10), (11).
Layer wise transformation:
(10) Final Output:
(11)
Deep neural networks are being created for use in the healthcare industry by the research group Google DeepMind Health. Algorithms developed by DeepMind Health may be used to identify diseases, predict patient outcomes, and create customized treatment regimens (Singh, 2024). SinhaRoy & Sen (2024) improved the early diagnosis of Alzheimer’s Disease (AD) by producing synthetic Magnetic Resonance Imaging (MRI) images with Deep Convolutional Generative Adversarial Networks (DCGANs). The authors highlighted the efficacy of GAN-based methodologies in mitigating data constraints and improving diagnostic accuracy. It emphasizes an innovative method for Alzheimer’s disease prediction, providing a robust instrument for early diagnosis and enhancing the use of deep learning in medical imaging.
Generative artificial intelligence
Generative AI, which can produce new data such as text, images, or music based on training data, offers several fascinating potential uses in diagnosing and predicting diseases. It can uncover hidden patterns by analyzing vast patient data, including genetic information, scans, and medical records, and finding links and patterns humans might overlook. This capability can lead to earlier diagnoses, more precise prognoses, and even the identification of new diseases and mathematical notation is outlined in Eqs. (12), (13).
General generative model likelihood:
(12) Variational Autoencoder (VAE) objective (commonly used in GAI):
(13)
Additionally, generative AI can simulate disease progression by building models that mimic the course of a disease in a specific patient, assisting medical professionals in determining the optimal course of action and predicting a patient’s potential response to various treatments. Moreover, generative AI can produce synthetic data that resembles actual patient data, which is invaluable for developing new diagnostic tools and training other AI models (LeewayHertz & Takyar, 2024). Employing creative ideas to surpass conventional methods is crucial in healthcare. The use of GAI has been steadily rising across several domains. Balas & Micieli (2023) used generative artificial intelligence technology to produce visuals depicting the visual perception of a patient with visual snow syndrome, using textual descriptions to facilitate the text-to-image translation process. Diverse models provided a clear image, including DALL·E2, midjourney, and Stable Diffusion.
Deep learning techniques for disease prediction and diagnosis
Deep learning techniques have the power to transform the medical industry. Deep learning models can assist in improving patient outcomes by giving clinicians early and more precise diagnoses. New disease therapies and cures are also being developed using deep learning techniques. Future deep learning applications in healthcare will probably increase as deep learning technology advances. Concerning chronic disease diagnosis using DL approaches, we thoroughly reviewed the studies on heart disease, cancer, diabetes, skin, contagious, Alzheimer’s, and Parkinson’s disorders.
Overview of the dataset
Various chronic diseases and data modalities from public and commercial sources comprise the datasets used in existing works. Table 2 shows that the datasets used in medical research vary significantly in nature, availability, and accessibility, reflecting the varying data requirements of different disease areas.
| Related work | Disease (category) | Dataset (type) | Image type | Number of Images |
|---|---|---|---|---|
| Asif et al. (2025) | Brain tumor (e.g., glioma, meningioma) | Public (Kaggle brain MRI dataset) | MRI (T1-weighted scans) | 2,870 (original images; ~10,000 with augmentation) |
| Guo et al. (2023) | Pneumonia (respiratory) | Public (Kermany Kaggle & RAIG X-ray) | Chest X-ray | 5,856 (training) + 3,900 (external test) |
| Vuran et al. (2025) | Monkeypox (skin infection) | Public (Mpox Skin Lesion v2.0) | Skin lesion images (clinical) | 755 |
| Malik et al. (2024) | Skin cancer (melanoma, BCC) | Public (ISIC dermoscopy archive) | Dermoscopic images | 3,762 (training) + 1,060 (testing) |
| El-Ghany, Mahmood & Abd El-Aziz (2024) | Gastrointestinal (ulcers, polyps) | Public (Kvasir-Capsule & Kvasir v2) | Wireless capsule endoscopy | ~35,468 (Kvasir-Capsule) + 8,000 (Kvasir v2) |
| Raza et al. (2024) | Alzheimer’s disease (neurological) | Public (ADNI) | MRI (structural sMRI) | 1,075 |
| Li et al. (2025) | Cardiac diseases (e.g., cardiomyopathy) | Public (ACDC challenge data) | Cardiac MRI | 150 patients (~28–40 frames each ≈5,000 images) |
| Thatha et al. (2025) | Breast cancer (histopathology) | Public (BreakHis histology dataset) | Histopathology (H&E stained) | 7,909 |
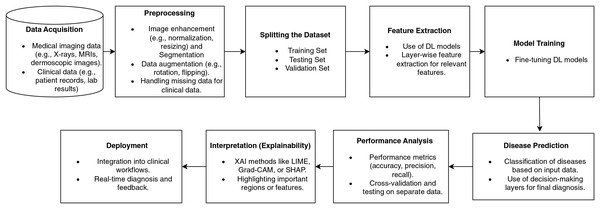
Figure 7 illustrates a disease diagnostic pipeline based on deep learning, beginning with the data collection from medical imaging and clinical records. Preprocessing improves the data by normalizing, augmenting, and handling missing variables. Feature extraction employs deep learning algorithms.
Figure 7: Disease diagnosis workflow pipeline based on deep learning.
The model undergoes training using data and is then precisely adjusted to enhance its accuracy. Disease classification predictions are assessed using performance metrics. Explainability approaches, such as Grad-CAM, emphasize significant characteristics, thereby improving the transparency of the model. The last stage involves incorporating the model into clinical operational processes, thus facilitating immediate diagnosis and bolstering healthcare decision-making. Figure 7 illustrates a comprehensive pipeline for illness detection using deep learning, highlighting the progressive integration of clinical data management, model development, and practical implementation. It distinctly integrates explainability and feedback mechanisms to enhance trust and decision-making in therapeutic environments.
Disease diagnosis and prediction
The study reviews numerous research studies that use DL methods to diagnose and forecast chronic diseases. These diseases include heart disease, cancer, diabetes, skin disorders, contagious diseases, and neurodegenerative diseases such as Alzheimer’s and Parkinson’s. The study focuses on advances in applying DL models and other deep learning architectures to improve diagnostic accuracy, early detection, and tailored treatment planning for various disorders. By evaluating varied datasets such as medical images, patient records, and genetic data, DL techniques have been demonstrated to considerably improve prediction results, allowing for quicker treatments and possibly lowering healthcare expenditures.
Heart disease
Electrocardiograms (ECGs), echocardiograms, blood pressure measurements, and imaging methods like Computed Tomography angiograms or MRI are used to identify cardiovascular diseases (CVDs), including heart disease, stroke, arrhythmias, and hypertension. Deep learning models have been used to predict cardiac disease and evaluate ECG signals to diagnose arrhythmias by analyzing numerical data such as blood pressure, heart rate, and cholesterol levels. Predicting heart disease often entails using DNNs to integrate clinical imaging (such as echocardiograms and CT scans) with patient history data (such as age, gender, and smoking status). Deep learning is used in stroke diagnosis and prediction to examine CT and MRI images and pinpoint brain regions that have been impacted by ischemia or bleeding. Deep learning models have also been used to spot minute changes in the heart’s structure, such as identifying coronary artery disease on CT scans (Rani et al., 2024). Baviskar et al. (2023) highlighted the necessity for an effective prediction approach to cope with the complications of important heart-related diseases. Uma Maheswari & Valarmathi (2023) used the Optimized Deep Belief Network to develop a prediction and recommendation model for heart disease prediction. Hybrid methods will extract the best characteristics from diverse data sources.
