RNA-Seq analysis reveals regulatory networks driven by EpCAM overexpression in esophageal adenocarcinoma cells
- Published
- Accepted
- Received
- Academic Editor
- Vladimir Uversky
- Subject Areas
- Bioinformatics, Genomics, Molecular Biology, Oncology, Pathology
- Keywords
- RNA-seq, Esophageal cancer, Overexpression, Adhesion, PPI
- Copyright
- © 2026 Mohtar et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2026. RNA-Seq analysis reveals regulatory networks driven by EpCAM overexpression in esophageal adenocarcinoma cells. PeerJ 14:e20877 https://doi.org/10.7717/peerj.20877
Abstract
Esophageal cancer is a formidable malignancy, presenting a significant health challenge due to its widespread prevalence and associated high mortality rates. Epithelial cell adhesion molecule (EpCAM), a pro-oncogenic glycoprotein, has been identified as an upregulated protein in esophageal adenocarcinoma (ESCA) through multi-OMICS platforms. However, its functional role in ESCA remains relatively understudied. Here, we investigated the contribution of EpCAM to ESCA pathogenesis using an EpCAM-null ESCA cell line, FLO-1, as a gain-of-function model. Introduction of a recombinant EpCAM–GFP fusion construct into FLO-1 cells resulted in enhanced cell migration, adhesion, clonogenic survival, and invasive capacity, supporting a pro-tumorigenic role for EpCAM. To define EpCAM-associated regulatory networks, RNA sequencing was performed on EpCAM-overexpressing cells, revealing 797 differentially expressed genes. Functional enrichment analyses indicated significant involvement of pathways related to cell adhesion, cell motility, transmembrane activity, and neuronal-associated processes, with enrichment in plasma membrane, focal adhesion, and neuron projection terminus compartments. Protein–protein interaction network analysis identified key hub genes, including SOX2, COL1A1, LOX, COL3A1, LUM, PXDN, BDNF, NCAM1, TLR2, and CCL5, linking EpCAM signaling to PI3K–Akt, ECM–receptor interaction, and focal adhesion pathways. Importantly, quantitative polymerase chain reaction (qPCR) validation of selected hub genes confirmed significant upregulation of the extracellular matrix components COL1A1 and PXDN in EpCAM-overexpressing FLO-1 cells, supporting the transcriptomic predictions and implicating ECM remodeling as a downstream consequence of EpCAM signaling. Collectively, these findings demonstrate that EpCAM promotes aggressive cellular phenotypes in ESCA and drives transcriptional programs associated with adhesion, invasion, and extracellular matrix regulation, highlighting potential therapeutic vulnerabilities in EpCAM-driven ESCA.
Introduction
Esophageal cancer stands as an alarming malignancy with a staggering global prevalence and mortality rate. As of 2020, it ranks as the fourteenth most common cancer worldwide and claims the position of the eighth leading cause of cancer-related deaths globally (Sung et al., 2021). This disease manifests primarily in two histologic subtypes: squamous cell carcinoma (ESCC) and adenocarcinoma (ESCA). ESCC, the more prevalent subtype, originates from the flat cells lining the upper esophagus, whereas ESCA typically arises in the lower esophagus near the stomach junction. The choice of treatment for esophageal cancer hinges upon various factors, including cancer stage, subtype, and the patient’s overall health (Harada et al., 2020). While surgical resection stands as the cornerstone of treatment for early-stage disease, comprehensive approaches combining chemotherapy, radiation therapy, targeted therapy, and immunotherapy hold promise for more advanced cases. Despite intensive research efforts aimed at refining early detection methods and devising novel treatment modalities, the identification of biomarkers associated with the disease remains limited. Notably, large-scale genomic studies have uncovered significant genomic alterations in both ESCC and ESCA (Cancer Genome Atlas Research Network et al., 2017). Gene expression analyses from these studies have elucidated distinct signaling pathways, with ESCA demonstrating enhanced E-cadherin signaling and upregulation of ARF6 and FOXA pathways, while ESCC exhibits activation of Wnt, syndecan, and p63 pathways.
Epithelial cell adhesion molecule (EpCAM) emerges as a prominent cell surface glycoprotein, exhibiting high expression in various cancers, particularly those of epithelial origin, such as esophageal cancer (Kimura et al., 2007; Mohtar et al., 2020). Recent high-throughput mass-spectrometry based proteomics uncovered EpCAM overexpression in primary ESCA tumour and lymp node metastases (O’Neill et al., 2017). Given its surface localization and immunogenic properties, EpCAM has become a prime target for cancer therapy. Indeed, numerous drugs have received approval and are currently undergoing clinical trials targeting EpCAM in cancer (Baeuerle & Gires, 2007; Simon et al., 2013; Kaplon & Reichert, 2019). While EpCAM’s classical role involves mediating cell-cell adhesion, notably without belonging to the cell adhesion molecule (CAM) family, its multifaceted functions continue to be elucidated (Litvinov et al., 1994; Balzar et al., 1999).
In our research, we previously identified an ESCA metastatic cell line lacking EpCAM expression, prompting an investigation into its functional significance using this cell model (Mohtar et al., 2018). Through the generation of stable cell lines overexpressing EpCAM-GFP, we demonstrated that EpCAM overexpression enhances cancer cell fitness phenotypically in ESCA cancer cell model. RNA-seq analyses combined with integrative bioinformatics analyses identified several significant differentially expressed genes that could be important in EpCAM signalling axis and ESCA pathogenesis.
Materials and Methods
Plasmids
The EpCAM-GFP recombinant plasmid encoding mature EpCAM fused to GFP was described previously (Mohtar et al., 2018). Briefly, EpCAM cDNA was polymerase chain reaction (PCR) amplified and cloned into pEGFP-N2 mammalian expression vector. All constructs were confirmed by DNA sequencing and analyzed using the SnapGene Viewer software v.5.0.4. The resulting plasmid EpCAM-GFP thus harbors an in-frame GFP fused to C-terminal of EpCAM.