Cancer disease
A group of conditions known as cancer is characterized by abnormal cell proliferation and the capacity to infiltrate or spread to various body locations. These stand in contrast to normal tumors, which remain stable. Large databases of chemical compounds can be screened using deep learning to find those that can destroy cancer cells. When cancer is diagnosed sooner, it can often be treated more successfully (Joshi & Aziz, 2024). Asthma, pneumonia, COPD, lung cancer, and other respiratory conditions are excellent prospects for deep learning applications. Deep CNNs are taught to recognize nodules and categorize them as benign or malignant to detect lung cancer using CT images. Positron Emission Tomography (PET) scans may also be performed with CNNs to find metabolically active cancers. A mix of imaging and pulmonary function tests is used to diagnose asthma and COPD. Deep learning models are increasingly used to examine chest CT images and X-rays for indications of emphysema and airway blockage. Deep learning is also utilized to predict respiratory failure or exacerbations in chronic respiratory disorders using time-series data from spirometers and pulse oximeters (Khandakar et al., 2024). Imaging, biopsy, and endoscopy are often used to identify gastrointestinal disorders, such as Crohn’s disease, colorectal cancer, and Irritable Bowel Syndrome (IBS). Images from colonoscopies are analyzed using deep learning models to look for polyps or other abnormalities that might be signs of colorectal cancer. Deep neural networks are used for automated image segmentation to identify benign or malignant tumors. Colorectal cancer is also staged using CT scans, and deep learning models use changes seen in radiological imaging to predict how the tumor will react to treatment. Deep learning is used to assess the thickness of the gut wall in MRI images for Crohn’s disease and to predict flare-ups in biomarkers such as blood and stool tests. Deep learning has also helped microbiome research; algorithms now examine 16S rRNA sequencing data to find bacterial signatures associated with Crohn’s disease or IBS (Sokouti & Sokouti, 2024). Pradhan, Chawla & Rawat (2023) created a novel lung cancer diagnostic model using an optimized deep learning technique and attribute correlation-based optimized weighted feature extraction. By comparing the suggested model to other optimization and machine learning techniques, it has been shown to perform better. Additionally, the suggested SA-SLnO-RNN model has the drawback of being unable to resolve combinatorial optimization issues.
Neurological diseases: Alzheimer’s disease
Deep learning algorithms greatly enhance the intricate data sets used in neurological disorders, which impact the brain, spinal cord, and nervous system, including neuroimaging (MRI, CT, and PET scans), electroencephalography (EEG), and genetic data. To identify early alterations in brain structure from Magnetic Resonance Imaging (MRI) images, for instance, deep learning-based image processing is often used to diagnose Parkinson’s disease (PD). There has been a recent uptick in the use of deep learning models for the diagnosis of Alzheimer’s disease. These models examine genetic information, CerebroSpinal Fluid (CSF) biomarkers, and patterns in neuroimaging data, such as atrophy in specific brain areas. Deep learning networks may be taught to recognize tiny lesions that may not be apparent to human observers, allowing MRI to segment brain lesions in diagnosing multiple sclerosis. Gait analysis and voice recognition are two examples of physiological data that may be used with these technologies to enhance the monitoring and early diagnosis of neurological diseases (Hussain & Nazir, 2024).
Alzheimer’s disease is a degenerative brain disease that impairs cognition and causes memory loss. LSTMs have been used to identify Alzheimer’s disease. An LSTM model was trained using a dataset of EEG recordings from patients with Alzheimer’s disease and healthy controls. 90% of the time, the model successfully identified people with Alzheimer’s disease. Nguyen et al. (2022) suggested an ensemble learning framework for AD detection that combines deep learning, machine learning, and a multi-model, uni-data approach. The deep learning model was developed using a 3D-ResNet to benefit from 3D structural properties in neuroimaging data. Transfer learning may be employed to reduce further overfitting from the sparse training data. Ahmed, Elsharkawy & Elkorany (2023) introduced a Deep Convolutional Neural Network (DCNN) architecture based on brain MRI images for AD diagnosis. Normal Controls (NC), Mild Cognitive Impairment (MCI), and AD are distinguished using a multiclass DCNN classifier. To categorize EEG spectrum images into three groups for early AD detection, Bi & Wang (2019) have proposed an allegedly advanced discriminative deep probabilistic model with multi-task learning. Their approach comprises a multi-task learning technique with an advanced discriminative deep convolutional generative model. The developed model performs well because it connects feature extraction and classification compared to other generative models.
Parkinson’s disease
Parkinson’s disease (PD) is a progressive disorder that causes uncontrolled shaking, stiffness, and balance and coordination problems. Tanveer et al. (2022) analyzed multiple modalities, datasets, architectural configurations, and experimental setups. Using time-series data, the hybrid CNN-RNN structures have also produced precise findings in diagnosing Parkinson’s disease. The approach taken by Alissa et al. (2022) to diagnose PD focuses on identifying movement abnormalities in patients using drawing tasks. They do this by utilizing a convolutional neural network, a deep neural network architecture, to distinguish between healthy controls and PD patients. It is possible to advance the compact model into an automated single-task diagnostic tool that operates offline in real time and can be conveniently implemented in a clinical environment. The suggested systems, the AE deep features-based system, and the Mel Frequency Cepstral Coefficients-Gaussian Mixture Models (MFCC-GMM) based system had great results, approaching 100%, as empirically proved and this method, based on voice, can detect PD without having a medical test (Khaskhoussy & Ayed, 2022).
Diabetes
Diabetes mellitus is a metabolic condition in which the body experiences persistently elevated blood sugar levels. It serves as the primary energy source for the brain. Diabetes, regardless of type, can result in excess blood sugar. Blood glucose levels, body mass index (BMI), and hormone testing are often used to identify diabetes, obesity, and thyroid conditions. Deep learning may enhance diagnosis and prediction in various domains by evaluating vast numerical data with medical imaging. For instance, by examining genetic, clinical, and demographic data, deep learning can predict the likelihood of type 2 diabetes. Ultrasound imaging and blood tests (such as TSH, T3, and T4 levels) identify thyroid diagnoses, such as hypothyroidism or hyperthyroidism. Thyroid nodules and other anomalies may be found using deep learning models trained on ultrasound images. Regarding obesity, deep learning models can detect fat distribution in the body and predict consequences like type 2 diabetes or cardiovascular disease by analyzing body scans (such as CT or MRI scans) (El-Bashbishy & El-Bakry, 2024). Önal, Güraksin & Duman (2023) presented a hybrid deep learning and image processing technique based on iris images for a more objective inspection and diabetes diagnosis. The recommended method identified the iris border and automatically retrieved the pancreatic area from the iridology chart. Kurt et al. (2023) proposed a novel and successful decision support model using RNN-LSTM and Bayesian optimization to diagnose patients in the gestational diabetes (GD) risk group with 95% sensitivity and 99% specificity on the generated dataset. By obtaining 98% AUC (95% CI [0.95–1.00] and p < 0.001, the model effectively diagnosed GD.