Cell culture
The human esophageal adenocarcinoma cell line, FLO-1 was obtained as a kind gift from Ted Hupp (University of Edinburgh, United Kingdom). Cells were routinely maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (Nacalai Tesque, Kyoto, Japan) and was supplemented with 10% fetal bovine serum (Gibco, Waltham, MA, USA) and 1% penicillin-streptomycin mixed solution (Nacalai Tesque, Kyoto, Japan). The cultures were grown in incubator at 37 °C in a humidified incubator with a 5% CO2.
Cell line authentication was performed using short tandem repeat (STR) profiling. Genomic DNA was analyzed across 22 STR loci, including the gender-determining Amelogenin locus and the male-specific DYS391 locus, using the GenePrint® 24 System (Promega, Madison, WI, USA). Fragment analysis was carried out on an ABI 3730XL Genetic Analyzer, and data were analyzed using GeneMapper® v5.0 software (Applied Biosystems™, Waltham, MA, USA). The STR profile matched the reference profile for the FLO-1 esophageal adenocarcinoma cell line, confirming cell line identity. All cell cultures were additionally confirmed to be mycoplasma-free using the Mycoplasma PCR Detection Kit (Cat# ab289834; Abcam, Cambridge, United Kingdom).
Antibodies
The antibodies used were EpCAM (Cat# VU-1D9; Thermo Fisher Scientific, Waltham, MA, USA) at 1:1,000 and anti-GFP (Cat# sc-9996; Santa Cruz Biotechnology (B-2), Dallas, Texas, USA). Secondary antibodies were horseradishperoxidase (HRP) conjugated anti-Rabbit IgG (Cat# P0217; Dako, Glostrup, Denmark) at 1:1,000 and HRP-conjugated anti-Mouse IgG (Cat# P0260; Dako, Glostrup, Denmark) at 1:1,000.
Generation of stable cell lines
The FLO-1 cells were seeded in 6-well plate at a density of 5 × 105 cells/well and cultured to a confluency of 80–90%. Cells were transfected with EpCAM-GFP or with GFP (pEGFP-N2) using Attractene (Qiagen GmbH, Hilden, Germany) as described by the manufacturer. After 48 h of transfection, the selection antibiotic, G418 (Geneticin) (Merck Millipore) were added to the cells at concentration of 1.5 mg/ml. For the next 2 weeks, the G418-containing medium were replaced every 3 to 4 days (or as needed). During the second week, the cells were monitored for distinct islands of surviving cells. The healthy colonies were isolated using pipette tips and transferred into wells of 96-well plates and continue to maintain cultures in medium containing the same concentration of antibiotic. The cells were then expanded into larger culture vessels before harvested for subsequent cell-based assays.
Western blotting
Cells were lysed in urea buffer (7 m urea, 0.1 m DTT, 0.1% Triton X-100, 25 mm NaCl, 20 mm HEPES–KOH pH 7.6, 5 mm NaF, 2 mm NA2VO4, 2.5 mm Na4P2O7). Proteins were quantified using the Pierce BCA Protein Assay Kits (Thermo Fisher Scientific, Waltham, MA, USA). Proteins were separated using 10–15% SDS-PAGE and then transferred onto a nitrocellulose membrane (GE Healthcare). The membranes were probed with primary antibodies, followed by secondary antibodies conjugated to HRP. Bound antibody was detected by enhanced chemiluminescence (ECL). and visualized using the ChemiDoc Go Imaging System (Bio-Rad., Hercules, CA, US).
Reverse transcriptase-quantitative PCR (RT-qPCR)
Total RNA was isolated from cells using the Monarch Total RNA Miniprep Kit (New England Biolabs, Ipswich, MA, USA) following the manufacturer’s protocol. RNA quantity and quality were assessed prior to downstream analysis. For cDNA synthesis, 1 µg of purified RNA was reverse transcribed using the Luna® Universal One-Step RT-qPCR Kit (New England Biolabs, Ipswich, MA, USA). Quantitative PCR analysis was performed using fluorogenic probe-based assays. Gene-specific primers and 5′-carboxyfluorescein (FAM)–labeled probes were obtained from PrimeTime qPCR Assays (Integrated DNA Technologies, Coralville, Iowa, USA), targeting COL1A1 (Hs00164004_m1), PXDN (Hs01034602_m1), NCAM1 (Hs00941830_m1), and the reference gene ACTB (Hs.PT.39a.22214847). Amplification reactions were carried out using Luna Universal Probe qPCR Master Mix (New England Biolabs, Ipswich, MA, USA) in combination with pre-designed TaqMan gene expression probes (Thermo Fisher Scientific, Waltham, MA, USA) on a CFX96 Touch Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA). Thermal cycling conditions and reaction setup were conducted in accordance with the manufacturers’ guidelines. Relative transcript abundance was determined by normalizing cycle threshold (Ct) values of target genes to ACTB, and differential gene expression between experimental groups was calculated using the comparative 2−ΔΔCt method.
Wound healing assay
FLO-1 stable cells (EpCAM-GFP and GFP) were seeded at 5 × 105 cells/ml in 6-well plates and culture until confluent. Cells were scratched using a (yellow) pipette tip under an angle of around 30 degrees to keep the scratch width limited by making a straight scratch to simulate the wound. After the scratched, the dish was viewed under the inverted microscope (Nikon, Tokyo, Japan) and the image was captured. The observation process was started by taking images several times throughout the following hours (0, 24, 48 and 72 h). The percentage of wound healing (%) was measured using the TScratch software (Gebäck et al., 2009).