Skin diseases
Clinical examination and biopsy are the primary methods to identify dermatological conditions, including melanoma, psoriasis, and eczema. Deep learning algorithms are being used more and more in medical images. Dermoscopy is often used to diagnose melanoma, a skin cancer, since deep learning algorithms may identify early cancer symptoms by classifying lesions based on color, shape, and texture patterns. Furthermore, image-classification networks may improve diagnosis accuracy by distinguishing benign moles from malignant melanomas. Skin biopsy and visual examination are the two methods used to diagnose the persistent skin disorder psoriasis. By examining characteristics, including skin lesions, inflammation, and scaling, deep learning has been used to categorize and track the severity of psoriasis in dermatological images. By examining dermatological images to find distinctive patterns linked to atopic dermatitis and other types of eczema, deep learning models may assist in automating the diagnostic process for eczema (Groh et al., 2024).
Contagious diseases
Blood tests, imaging, and PCR-based diagnostics are the primary methods used to detect infectious disorders such as TB, hepatitis, HIV/AIDS, and COVID-19. For instance, RT-PCR testing and chest X-rays/CT scans diagnose COVID-19. Deep learning algorithms identify the virus from radiological images by identifying bilateral lung infiltrates and distinctive ground-glass opacities. Chest X-rays and sputum smear microscopy are often employed to diagnose tuberculosis, and deep learning models are utilized to identify radiological abnormalities in the lungs and categorize them as suggestive of TB infection. While genomic sequencing is increasingly used to find viral mutations, serological testing is still employed to find biomarkers in hepatitis diagnosis. By examining genetic sequences and patient data, deep learning models are also used to predict medication resistance in hepatitis B and C (Ajagbe & Adigun, 2024).
In summary, deep learning technologies are revolutionizing the detection and categorization of many diseases in several medical areas. Deep learning models provide practical tools for early disease identification, individualized therapy, and prognosis prediction by evaluating complex datasets, including clinical and demographic data, medical imaging, genetic data, and electrophysiological signals. Deep learning’s potential to enhance healthcare outcomes expands as more diseases are researched and more varied datasets become accessible. In addition to improving diagnostic precision, this strategy helps uncover subtle patterns in data that conventional approaches can miss.
Algorithmic steps to implement deep learning models
The disease diagnosis and prediction process using deep learning encompasses many stages, including data collection, preprocessing, model selection, training, and deployment. This improves the precision of diagnoses and the ability to make predictions in medical environments. The algorithmic steps show the fundamental disease diagnosis and prediction stages for implementing deep learning models. This survey is a valuable guide for researchers and clinicians aiming to leverage deep learning in healthcare applications using algorithmic steps to design the DL model (Algorithm 1).
Model optimization and performance metrics
Hyperparameters are essential in influencing the performance, generalization, and efficiency of deep learning models in disease diagnosis. The Learning Rate (LR) is a crucial hyperparameter that regulates the magnitude of weight updates. A minimal learning rate (e.g., 0.001 for Adam or 0.01 for SGD) facilitates stable convergence and mitigates overshooting, whereas a greater rate accelerates learning but poses a danger of divergence. To maximize this parameter, researchers commonly apply tuning procedures such as grid search, cosine annealing, or cyclical learning rate scheduling. Another essential hyperparameter is the batch size, often ranging from 16 to 128 in medical imaging investigations. Reduced batch sizes enhance generalization due to increased stochasticity in gradient updates, whereas higher batch sizes optimize computational efficiency and use GPU parallelization. Researchers frequently determine batch size based on dataset dimensions and hardware limits, occasionally employing gradient accumulation when memory constraints inhibit bigger batches.
The number of epochs specifies the frequency with which a model traverses the dataset. The selection of epochs, typically ranging from 50 to 200 in medical deep learning research, is contingent upon the size and complexity of the dataset. Inadequate epochs may result in underfitting, while an excessive number of epochs poses a danger of overfitting. To mitigate this, strategies like early halting and monitoring validation loss are commonly utilized. The optimizer directly influences convergence. Adam, SGD with momentum, and RMSProp are commonly employed in disease diagnostic applications. Adam is renowned for its adjustable learning rate mechanism, which is effective on diverse medical datasets, although SGD offers more steady convergence, especially when utilized with momentum. Comparative testing is frequently used to identify the best appropriate optimizer.
Regularization methods, including dropout (rates of 0.2 to 0.5) and L1/L2 penalties (λ values from 0.0001 to 0.01), are employed to alleviate overfitting, a prevalent challenge in medical domains with insufficient data. Dropout randomly disables neurons during training, encouraging the network to acquire more resilient representations, whereas weight decay diminishes dependence on substantial weights. Cross-validation is frequently employed to optimize these parameters. The weight initialization approach is essential. Xavier (Glorot) and the initialization are commonly utilized to stabilize gradient propagation in CNNs, GANs, and DNNs, hence mitigating the hazards of disappearing or bursting gradients. Although they are frequently established by architecture, they substantially influence initial training dynamics.
In non-linear representation learning, activation functions like Rectified Linear Unit (ReLU) and Leaky ReLU are frequently employed in CNNs and RNNs, whereas Gaussian Error Linear Unit (GELU) and Swish have gained prominence in Transformer-based architectures like ViTs because of their smoother activation characteristics. The output layer often uses softmax for classification or sigmoid for binary jobs. In vision-based illness diagnosis, the size of the input picture is a significant hyperparameter. Standard dimensions, like 224 × 224 (ResNet, ViT) and 299 × 299 (Inception), are employed to reconcile spatial detail with computing efficiency. Larger medical images, such as 512 × 512 MRI or CT scans, are frequently shrunk or cropped according to model capability. In Vision Transformers, patch size (often 16 × 16 or 32 × 32) is a critical hyperparameter; smaller patches retain intricate features of lesions, whilst bigger patches decrease processing demands.
Learning rate schedulers, including step decay, cosine annealing, and ReduceLROnPlateau, are frequently utilized to dynamically modify the learning rate throughout the training process. These strategies promote convergence, mitigate stagnation, and improve generalization, especially in multimodal and unbalanced datasets. These hyperparameters together impact predicted accuracy, training efficiency, robustness, and clinical application. Meticulous selection, informed by empirical tuning methodologies such as cross-validation, Bayesian optimization, or adaptive scheduling, is crucial to enhancing the efficacy of deep learning models in disease detection.
Gupta et al. (2025) employed fivefold cross-validation (performed five times) on a coronary CT dataset, documenting average accuracy and AUC throughout the folds. The authors specifically conducted fivefold cross-validation five times to stabilize their results. Kong et al. (2025) employed 5-fold cross-validation to train deep MRI classifiers for the selection of appropriate image inputs. Formal hypothesis testing is also utilized. The glioma/MRI study employed t-tests and Mann–Whitney U-tests to compare patient groups, whereas other studies utilized paired t-tests or bootstrap tests on cross-validated ratings for comparing model variants. Huang et al. (2023) Performance indicators (accuracy, AUC, F1, etc.) are invariably accompanied by uncertainty estimates, typically represented as 95% confidence intervals. The CT model attained an AUC of around 0.95 with a 95% confidence interval of 0.92–0.97, while a transformer-based EHR model demonstrated an AUROC of 0.844 (95% CI [0.838–0.851]) on the internal test set and 0.849 (95% CI [0.846–0.851]) on an independent cohort.