Adhesion assay
Cell adhesion was evaluated to determine the effect of EpCAM overexpression on the attachment capacity of FLO-1 cells. FLO-1 EGFP and FLO-1 EpCAM-EGFP stable cells were grown to confluence and detached using 1× Trypsin–EDTA (0.5%) (Nacalai Tesque) pre-warmed to 37 °C. Trypsinization was neutralized by adding DMEM containing 10% FBS at a volume four times that of the trypsin. The resulting cell suspension was centrifuged, resuspended, and seeded into new 6-well plates at a 1:5 dilution using complete DMEM. Cells were allowed to adhere for 2, 4, and 6 h in separate wells. At each time point, non-adherent cells were gently removed by washing with PBS, and adherent cells were fixed and stained with 0.5% crystal violet. Plates were then air-dried and imaged using the ChemiDoc XRS+ Imaging System (Bio-Rad, Hercules, CA, USA). For quantification, the crystal violet bound to adherent cells was solubilized with 20% acetic acid, transferred into a 96-well microplate, and absorbance was measured at 600 nm using the Varioskan LUX Multimode Microplate Reader (Thermo Fisher Scientific, Waltham, MA, USA). Adhesion capacity was expressed as the relative percentage of adherent cells in EpCAM-EGFP–expressing FLO-1 cells compared to EGFP controls.
Invasion assay
Cell invasion was assessed using the QCM ECMatrix Cell Invasion Assay (ECM551; Merck Millipore, Burlington, MA, USA), following the manufacturer’s protocol with minor modifications. FLO-1 EGFP and FLO-1 EpCAM-EGFP cells were first serum-starved for 18 h in serum-free DMEM to minimize proliferation-driven movement. Following starvation, cells were washed twice with 1× PBS, after which PBS was removed completely. Cells werethen detached using 1× Trypsin-EDTA (0.5%) (Nacalai Tesque, Kyoto, Japan) pre-warmed to 37 °C, and returned briefly to the incubator until detachment was achieved. The trypsinized cells were neutralized with serum-free DMEM supplemented with 5% BSA containing Mg2+ and Ca2+ (quenching medium), and centrifuged at 2,000 rpm for 5 min. The resulting pellet was resuspended and counted to obtain a final working concentration of 6.0 × 105 cells/mL. ECMatrix-coated Boyden chamber inserts were equilibrated according to kit instructions before seeding. A total of 300 μL of the prepared cell suspension was added to the upper chamber, while the lower chamber was filled with DMEM containing 10% FBS to act as a chemoattractant. Cells were allowed to invade for the recommended incubation period. Following incubation, non-invading cells on the upper surface of the insert were carefully removed using a cotton swab. Invaded cells adhering to the underside of the membrane were stained with the Cell Stain Solution provided in the kit, washed with sterile water, and visualized using phase-contrast microscopy at 100× magnification. Representative fields were imaged for qualitative assessment. For quantitative analysis, the stained invaded cells were solubilized using the extraction buffer included in the kit, transferred to a 96-well microplate, and absorbance was measured at 560 nm using a multimode microplate reader. Invasion ability was expressed as relative absorbance values compared between EpCAM-EGFP and EGFP control FLO-1 cells.
Clonogenic analysis
The clonogenic potential of FLO-1 GFP and FLO-1 EpCAM-GFP stable cells was assessed using a standard colony formation assay. Cells were seeded at a density of 1,000 cells/well in 6-well plates containing complete DMEM and cultured for approximately 10 days, with medium renewal every 3 days. Colony formation was monitored until control wells produced colonies of adequate size, with ≥50 cells per colony considered the minimum threshold for scoring. At the end of the incubation period, colonies were washed twice with PBS and fixed using an acetic acid:methanol (1:7, v/v) solution for 10 min at room temperature. After fixation, wells were stained with 0.5% crystal violet for 2 h, washed gently with water, and air-dried. Colony images were subsequently captured, and quantification was performed across three biological replicates for each condition.
RNA-sequencing
The samples were prepared in biological replicates according to the MGIEasy RNA Directional library preparation guide with the Dynabeads® mRNA purification kit for mRNA enrichment. The libraries that passed QC were pooled based on library concentration and target data volume for sequencing on the MGI platform, DNBSEQ-G400 set at 30 million total raw reads. The mRNA sample enrichment was performed with Dynabeads® mRNA Purification Kit before being subjected to RNA fragmentation. The fragmented mRNA was reverse transcribed to cDNA followed by a second strand synthesis, end-repair and A-tailing, adapter ligation, and PCR amplification. The amplified library then went through a denaturation step, circularization, and digestion for subsequent sequencing on the DNBSEQ-G400 sequencer. The statistical power of this experimental designed was performed using the RNASeqPower model (Hart et al., 2013), assuming a biological coefficient of variation (BCV) of 0.2, a significance level of 0.05, and a minimum detectable log2 fold change of 1. Based on this, the estimated power for detecting differential expression with two replicates per group was 0.45.
Mapping and identification of differentially expressed genes
To analyze the expression of genes, RNA-seq raw reads were subjected to Trim Galore v0.6.7 for quality control and trimming. The paired-end reads were trimmed using Cutadapt v3.5.0, followed by quality inspection of clean reads using FastQC v0.11.8. Low-quality reads below the Phred score of 20 were discarded from the original data, and MultiQC v1.14 was used to compile a single report of cleaned reads. In this study, STAR aligner v2.7.10a was employed to align the clean reads to the human reference genome (GRCh38.p13), which was obtained from the Gencode human database (https://www.gencodegenes.org/human/). The chimeric reads were detected by STAR with the parameters ‘–chimSegmentMin and –chimOutType Within BAM’, and were discarded to ensure the reliability of the sequencing data by removing fragments of RNA sequences that may arise from errors during the sequencing process. The aligned RNA-seq reads were subjected to featureCounts v2.0 to quantify the gene expression levels. Differential gene expression analysis was performed using EdgeR with the parameters; mean read counts >10, p-adjusted value <0.05 and log2 FC > 1 and <−1.
Gene ontology and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis
Enrichment analyses of Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were conducted using ClusterProfiler to investigate the biological functions associated with the DEGs (reference). GO enrichment analysis was categorized into three ontology domains: biological process (BP), molecular function (MF), and cellular component (CC). The parameters used for GO and pathway enrichment were as follows: minimum number of DEGs (minGSSize) = 3, maximum number of DEGs (maxGSSize) = 800, showCategory = 10, and pvalueCutoff = 0.05. For GO enrichment analysis, the DEGs were enriched using the ‘org.HS.eg.db’ Bioconductor package, whereas the ‘kegg_organism’ parameter was set to ‘hsa’ to specifically enrich the DEGs to KEGG human pathways. REVIGO, a software tool designed to analyze and visualize enriched Gene Ontology (GO) terms (Supek et al., 2011) was employed. REVIGO facilitates the exploration of large GO term sets by reducing redundancy and identifying the most significant terms. This approach, utilizing the TreeMap function within REVIGO, enabled the generation of a concise visual summary of the enriched GO terms.
Protein-protein interaction network analysis and hubs detection
The PPI network was constructed to evaluate the relationship among the DEGs after EpCAM overexpression. The PPI bulk data was downloaded from HIPPIE v2.3 (http://cbdm-01.zdv.uni-mainz.de/~mschaefer/hippie/). HIPPIE is a comprehensive database that integrates protein interactions from various sources, and the reliability of the interactions is supported by confidence scores ranging from 0 to 1, where the cut-off range for the most reliable interactions is 0.73 to 1.0. The PPI bulk data comprising 19,679 proteins and 831,933 interactions (accessed on 29th March 2023) were visualized using Cytoscape v3.9.1. To construct the EpCAM-GFP overexpressed cells PPI network, the DEGs with the log2FC > 1 and <−1 were mapped onto the existing network map, and all the individual nodes that did not interact with any DEGs were dismissed. To gain deeper insights into the impact of EpCAM overexpression and its DEG-encoded neighbouring proteins, we specifically focused on the upregulated PPIs. The cytoHubba was employed to predict the important DEGs by predicting the top 10 highly connected genes (also called hub genes) in EpCAM-overexpressed cells’ PPI using the Maximal Clique Centrality (MCC) method. To complement the analysis of enriched GO terms, the visualization of hub genes was undertaken using a chord diagram generated by SRPlot (Tang et al., 2023). Chord diagrams excel at depicting the interrelatedness between genes and biological concepts. In this instance, SRPlot was employed to construct a visual representation of the hub genes and their connections. This visualization strategy facilitated the identification of co-occurring genes across different functional groups.
Statistical analysis
Statistical analysis was performed using GraphPad Prism v7 (GraphPad Software, La Jolla, CA, USA). The quantitative data were presented as the mean ± standard deviations (SD). Differences were considered statistically significant at values of p < 0.05.
Results
EpCAM expression and function in esophageal adenocarcinoma (ESCA)
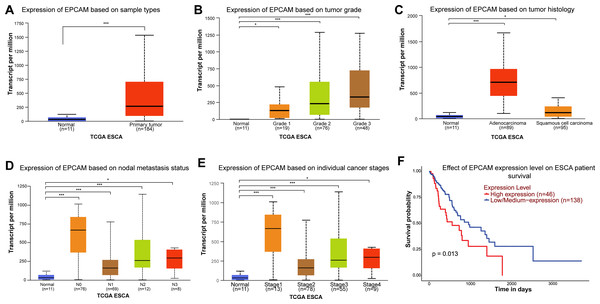
Previous high-throughput OMICS studies have identified EpCAM as one of the highly expressed molecules in ESCA (Went et al., 2004; O’Neill et al., 2017). Here, we analyzed publicly available RNA-Seq data (TCGA and GTEX) and confirmed that EpCAM was significantly upregulated in ESCA (Fig. 1A). Its overexpression also increases with higher grade of ESCA and adenocarcinoma subtype (Figs. 1B, 1C). Expression of EpCAM was highest in N0 nodal metastasis and stage 1 ESCA (Fig. 1D). EpCAM expression was found to be significantly increased in patients at stage 1 ESCA (Fig. 1E). Importantly, patients with high EpCAM expression were associated with poorer overall survival (Fig. 1F).
Figure 1: Expression of EpCAM in ESCA and its impact on patient survival.
(A) Expression of EpCAM in ESCA based on publicly available transcriptomics datasets UALCAN. The box plot compares the expression levels of EpCAM between normal tissues (n = 11) and primary tumour tissues (n = 184) in ESCA samples from The Cancer Genome Atlas (TCGA). The expression is significantly higher in primary tumors (***p < 0.001). (B) Expression of EpCAM in ESCA based on tumor grade. This box plot shows EpCAM expression across different tumor grades in ESCA. Expression levels increase from grade 1 (n = 9) to grade 3 (n = 84), with significant differences compared to normal tissues (n = 11). (*p < 0.05; ***p < 0.001) (C) Expression in EpCAM in ESCA based on tumor histology. The box plot illustrates EpCAM expression in ESCA based on tumor histology. Both adenocarcinoma (n = 89) and squamous cell carcinoma (n = 95) show higher EpCAM expression compared to normal tissues (n = 11), with adenocarcinoma exhibiting the highest levels (*p < 0.05); ***p < 0.001. (D) Expression of EpCAM in ESCA based on nodal metastasis status. This plot presents EpCAM expression across different nodal metastasis statuses in ESCA. Samples with N2 (n = 26) and N3 (n = 8) status show higher expression levels compared to normal tissues (n = 11) and N0 (n = 70) (*p < 0.05; ***p < 0.001). (E) Expression of EpCAM in ESCA based on individual cancer stages. EpCAM expression is shown for different cancer stages. Stage 4 (n = 9) samples have higher expression levels compared to earlier stages and normal tissues (n = 11) (*p < 0.05; ***p < 0.001). (F) Effect of EpCAM expression level on ESCA patient survival. Kaplan-Meier survival curves show the impact of EpCAM expression on overall survival in ESCA patients. High EpCAM expression (n = 46) is associated with poorer survival compared to low/medium expression (n = 138), with a significant p-value of 0.013.To investigate the role of EpCAM in ESCA, we previously screened three ESCA cell lines using western blot analysis and identified that OE19 and OE33 cell lines expressed endogenous EpCAM, whereas FLO-1 cells lack detectable EpCAM expression (Mohtar et al., 2018). Earlier studies have demonstrated that following subcutaneous inoculation in mice, FLO-1 cells develop spontaneous metastases with dissemination patterns closely resembling those observed in ESCA patients (Liu et al., 2016). Given that FLO-1 is the only well-characterized ESCA cell line that naturally lacks EpCAM expression, it provides a unique and unconfounded system to assess the direct functional consequences of EpCAM reintroduction without interference from endogenous protein. This feature makes FLO-1 particularly suitable as a gain-of-function model to interrogate EpCAM-mediated phenotypes. Accordingly, we employed aprotein overexpression strategy and generated stable FLO-1 cells overexpressing EpCAM-GFP or GFP alone as a control (Fig. 2A). Cells were transfected with vectors encoding EpCAM fused in-frame to a C-terminal GFP tag. A single representative clone expressing EpCAM-GFP and a matched GFP control clone were selected for downstream analyses. Stable expression of EpCAM-GFP and GFP was confirmed using fluorescence microscopy (Figs. 2B, 2C). EpCAM-GFP fluorescence was predominantly localized at the plasma membrane, with a subset of cells displaying punctate cytosolic staining (Fig. 2C). Importantly, transfection of EpCAM-GFP into ESCA cells that endogenously express EpCAM (MCF-7) resulted in clear plasma membrane localization (Fig. S1), indicating that the GFP fusion does not disrupt EpCAM trafficking. The partial cytoplasmic localization observed in FLO-1 cells likely reflects cell line–specific differences in protein folding or membrane trafficking pathways in the absence of endogenous EpCAM. To further validate expression of the EpCAM-GFP fusion protein at the protein level, western blot analysis was performed using a monoclonal anti-GFP antibody. Immunoblotting detected a higher molecular weight band corresponding to EpCAM-GFP in EpCAM-GFP–expressing FLO-1 cells, while GFP-only control cells showed the expected lower molecular weight GFP band (Fig. 2D). Detection using Anti-EpCAM antibody detected the presence of EpCAM-GFP only but not GFP-only control cells (Fig. 2E). These data confirm successful and stable expression of the EpCAM-GFP fusion construct and validate this EpCAM-null model for subsequent functional and transcriptomic investigations.
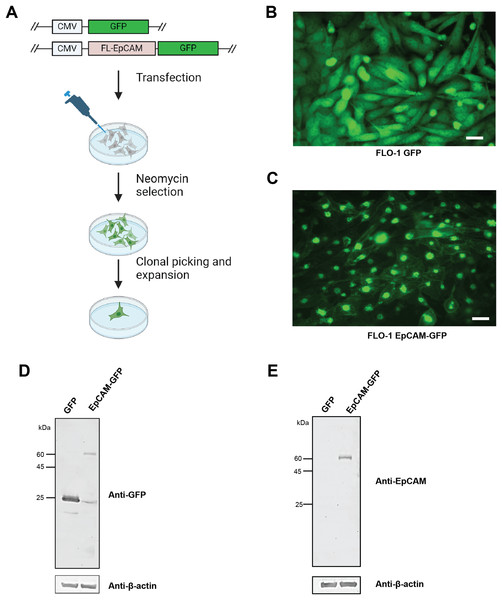
Figure 2: Generation and validation of FLO-1 cell lines expressing GFP and EpCAM-GFP.
(A) Schematic overview of the transfection and selection strategy used to generate stable FLO-1 cell lines. FLO-1 cells were transfected with either a GFP control vector or a full-length EpCAM–GFP (FL-EpCAM-GFP) expression vector, both driven by the CMV promoter. Transfected cells were subjected to neomycin selection, followed by clonal isolation and expansion to obtain stable GFP- or EpCAM-GFP–expressing cell lines. (B) Representative fluorescence microscopy image of FLO-1 cells stably expressing GFP, showing diffuse cytoplasmic GFP fluorescence. (C) Representative fluorescence microscopy image of FLO-1 cells stably expressing EpCAM-GFP, demonstrating expression of the EpCAM–GFP fusion protein with partial localization at the cell surface and punctate intracellular distribution. Scale bar = 50 μm. (D) Western blot analysis confirming expression of GFP and EpCAM–GFP proteins in stable FLO-1 cell lines using a monoclonal anti-GFP antibody. The higher molecular weight band corresponds to the EpCAM–GFP fusion protein. β-actin was used as a loading control. (E) Western blot analysis of EpCAM expression in stable FLO-1 GFP and EpCAM-GFP cell lines using a monoclonal anti-EpCAM antibody. β-actin was used as a loading control.EpCAM overexpression enhances adhesive, clonogenic, and invasive properties in ESCA cells
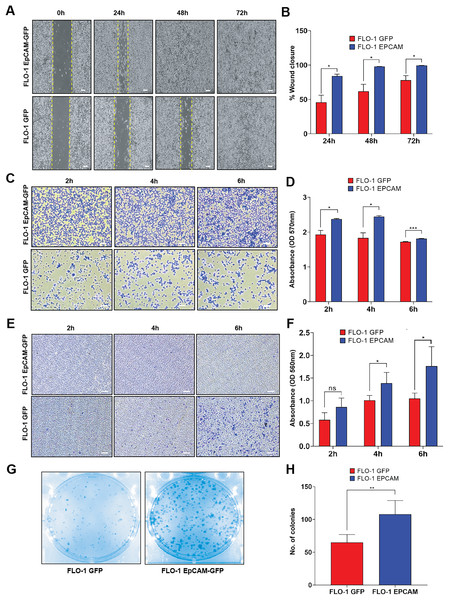
We investigated whether overexpression of EpCAM could alter cancer cell fitness in ESCA. A wound healing assay was performed to evaluate the migration properties of EpCAM-GFP-expressing FLO-1 cells compared to GFP-expressing cells. Our results demonstrated a significant increase in migration in EpCAM-GFP cells in all timepoints (24–72 h), indicating a role for EpCAM in promoting cell growth (Figs. 3A, 3B). To further elucidate the effects of EpCAM on cell behavior, we examined its impact on cell adhesion. Cells were trypsinized and allowed to bind to a plastic substratum up to 6 h timepoint. Subsequently, we stained the attached cells with crystal violet to quantify the number of cells adhering to the plastic surface. EpCAM-GFP stable cells exhibited enhanced adhesion properties and attached more rapidly to the substratum in all timepoints, suggesting that EpCAM expression contributes to increased cell stickiness (Figs. 3C, 3D). To directly assess whether EpCAM overexpression also influences invasive behavior, we next performed a transwell invasion assay. EpCAM-GFP–expressing FLO-1 cells showed a significantly higher number of invaded cells at both 4 and 6 h timepoints compared to GFP control cells, demonstrating that EpCAM overexpression enhances invasion capacity in this ESCA model (Figs. 3E, 3F). Finally, we assessed the effect of EpCAM overexpression on cell survival using a clonogenic assay. EpCAM-GFP–expressing FLO-1 cells exhibited a significantly increased colony-forming ability compared to GFP control cells, indicating that EpCAM promotes cell survival and clonogenic growth (Figs. 3G, 3H).