Importantly, the majority of studies incorporate external validation using independent datasets. Models are evaluated on reserved cohorts or distinct hospital datasets to exhibit generalization and robustness. To identify the location of kidney stones, Ma et al. (2020) proposed a Heterogeneous Modified Artificial Neural Network (HMANN) approach that lowers noise and aids in kidney image segmentation. Three classifiers were tested to arrive at the correct prediction results for Chronic Kidney Disease (CKD), multilayer perceptron, artificial neural networks, and support vector machines and AUC, while Gupta et al. (2025) attained consistent AUCs with narrow confidence intervals on an unobserved cardiac dataset. Authors frequently observe that external testing “exhibits robustness and generalizability.” Certain research enhance reliability by conducting repeated experiments (e.g., multiple cross-validation runs) or use bootstrapping techniques. Gupta et al. (2025) specifically conducted fivefold cross-validation five times to stabilize their results. Statistical significance tests, external test-set performance, 95% confidence intervals for AUC/accuracy, and k-fold cross-validation data are all used by contemporary deep-learning diagnostic investigations to prove that their findings are reliable and repeatable.
Table 3 presents a comprehensive summary of the performance evaluation measures often utilized in deep learning-based illness detection. The table delineates classification and regression metrics, as well as segmentation and calibration measures, providing a comprehensive overview of model assessment in medical contexts. In classification tasks, accuracy, precision, recall (also known as sensitivity), specificity, F1-score, AUC-ROC, and AUC-PR are commonly used to assess correctness, sensitivity in disease detection, and threshold-independent separability of predictions. Regression measures such as Mean Absolute Error (MAE), Root Mean Square Error (RMSE), and the R² score are employed in scenarios like illness progression tracking or severity grading. Image-based applications, such as tumor and lesion detection, depend on spatial overlap metrics like the Dice Similarity Coefficient (DSC) and Intersection over Union (IoU), which quantitatively assess the concordance between predicted and actual anatomical areas.
| Metric | Formula | Clinical relevance | Common use case |
|---|---|---|---|
| Accuracy | (TP + TN)/(TP + TN + FP + FN) | Overall correctness; limited for imbalanced data | General classification |
| Precision (PPV) | TP/(TP + FP) | Reliability of positive diagnosis | Cancer detection, anomaly detection |
| Recall (Sensitivity/TPR) | TP/(TP + FN) | Ability to detect actual disease cases | Screening tasks (e.g., COVID-19, Alzheimer’s) |
| Specificity (TNR) | TN/(TN + FP) | Ability to correctly identify healthy cases | Avoiding false alarms in clinical diagnosis |
| F1-score | 2 × (Precision × Recall)/(Precision + Recall) | Balanced trade-off between precision and recall | Imbalanced disease datasets |
| AUC-ROC | Area under ROC curve | Threshold-independent measure of separability | Binary/multi-class diagnosis |
| AUC-PR | Area under Precision–Recall curve | Focused on performance of minority (disease) class | Rare disease detection |
| Confusion matrix | Counts of TP, TN, FP, FN | Provides detailed error distribution | Model bias analysis |
| MAE (Mean Absolute Error) | Average deviation in predictions | Disease progression (e.g., Parkinson’s severity score) | |
| RMSE (Root Mean Square Error) | Penalizes larger errors | Clinical risk score prediction | |
| R2Score | Measures variance explained by model | Disease severity regression | |
| Dice Similarity Coefficient (DSC) | Overlap of predicted vs. true regions | Tumor/lesion segmentation | |
| IoU (Intersection over Union) | Segmentation boundary accuracy | MRI/CT-based lesion detection | |
| Brier score | Probability calibration | Risk prediction models | |
| Calibration curve | Plot predicted vs. observed probabilities | Evaluates the probability reliability | Clinical decision support |
Furthermore, probabilistic metrics such as the Brier Score and calibration curves assess the dependability of anticipated probabilities, hence enhancing confidence in clinical decision-making. By summarizing the formulae, clinical significance, and popular use cases, the table not only gives clarity on metric selection but also stresses the need of employing many complementing measures for a full and clinically useful evaluation of deep learning models.
Comprehensive comparative analysis of deep learning approaches
Deep learning models can uncover intricate relationships and patterns in data that would be challenging or impossible for conventional machine learning models to discover. Table 4 compares deep learning algorithms, including approaches, datasets, models, performance indicators, contributions, and limitations for other diseases. This comparative analysis highlights numerous methodological advancements observed in recent disease diagnostic research, such as improved model architectures, enhanced feature extraction strategies, integration of transfer learning, and the adoption of hybrid and multimodal learning techniques. It elucidates domain-specific patterns, model intricacies, and the trade-offs among accuracy, interpretability, and scalability. It highlighted persistent constraints, including insufficient explainability, restricted dataset variety, and the need for multimodal or privacy-preserving methodologies. This study demonstrates that deep learning models like DBN, LSTM, CNN, and GANs are highly effective in diagnosing and predicting various medical conditions. The models can handle complex medical data with ease and provide superior results. Overall, DL approaches produced better performance, achieving top accuracy in prostate (99.07%), lung (99.8%), and potentially breast cancer (98.96%). Notably, a machine learning method achieved high accuracy in Alzheimer’s disease (98.68%). Overall, models that use advanced optimization techniques, transfer learning, and neural network designs, such as CNNs, RNNs, and hybrid models, are better at handling medical datasets and achieving high performance levels across various disorders.
| Disease type/Related work | Dataset and type of data used | DL models | Performance metrics (%) | Contributions | Drawbacks |
|---|---|---|---|---|---|
| Heart Disease (Uma Maheswari & Valarmathi, 2023; Almazroi et al., 2023; DeGroat et al., 2024; Alshraideh et al., 2024) | Cleveland Heart Disease Dataset (Numeric Format) | DBN with SVM | Accuracy: 97.91% | • DBN-SVM integrates DBNs for feature extraction and SVMs for classification. | • SVM scalability issues |
| • Excellent at processing organized numerical data and obtaining high accuracy. | • High computational complexity | ||||
| UCI Repository (Numeric Format) | LSTM | Accuracy: 82.49%, | • LSTM captures temporal patterns but struggles with identifying non-disease instances. | • Poor performance with imbalanced datasets, leading to false positives for non-disease cases. | |
| Precision: 77.08%, | • Effective at capturing temporal connections in health data. | • Class imbalance | |||
| Recall: 87.24%, F1-score: 84.41% | |||||
| UCI Repository (Numeric Format) | CNN with Bi-LSTM | Accuracy: 94.5%, Precision: 94%, Recall: 94%, | • The hybrid model combines sequential and spatial data for balanced performance. | • Higher computational cost due to the hybrid architecture. | |
| F1-score: 94% | • Combines the advantages of CNN and LSTM for spatial and sequential data. | • Complexity in model design and tuning | |||
| Real-time dataset (Numerical Format) | SVM with PSO | Accuracy: 94.3% | • Used various ML models to predict heart disease | • Lack of Retrospective Data and additional evaluation metrics | |
| • Findings significantly impact early disease detection, diagnosis, and customized therapy, which might help healthcare providers make wise choices and enhance patient outcomes. | • Data dimensionality and model interpretability | ||||
| Breast Cancer (Saber et al., 2021; Alloqmani, Abushark & Khan, 2023; Maria et al., 2023; Fatima et al., 2024) | DDSM, MIAS, and Private Dataset (Images) | Transfer Learning | Accuracy: 98.96% | • Pre-trained models enhance accuracy for small mammography datasets. | • It relies on high-quality pre-trained models; |
| • Using pre-trained models significantly decreases training time. | • Performance depends on data preprocessing. | ||||
| INbreast and MIAS (Images) | CNN | MIAS AUC: 97.36%, INbreast AUC: 94.25% | • CNNs excel at identifying malignant areas with consistent performance. | • Requires extensive training data to avoid overfitting. | |
| • Effective for spotting fine-grained patterns in image datasets. | • Lack of interpretability | ||||
| BI-RADS Dataset and SRM Medical Hospital (Images) | DMD-CNN | Accuracy: 98.2% | • Customized CNNs like DMD-CNN demonstrate high diagnostic accuracy in clinical applications. | • Limited generalizability to other datasets without retraining. | |
| • Tailored architectures improve task-specific performance. | • Increased computational demands | ||||
| Public dataset (images) | Neural Network | Accuracy: 93% | Breast Cancer Prediction Analysis • Provided comprehensive analysis. • Identified key predictors for accurate diagnosis. • This study compared traditional ML with the DL model | • A small dataset was used, and various ML models should have been explored | |
| Precision: 98% Recall: 87%, | • Overfitting risks in DL models | ||||
| F1-score: 92% | |||||
| Lung Cancer (Pradhan, Chawla & Rawat, 2023; Demiroğlu et al., 2023; Yamuna Devi et al., 2024) | Kaggle Lung Cancer Dataset (Numeric Format) | SA-SLnO | Accuracy: 96.66% | • Optimization methods like SA-SLnO improve DL models by enhancing parameter tuning and feature selection. | • Optimization approaches can be computationally expensive. |
| • Enhances feature selection and parameter optimization. | • Complexity integration with DL models | ||||
| UCI Repository (CT Images) | DarkNet-53 and DenseNet-201 with Neighborhood Component Analysis | Accuracy: 98.86% | • DenseNet-201 performs well with complex image data and integrated neighborhood analysis. | • High computational resources are needed for training. | |
| • Combines deep layers with robust analysis to increase accuracy. | • Difficult to train with limited data | ||||
| Bronchoscopy and Hamlyn lung datasets (Images) | Proposed CNN | Accuracy: 99.8% | • The proposed method is used for the early identification of lung cancer. | • Concentrated only on segmentation and classification | |
| Precision: 99.8% | • Adaptive median filter for segmentation | • Lack of model interpretability | |||
| Recall: 100% | • Investigated the malignant tumor detection method over a few currently used structures | ||||
| F1-score: 99.9% | • Focused on the edge-segmentation process | ||||
| • Classification is done using the cluster technique | |||||
| Colorectal Cancer | MCO Dataset (Images) | CRCNet | Confidence Interval (CI): 95% | • The small confidence intervals provide high dependability. | • Dependency on image quality and data preprocessing. |
| (Li et al., 2022; Mulenga et al., 2021; Sharkas & Attallah, 2024; Millward et al., 2025) | • CRCNet is reliable and well-suited for accurate diagnostic tasks. | ||||
| UCI Repository (CT Images) | t-SNE | AUC: 94% | • Effective for showing multidimensional data patterns. | • t-SNE’s outputs are non-deterministic, making it unsuitable for large-scale models. | |
| • t-SNE aids in visualizing and classifying intricate patterns in microbiome data. | |||||
| CRC-based Microbiome Datasets (Numeric Format) | DNN | Accuracy: 95% | • Flexible with both organized and unstructured data types. | • Requires significant hyperparameter tuning to achieve optimal results. | |
| • DNN models efficiently process diagnostic tasks with structured datasets. | |||||
| Austin-CRC (n = 353), RNSH-CRC (n = 1,070), MCO-CRC (n=885). (Images) | SegFormer-B0 for broad tumor segmentation. | Broad tumor segmentation F1-score: 0.95. | • Created a completely automatic and comprehensible iTIL rating methodology. | • Erroneous identification of tangentially sectioned tumor nuclei as Tumor-Infiltrating Lymphocytes (TILs). | |
| - SegFormer-B1 for TIL detection. | - TSN segmentation F1-score: Tumor 0.88, Stroma 0.91, Necrosis 0.64. | • Generalizes across many independent cohorts without the need for retraining. | • Inability to differentiate mucin from stroma. | ||
| - TIL detection F1-score: 0.59, Average Precision: 0.52. | • Pioneered pixel-level segmentation for both tumor and tumor-infiltrating lymphocytes in whole slide images of colorectal cancer. | • Marginally decreased stratification efficacy in MCO-CRC Stage III instances. | |||
| - Prognostic stratification (5-Year OS HR: 1.67, Multivariate HR: 1.37). | |||||
| Prostate Cancer (Salman et al., 2022; Yildirim et al., 2022; Iqbal et al., 2021) | Sakarya University Research Hospital Dataset (Images) | CNN | Accuracy: 97% | • Extracted essential image features with great precision. | • Overfitting risk when applied to small datasets without augmentation. |
| • CNN’s feature extraction abilities are highly effective for prostate cancer detection. | • Large data requirements | ||||
| mp-MRI Data (T2W, DWI, and ADC) (Images) | Pre-trained CNN | Accuracy: 96.09% | • Reduces training time while ensuring precision. | • Dependency on the selection of suitable pre-trained models. | |
| • Pre-trained CNNs utilize MRI-specific features efficiently, reducing retraining effort. | • Retraining is still required for optimal precision | ||||
| Public Dataset (Images) | LSTM and ResNet-101 | Accuracy: 99.07% | • High precision is achieved via the complementary qualities of ResNet and LSTM. | • Computational complexity increases significantly with model size | |
| • Combines ResNet-101’s deep-layer capabilities with LSTM for sequential processing, ensuring exceptional precision. | • Lack of model transparency | ||||
| RMSD: 2.5 mm | |||||
|
Alzheimer’s Disease (Alghamdi et al., 2025; Bi & Wang, 2019; Raghavaiah & Varadarajan, 2022; Matlani, 2024) |
ADNI Dataset (Images) | Black Widow Optimization (BWO) with Fuzzy C-Means Clustering (FCM) | Accuracy: 98.68%, Sensitivity: 97.72%, Specificity: 97.19% | • Combines optimization and clustering to enhance diagnostic sensitivity and specificity. | • There is a high dependency on parameter tuning for the optimization algorithm. |