Figure 3: Functional cell-based assays of FLO-1 cell lines expressing GFP and EpCAM-GFP.
(A) Representative phase-contrast images of the wound-healing assay performed in FLO-1 GFP and FLO-1 EpCAM-GFP cells at 0, 24, 48, and 72 h post-scratch. Yellow dashed lines indicate the wound edges. Scale bar = 100 µm. (B) Quantification of wound closure expressed as percentage of closure over time. FLO-1 EpCAM-GFP cells exhibit significantly increased wound closure at 24, 48, and 72 h compared to GFP control cells. (C) Representative images of the adhesion assay showing attached FLO-1 GFP and FLO-1 EpCAM-GFP cells at 2, 4, and 6 h after replating. Scale bar = 100 µm. (D) Quantification of cell adhesion based on crystal violet staining, measured by optical density (OD) at 570 nm. Significantly enhanced adhesion was observed in EpCAM-GFP cells at 4 and 6 h. (E) Representative images of invaded cells from the transwell invasion assay (QCM ECMatrix) showing FLO-1 GFP and FLO-1 EpCAM-GFP cells following the invasion period. Scale bar = 100 µm. (F) Quantification of cell invasion based on extraction of stained invaded cells and measurement of optical density (OD) at 560 nm, demonstrating a significant increase in invasion in EpCAM-GFP cells compared to GFP controls. (G) Representative images of the clonogenic assay showing colony formation in FLO-1 GFP and FLO-1 EpCAM-GFP cells after ~10 days of growth. (H) Quantification of the number of colonies formed. All data represent mean ± SD from three independent biological replicates. Statistical significance was determined as indicated: *p < 0.05, **p < 0.01, ***p < 0.001, ns, not significant).Gene expression profiles in EpCAM overexpressed ESCA cells
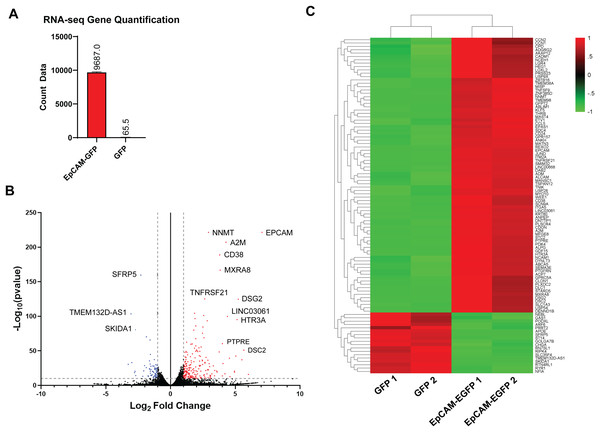
To identify potential EpCAM signalling constituents in ESCA, high-throughput RNA-seq analysis approach was used to profile the transcriptome of EpCAM-GFP and GFP ESCA cells. RNA-seq analysis were performed in biological duplicates and generated approximately between 33.92 to 41.96 million raw sequencing reads in each sample, with Phred quality score (Q30 level) surpassing 92%. Gene counts from EpCAM-GFP cells showed a significant upregulation of EpCAM transcripts compared to GFP control cells further confirming our EpCAM overexpression cell model (Fig. 4A). Volcano plot further revealed that EpCAM as one of the highest expressed gene compare to GFP control cell (Fig. 4B). We identified 797 differentially expressed genes (DEGs) in EpCAM-GFP, compared to GFP cells (p-value < 0.05 and log2 fold change >1 and <−1) (Fig. 4B). The differential expression analysis comparing EpCAM-GFP overexpressing cells to GFP-only expressing cells revealed significant changes in gene expression. The base mean expression value across all genes ranged from 14.45 to 105,189.97, with a mean of 794.76. The log2 fold change (FC) values, indicating the magnitude of differential expression, ranged from −4.60 to 9.43 with a mean of 1.03, suggesting that on average, the genes were upregulated in EpCAM-GFP overexpressing cells compared to GFP-only cells. Of this, 571 DEGs were upregulated whereas 226 were downregulated (Table S1). Hierarchical clustering analysis highlighted that the top 100 significant genes were enriched with upregulated genes compared to the downregulated genes (Fig. 4C). This significant gene list includes both upregulated and downregulated genes. Among the top upregulated genes, EpCAM (log2 FC: 7.06), NNMT (log2 FC: 2.93), and A2M (log2 FC: 4.28) were notably elevated in EpCAM overexpression cells. Other significantly upregulated genes include GDF15 (log2 FC: 3.68), SERPINE1 (log2 FC: 3.45), and S100A4 (log2 FC: 2.78). Conversely, substantial downregulation was observed in genes such as MXRA8 (log2 FC: −3.84), CD38 (log2 FC: −3.79), and TNFRSF12A (log2 FC: −2.65). Additional significantly downregulated genes include ADAM12 (log2 FC: −2.50), KRTAP4-12 (log2 FC: −2.45), and BHLHE40 (log2 FC: −2.30). These findings indicate significant transcriptional changes due to EpCAM overexpression, affecting various pathways and biological processes.