| • Balances sensitivity and specificity for greater diagnostic accuracy. | • Dependence on the quality of data | ||||
| ADNI Dataset (Images) | Deep Convolutional Neural Network (DCNN) | Accuracy: 93.86% | • DCNN outperforms competitors, though hybrid techniques may better balance accuracy and interpretability. | • Lack of interpretability compared to simpler or hybrid models. | |
| • It extracts rich hierarchical characteristics from neuroimaging data. | • Small dataset | ||||
| Public Dataset (Images) | Contractive Slab and Spike Convolutional Deep Boltzmann Machine (CssCDBM) | Accuracy: 95.04% | • Effectively handles noise in neuroimaging datasets. | • Computationally intensive and prone to overfitting in small datasets. | |
| • CssCDBM shows resilience against noisy neuroimaging data, delivering solid performance. | • Interpretability issues | ||||
| MRI images (ADNI and OASIS dataset) | Hybrid Bi-directional Long Short-Term Memory with Artificial Neural Network | Accuracy of 99.22% | • Used Improved Wild Horse Optimization algorithm (IWHO) | ||
| (BiLSTM-ANN) | Precision: 98.26 | • Automatic Alzheimer’s Disease Diagnosis using Hybrid Deep Learning | • Comparable with other machine learning models. | ||
| Recall: 98.06-ADNI and Accuracy: 98.96% Recall | • Utilizes Improved Adaptive Weaver Filtering (IAWF) for image pre-processing. | • Future use of ensemble-based feature extraction techniques. | |||
| Sensitivity: 98.32 | • Principal Component Analysis extracts significant features from images using a Normalized Global Image Descriptor (PCA-NGIST). | ||||
| Specificity: 99.21 for the OASIS | |||||
|
Parkinson’s Disease (Alissa et al., 2022; Khaskhoussy & Ayed, 2022; Kumar & Ghosh, 2024; Hadadi & Arabani, 2024) |
Leeds Teaching Hospitals NHS Trust (Images) | CNN | Accuracy: 93.5% | • Strong at extracting features from handwriting images. | • Struggles with temporal patterns without complementary sequential models. |
| • Performs well in analyzing image-based handwriting data for Parkinson’s diagnosis. | • Missing Clinical settings | ||||
| PaHaW Dataset (Images) | Bi-LSTM | Accuracy: 100% | • Exceptional at identifying temporal relationships in sequential data. | • It may overfit small datasets without adequate validation. | |
| • Bi-LSTM achieves perfect accuracy in sequential handwriting data. | • The severity of PD classification should have been explored | ||||
| Public Dataset & MFCC-GMM (Numeric Format) | Deep features-based Autoencoder with MFCC-GMM | Accuracy: 99% | • Effective feature extraction from complex voice data. | • Requires preprocessing expertise and may underperform in noisy datasets. | |
| • It combines autoencoders and GMM to process speech features effectively for Parkinson’s prediction. | • Class imbalance | ||||
| NewHandPD dataset (Images) | Harris Hawks Optimization (HHO) with pre-trained models | Accuracy of 94.12% | • Implemented Harris Hawks Optimization algorithm for optimal hyperparameter values. | • Continuous analysis of handwriting changes can provide insights into disease control and drug effects. | |
| Precision: 94.1%, | • Outperforms other methods in 10 iterations. | • Small size of the data used | |||
| Recall: 94.24%, | • Combines a deep neural network and the Harris Hawks Optimization algorithm. | ||||
| F1-score: 94.11%, and AUC: 0.98 | |||||
| Multiple Sclerosis (Schwab & Karlen, 2020; Kaur et al., 2023; Balgetir et al., 2021; Yaghoubi et al., 2024) | Floodlight Open Study (Real-time Numeric Dataset) | Multilayer Perceptron with Neural Soft Attention | Accuracy: 88% | • Attention processes emphasize critical elements for better decision-making. | • Accuracy is moderate compared to other advanced DL methods. |
| • Neural attention enhances focus on relevant features, improving diagnostic accuracy. | • The data sample is limited in size. | ||||
| Public Dataset (Images) | DeepMS2G | Accuracy: 83% | • Excel at analyzing dynamic gait data for multiple sclerosis. | • Limited accuracy indicates scope for better architecture or training techniques. | |
| • Captures gait dynamics effectively but leaves room for performance improvement. | |||||
| Public Dataset (Numeric Format) | VGG19-SVM | Accuracy: 89.23% | • Hybrid approaches increase classification efficiency. | • Performance heavily depends on dataset quality and feature extraction. | |
| • Combines deep feature extraction with SVM for enhanced classification. | • Model Explainability | ||||
| Kashani Comprehensive MS Center in Isfahan (Images) Real-time data collected | Accuracy: 97.44% | • Mobile Net V2 Model for MS Diagnosis | • Experimentation with diverse algorithms for enhanced performance. | ||
| Transfer learning | Precision: 100% | • Introduced a transfer learning network for MS diagnosis in SLO images. | • Potential integration with other deep-learning methods. | ||
| Recall: 91.67% | • MS Diagnosis Using SLO Images and Computer Technology | ||||
| F1-score: 95.65% | • Converting color images to gray using Taylor-Coye and DWT algorithms. | ||||
| • Segmenting images and extracting retinal vessels. | |||||
| • Used two classification techniques: classic Machine Learning and transfer learning models. | |||||
| Diabetes (Naz & Ahuja, 2020; Önal, Güraksin & Duman, 2023; Gao et al., 2018; Kurt et al., 2023) | UCI Diabetes Dataset (Numeric Format) | ANN | Accuracy: 97.14% | • High flexibility to structured datasets and good prediction capability. | • Struggles with noisy or incomplete datasets and simple neural networks are used. |
| • Structured datasets are well-suited for ANN, showcasing high diabetes prediction accuracy. | |||||
| International Diabetes Federation (Images) | Deep CNN | Accuracy: 80% | • When correctly designed, CNNs are capable of learning complicated features. | • Low accuracy indicates inadequate handling of specific diabetes dataset challenges. | |
| • Deep CNN needs additional feature engineering to improve performance in diabetes datasets. | |||||
| Clinic Dataset (Numeric Format) | RNN-LSTM with Bayesian Optimization | Accuracy: 98% | • Bayesian optimization improves hyperparameter tweaking, resulting in more excellent model performance. | • Computationally expensive and sensitive to hyperparameter initialization. | |
| • Fine-tuned RNN-LSTM achieves top-tier clinical prediction accuracy for diabetes. | |||||
| Gestational Diabetes dataset (Numerical data) real-time dataset collected | RNN-LSTM with Bayesian optimization | Sensitivity: 95% | • Employed deep learning and Bayesian optimization. | • Limited sample size | |
| Specificity: 99% | • Compared SVM, RF, and RNN-LSTM methods on original and resampled datasets. | • Lack of model transparency | |||
| • RNN-LSTM with Bayesian optimization developed the most effective prediction model. |
These findings demonstrate that deep learning algorithms accurately identify and predict diseases. It is crucial to remember that these findings come from studies that utilized datasets and models. Deep learning models’ performance may change depending on the dataset and model utilized. Deep learning is a promising technology for predicting and diagnosing diseases. More research is necessary to enhance the functionality of deep learning models and increase their accessibility. Also, Table 4 shows that deep learning models are potent tools for disease detection, providing elevated accuracy, versatility, and adaptation across diverse datasets. These results indicate that sophisticated deep learning approaches, with customized data and feature engineering, may enhance model performance and clinical relevance. Hybrid architectures, optimization approaches, and transfer learning improve model performance and tackle data-specific issues. However, future initiatives must prioritize enhancing model generalizability, explainability, computing requirements, misclassification, overfitting, and handling unbalanced datasets to facilitate practical clinical use. The healthcare industry may use these findings to develop AI-driven diagnostic tools to enhance patient outcomes.
Research challenges and open issues in disease diagnosis and prediction
Despite their potency and adaptability, deep learning models encounter several substantial hurdles. Confronting these challenges necessitates a multidisciplinary strategy that encompasses data collection and preprocessing techniques, algorithmic improvements, fairness-conscious model training, interpretability methodologies, secure learning, resilient models against adversarial attacks, and collaboration with domain experts and impacted communities to advance deep learning and achieve its maximum potential. These problems are concisely summarized below (Talaei Khoei, Ould Slimane & Kaabouch, 2023).
Large dataset
Deep learning models need substantial computational resources. A basic deep learning model requires more network parameters in its architecture. The efficacy of deep learning models is significantly contingent upon the size of the training dataset. Due to their automated feature extraction capability, deep learning models need exposure to several variations of instances within the corresponding classes in the dataset to achieve effective generalization. This explains the success of deep learning in domains where large quantities of data can be readily gathered. A substantial volume of data may be amassed across several areas, such as natural language processing and computer vision, resulting in the increasing popularity of deep learning applications in these fields. Nonetheless, producing vast quantities of data poses a considerable challenge in medicine. Consequently, we cannot adequately train deep learning models using limited datasets to attain the intended objectives. Moreover, medical data analysis is more complex than other fields, such as image or voice recognition. These complexities impair the model’s capacity for effective generalization. A substantial amount of training data is hence necessary to mitigate this problem. Innovative data generation methodologies must enhance the volume of data collected and utilized for dataset formulation in the medical domain (Lopez Alcaraz et al., 2025).
Deep learning methods need extensive and varied datasets for successful generalization, since exposure to various examples aids in the acquisition of strong representations of fundamental patterns. In medical fields, databases frequently exhibit limited sizes and imbalances due to difficulties in acquiring annotated patient information. Insufficient data heightens the likelihood of overfitting, causing models to remember training samples instead of identifying genuine disease-related characteristics, which leads to diminished test performance and skewed predictions for underrepresented populations. The absence of generality is especially alarming in healthcare, since it may result in erroneous diagnosis or misclassification of uncommon illnesses. Research indicates that limited datasets “fail to generalize patterns,” resulting in unstable decision limits and diminished dependability when utilized with unfamiliar patient data.
To solve these issues, researchers apply methodologies such as transfer learning, regularization, ensemble models, and data augmentation. Data augmentation is very efficacious in improving model performance when working with limited datasets. Through artificial dataset expansion, augmentation introduces increased variability to models while maintaining diagnostic labels. Prevalent methodologies encompass geometric and photometric alterations (e.g., rotation, flipping, brightness modifications) for medical imaging, as well as Synthetic Minority Over-sampling Technique (SMOTE) or Conditional Tabular Generative Adversarial Network (CTGAN) for tabular clinical datasets. Generative models, such as GANs, are employed in advanced techniques to generate realistic MRI or X-ray images, which have been shown to improve the accuracy of Alzheimer’s and cancer identification. Data augmentation is a critical tool for the development of dependable deep learning systems in healthcare contexts with limited data, as it mitigates overfitting, improves generalization, and equilibrates class distributions.
Availability of public and real-time disease diagnosis datasets and ethical considerations
There are publicly accessible databases for various diseases; however, collecting or obtaining real-time data remains challenging, owing to ethical concerns. Patient permission is required before data collection to guarantee privacy and compliance with ethical guidelines. Real-time medical data can only be released as open source after successfully deploying the DL models, protecting patient anonymity. Access to public and real-time disease diagnosis databases is essential for improving healthcare research, especially in creating AI-driven diagnostic systems. Many datasets exist across many disease areas, each providing distinct insights into certain diseases. Access to these datasets often requires compliance with ethical principles and data-sharing agreements to safeguard patient privacy and data security. These resources provide a basis for pioneering research in disease diagnosis, enabling the creation of more precise, real-time diagnostic instruments that may eventually enhance patient outcomes (Borda et al., 2022).
Quality and availability of data
Deep learning models demand substantial quantities of labeled training data for efficient learning. Acquiring enough high-quality labeled data may be costly, time-intensive, or complex, especially in specialist fields or when handling sensitive information such as healthcare and cybersecurity. Despite several methods, like data augmentation, to produce substantial data, generating sufficient training data to meet the demands of deep learning models may sometimes be arduous. Moreover, a limited dataset might result in overfitting, causing deep learning models to excel on training data but struggle to generalize to novel data. Balancing model complexity and regularization methods to prevent overfitting while attaining effective generalization is difficult in deep learning. Furthermore, investigating methodologies to enhance data efficiency, such as few-shot, active, or semi-supervised learning, remains a prominent study domain (Talaei Khoei, Ould Slimane & Kaabouch, 2023).
Data enrichment methods and the time complexity of the model
Employing conventional data augmentation techniques facilitates the introduction of significant variances in data samples for training. Nonetheless, the quality of the enhanced instances deteriorates beyond a certain threshold of augmentation. Consequently, novel data enrichment procedures are necessary for disease diagnostic tools. Generative Adversarial Networks (GANs) are used throughout several fields, including medical image synthesis, to provide high-quality data samples. Additional strategies are necessary to synthesize high-quality data. Deep learning models for medical diagnoses must manage large amounts of data, leading to increased network complexity. This requires additional processing resources and raises the model’s computational complexity. To address model complexity, deep learning models need high-end computers with sufficient CPUs and GPUs (Shyamala & Navamani, 2024).
Lack of cohesive frameworks and suboptimal utilization of transformer architectures
Existing research works concentrated on attaining high accuracy within certain domains (e.g., lung cancer or heart disease) using single-modality data and model types. Nevertheless, they lacked a cohesive framework that amalgamated essential components such as interpretability (XAI), privacy-preserving techniques, Transformer-based topologies, and cross-modal data fusion. This fragmentation hinders the development of resilient, practically deployable systems. Despite their shown efficacy in sequential biological data, Transformers have been little investigated. Research works partly integrate attention processes; nevertheless, no research has comprehensively used Transformer models for EEG, ECG, or text-based features. This indicates a lost chance to represent intricate temporal and contextual linkages in clinical data (Shatnawi, Abuein & Al-Quraan, 2025).
Inadequate hyperparameter tuning and uneven datasets
Numerous research studies failed to provide detailed documentation of tuning methodologies. While a few research works exhibited performance enhancements via systematic tuning, others depended on default configurations, compromising repeatability and possible performance advancements. Data imbalance was a considerable challenge, particularly in the lung cancer and dermatological datasets. While CTGAN and SMOTE were used in some instances to mitigate this issue, most research failed to implement thorough class-balancing procedures or efficiently utilize data augmentation. This raises problems over the model’s generalizability to diverse real-world populations (Alzahrani, 2025).
Absence of privacy-enhancing techniques and limited external validation and generalizability
Only a few research works have investigated federated learning or blockchain as methods to facilitate collaborative learning while safeguarding patient data privacy. Considering the sensitivity of medical data, this is a significant oversight that must be rectified in further study. External validation was either absent or inadequately highlighted in the majority of research. While research works validated across many datasets, most mainly depended on single-dataset, training-testing split assessments, limiting confidence in the models’ efficacy on unobserved populations and diverse clinical environments (Abbas et al., 2025).