Figure 4: RNA-seq analysis of EpCAM-GFP overexpression FLO-1 cells.
(A) RNA-seq Gene Quantification. The bar plot represents the quantification of EpCAM gene expression in EpCAM-GFP and GFP samples. The count data shows a significant upregulation of EpCAM in EpCAM-GFP ESCA cells compared to GFP cells. (B) Volcano plot of treated (EpCAM-GFP) vs. control (GFP). The volcano plot displays the log2 fold change (FC) against the −log10 p-value for gene expression levels in treated (EpCAM-GFP) vs. control (GFP) cells. Points in red indicate significantly upregulated genes with a p-adjusted value <0.05 and log2 fold change (log2FC) > 1. Points in blue indicate significantly downregulated genes with a p-adjusted value <0.05 and log2FC < −1. Key genes, such as EpCAM, NNMT, A2M (upregulated), and SFRP5, SKIDA1 (downregulated), are highlighted. The dashed lines mark the thresholds for significance. (C) Heatmap of differential gene expression. The heatmap illustrates the hierarchical clustering of gene expression profiles between EpCAM-GFP and GFP samples. Each row represents a gene, and each column represents a sample. Red indicates upregulation, and green indicates downregulation. Key differentially expressed genes such as EpCAM and others are prominently displayed, demonstrating distinct expression patterns between the two conditions.Functional enrichment and validation of EpCAM-associated gene networks in ESCA
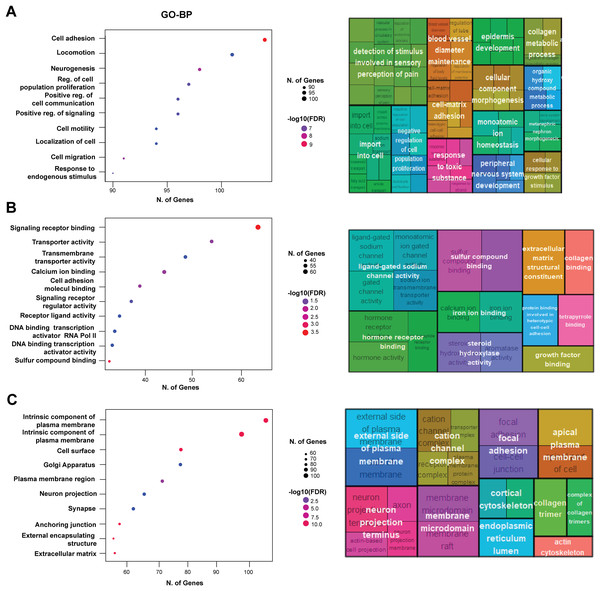
Next, we performed functional enrichment of the DEGs using GO annotations. The GO-biological process (GO-BP) analysis revealed significant enrichment of genes associated with cell adhesion, locomotion, and neurogenesis, indicating that EpCAM overexpression substantially impacts transcriptional programs governing cellular movement and interaction (Fig. 5A). Additional enriched biological processes included cell population proliferation, cell communication, collagen organization, angiogenesis, and signaling regulation, suggesting broad effects of EpCAM on cancer-associated cellular functions. GO molecular function (GO-MF) analysis showed significant enrichment of signaling receptor binding, transporter activity, and transmembrane transporter activity (Fig. 5B). Other notable functional categories included cell adhesion molecule binding, receptor ligand activity, hormone receptor binding, collagen binding, growth factor binding, and ligand-gated ion channel activity, further supporting the involvement of EpCAM in modulating signaling and adhesion-related pathways. Analysis of cellular component (GO-CC) annotations demonstrated enrichment of genes localized to the plasma membrane, focal adhesions, and neuron projection termini (Fig. 5C), consistent with the observed phenotypic changes in EpCAM-overexpressing cells. To further identify key molecular players downstream of EpCAM, we constructed a protein–protein interaction (PPI) network using the upregulated gene set and ranked nodes based on connectivity to identify highly connected hub genes. We reasoned that genes functionally intersecting with EpCAM signaling would be preferentially upregulated following EpCAM overexpression. Consistent with this hypothesis, EpCAM showed strong connectivity within the upregulated PPI network, whereas limited interaction was observed among downregulated genes. The top 10 upregulated hub genes identified were SOX2, COL1A1, LOX, COL3A1, LUM, PXDN, BDNF, NCAM1, TLR2, and CCL5 (Fig. 6A). Within this network, PXDN and SOX2 occupied central positions, exhibiting multiple interactions with other nodes and indicating their prominence in the predicted EpCAM-associated protein network. In addition to these central hubs, several peripheral yet highly connected genes including COL1A1, COL3A1, LOX, LUM, BDNF, NCAM1, CCL5, and TLR2. Functional annotation of these hub genes revealed enrichment in processes associated with cell adhesion, migration, extracellular matrix organization, and cancer-related signaling (Fig. 6B). KEGG pathway enrichment further linked these genes to major oncogenic pathways, including PI3K–Akt signaling, Hippo signaling, focal adhesion, and cell adhesion molecule pathways (Fig. 6C).
Figure 5: Functional enrichment analysis of differentially expressed genes.