Transparency
Deep learning’s interpretability and explainability are significant roadblocks to deciphering complicated models. The decision-making procedures of increasingly complex deep neural networks, with their many layers and parameters, might seem like “black boxes,” making it hard to understand the reasoning behind individual predictions. The lack of transparency in areas with high stakes, such as healthcare and finance, makes it harder to trust and use these models. Finding an acceptable balance between model performance and readability is essential for researchers, regulators, and end-users to understand the model’s logic, which is necessary for making informed decisions and holding models accountable in the complex world of contemporary deep learning (Hadadi & Arabani, 2024).
The present investigation exhibits encouraging outcomes; nonetheless, it is constrained by many limitations. The dataset used was constrained in size and variety owing to the availability of annotated data, potentially impacting the model’s generalizability to wider populations. Secondly, although performance was assessed using standard measures, external validation using unobserved clinical datasets was not conducted because of limitations in data availability. Third, computing resource limits hindered the deployment of more complicated Transformer-based structures and significant hyperparameter tuning. Additionally, privacy-enhancing approaches such as federated learning or differential privacy were not implemented in this model version but are planned for future work to ensure compliance with data confidentiality regulations. Despite these difficulties, deep learning can potentially enhance disease diagnosis and prognosis. Large quantities of patient data can be analyzed using deep learning to find patterns that are invisible to human clinicians. This can help diagnose people at risk of acquiring specific diseases and result in more precise and prompt diagnoses.
Recent advancements in deep learning for disease diagnosis and prediction
The rapid growth of deep learning in medical diagnostics offers promising opportunities, although there are still obstacles to overcome. This section examines many significant advancements that tackle these difficulties and drive the field ahead. We explored transfer learning methods that utilize current knowledge to address the limitations of medical datasets. In addition, we examined techniques for handling unbalanced data sets in which specific diseases are infrequent. Furthermore, we investigated how XAI methodologies promote confidence and clarity in deep learning models used in healthcare applications. Transfer learning has substantially influenced medical imaging and diagnostics by allowing researchers to utilize pre-trained models from different tasks, such as generic X-ray categorization or ImageNet datasets, to train deep-learning models. This enables enhanced efficiency on smaller datasets unique to diseases and pre-trained models relevant to domains. To improve diagnosis accuracy even with minimal data, information from massive medical datasets uses transfer learning and foundation models like BioBERT and MedGPT. The amalgamation of Vision Transformers (ViTs) with Graph Neural Networks (GNNs) has significant promise. ViTs may improve feature extraction in high-resolution imaging techniques such as CT scans and fundus photography. At the same time, GNNs excel in modeling intricate relational frameworks in patient histories, disease ontologies, and genetic connections (Sadr & Nazari Soleimandarabi, 2022). Federated learning enables medical institutions to collectively train models without sharing raw data, thus safeguarding patient privacy while promoting the development of exact models. Its capacity to develop strong models on various datasets guarantees wider application across multiple populations and improves diagnostic accuracy. Better early disease identification, individualized treatment strategies, and real-time monitoring are all possible results of integrating federated learning into healthcare, eventually enhancing patient outcomes (Bebortta et al., 2023).
An imbalance in medical datasets can lead algorithms to prioritize the majority class, potentially overlooking essential trends. Techniques such as oversampling the underrepresented class or undersampling the overrepresented are used to balance the classes. Class-weighted loss functions can prioritize the minority class more during model training. Model generalization can be enhanced by addressing class imbalance in medical datasets using strategies such as adaptive loss functions, oversampling, and synthetic data generation (Alzahrani, 2025; Hariprasad et al., 2023). The intricate nature of deep learning models, sometimes referred to as the “black box” problem, presents a hurdle to their use in clinical environments. Scientists are now focusing on creating XAI methodologies to offer a deeper understanding of the internal mechanisms of these models. This will help increase confidence and make incorporating AI technologies into healthcare operations easier. Interpretability approaches, such as Grad-CAM, SHAP, and attention-based methods, are being used to improve confidence in deep learning models by offering insights into model decision-making (Lopez Alcaraz et al., 2025; Saravanan et al., 2023). These are just a handful of fascinating advancements in deep learning that are helping to overcome the difficulties associated with disease prediction and diagnosis. We may anticipate even more developments as research goes on, resulting in more precise, understandable, and eventually life-saving uses of AI in healthcare (Rahal et al., 2024).
Conclusion and future work
This study demonstrates that DL has the potential to revolutionize the diagnosis and prediction of diseases. It can provide significant improvements in diagnostic accuracy, early identification, individualized therapy, and overall patient outcomes, while also contributing to the reduction of healthcare costs. Recent literature syntheses indicate that a variety of architectures, including CNNs, RNNs, LSTMs, GANs, DNNs, and hybrid models, have been effectively implemented in a variety of modalities, including medical imaging, time-series physiological data, clinical records, and genetic datasets. These models have demonstrated exceptional performance in the diagnosis of cardiovascular illnesses, malignancies, neurological disorders, including Alzheimer’s and Parkinson’s disease, diabetes, dermatological afflictions, and infectious diseases. Despite these developments, the review emphasizes the persistent obstacles that impede clinical adoption, including the inadequate interpretability of model predictions, the dependence on extensive and high-quality annotated datasets, the suboptimal integration of multimodal data sources, apprehensions regarding generalizability across diverse patient populations, and the constraints of computational resources. Additionally, the absence of cohesive frameworks that integrate interpretability, scalability, and security, incomplete external validation, and restricted implementation of privacy-preserving methodologies continue to present significant obstacles to their widespread adoption in healthcare systems.
Future research must focus on overcoming these limitations by developing multimodal and context-aware learning frameworks that can incorporate genetic, textual, signal, and imaging data to offer enhanced, context-specific diagnostic insights. This will be necessary to address dataset scarcity and imbalance by enhancing data efficacy through generative AI, transfer learning, and self-supervised learning. The incorporation of explainable AI methodologies, such as Local Interpretable Model-agnostic Explanations (LIME), SHapley Additive exPlanations (SHAP), Grad-CAM, ProtoPNet, and SENN, directly into model pipelines is necessary to ensure transparency, cultivate clinician confidence, and comply with regulatory norms, thereby enhancing interpretability. The improvement of privacy-preserving methodologies, such as federated learning, differential privacy, blockchain, and secure multi-party computing, is equally important in order to facilitate collaborative training while protecting sensitive patient data. The practical applicability and accessibility of adaptive, continuously learning systems will be substantially enhanced by the development of lightweight and resource-efficient models for deployment on edge devices and in low-resource environments. The future of AI-driven diagnostic instruments will be influenced by the exploration of innovative paradigms such as Vision Transformers for medical imaging, Graph Neural Networks for relational biomedical data, and foundational models like MedGPT and BioBERT for clinical text analysis. Ultimately, the integration of forthcoming diagnostic systems into healthcare workflows must be precise, transparent, equitable, and seamless, necessitating ongoing collaboration among AI researchers, healthcare professionals, policymakers, and patients to bridge the gap between research innovation and clinical practice.