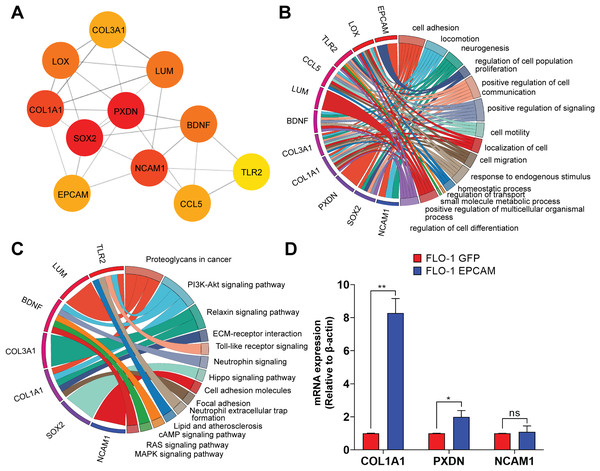
(A) GO Biological Process (GO-BP). The left panel shows scatter plot displaying the number of genes (N. of Genes) involved in significant biological processes, with the size of the dots representing the number of genes and the color indicating the −log10 of the False Discovery Rate (FDR). Key processes include cell adhesion, locomotion, neurogenesis, and regulation of cell population proliferation. The right panel shows treemap generated using REVIGO, providing a visual summary of the enriched GO-BP terms, highlighting the most prominent processes in larger and more centrally located blocks, with similar terms clustered together to reduce redundancy. (B) GO Molecular Function (GO-MF). The left panel shows scatter plot displaying the number of genes (N. of Genes) involved in significant molecular function which include signaling receptor binding, transporter activity, transmembrane transporter activity, and calcium ion binding. The right panel shows treemap illustrating the enriched GO-MF terms. (C) GO Cellular Component (GO-CC). Notable components include the intrinsic component of the plasma membrane, cell surface, Golgi apparatus, and synapse. The right panel shows treemap visualizing the enriched GO-CC terms.Figure 6: Protein-protein interaction network highlighting the top 10 upregulated hub-genes and their functional enrichment.
(A) Network of top 10 upregulated hub-genes. The network diagram displays the interactions between the top 10 upregulated hub-genes identified in the dataset, along with the EpCAM gene, which was added to observe its interaction with the hub-genes. The color intensity of the nodes indicates the number of interactions, with darker colors representing more interactions. The hub-genes included are SOX2, COL1A1, LOX, COL3A1, LUM, PXDN, BDNF, NCAM1, CCL5, and TLR2. Edges represent the predicted interactions between these the genes and EpCAM. (B) Hub-genes enrichment analysis GO-BP. The circular chord diagram illustrates the enrichment of the top 10 upregulated hub-genes in various biological processes. The width of the ribbons represents the number of genes involved in each process. Key processes include cell adhesion, locomotion, neurogenesis, regulation of cell population proliferation, and positive regulation of cell communication. The color gradient indicates the log2FC, with blue representing lower and red representing higher fold change. (C) KEGG pathway enrichment analysis. The circular chord diagram shows the enrichment of the top 10 upregulated hub-genes in KEGG pathways. Significant pathways include Proteoglycans in cancer, PI3K-Akt signaling pathway, ECM-receptor interaction, and neurotrophins signaling pathway. The color gradient represents the log2FC, with blue indicating lower and red indicating higher fold changes.(D) Quantitative PCR (qPCR) analysis of COL1A1, PXDN, and NCAM1 expression in FLO-1 EpCAM-GFP cells relative to FLO-1 GFP control cells. Gene expression levels were normalized to the housekeeping gene ACTB, and relative expression was calculated using the 2−ΔΔCt method. Data are presented as mean ± SD from three independent biological replicates. Statistical significance was determined as indicated (*p < 0.05, **p < 0.01, ns, not significant).To experimentally validate selected candidates from the EpCAM-associated PPI network, we prioritized COL1A1, PXDN, and NCAM1 based on their network connectivity, centrality within the PPI network, and functional relevance to cell adhesion and extracellular matrix regulation. Quantitative PCR (qPCR) analysis confirmed that COL1A1 and PXDN were significantly upregulated in EpCAM-GFP–expressing FLO-1 cells compared to GFP controls, consistent with the RNA-seq data (Fig. 6D). In contrast, NCAM1 exhibited a modest increase in expression that did not reach statistical significance, indicating partial concordance with the transcriptomic findings. These results validate COL1A1 and PXDN as robust downstream components of EpCAM-associated transcriptional programs, while highlighting gene-specific differences in validation outcomes.
Discussion
The upregulation of EpCAM has been widely reported in various epithelial cells-derived cancer types including esophageal cancer. Increased in EpCAM expression promotes tumor growth and aggressiveness, and is linked to poor clinical outcomes. Nonetheless, loss of EpCAM expression has been shown to mediate EMT that is the critical step in metastasis initiation (Hyun et al., 2016; Liu et al., 2021). The change into mesenchymal phenotype increases the cancer cells motility and invasiveness potentials, thus enabling the cells to intravasate into circulation and spread to the distant sites. The disseminated cancer cells will subsequently undergo mesenchymal-to-epithelial transition (MET), marked by increased in EpCAM expression, which this process facilitates tumor growth and colonization at the secondary sites. Therefore, the dynamic and temporal expression of EpCAM is crucial in driving the tumor metastasis, which this process are also orchestrated by intricate crosstalk between EpCAM and its interactome (Brown, Sankpal & Gillanders, 2021).
This present study aims to provide additional insights into EpCAM functional significance and its potential interacting partners that involve in promoting ESCA pathogenesis. To this end, we utilized the EpCAM-null metastatic FLO-1 ESCA cells, which represent an ideal model to interrogate EpCAM function via the reintroduction of exogenous EpCAM. The absence of EpCAM expression in these FLO-1 cells is consistent with the reported loss of EpCAM in mesenchymal-like metastatic or disseminated cancer cells (Lin et al., 2021). Introduction of EpCAM in this EpCAM-null FLO-1 background resulted in cell surface localization with punctate cytoplasmic staining, possibly reflecting altered EpCAM trafficking or signalling competence in these cells. Functional characterization of EpCAM-overexpressing FLO-1 cells demonstrated increased proliferation, adhesion, and clonogenic and invasive capacity, consistent with the established role of EpCAM in mediating cell-cell adhesion, metastasis and promoting tumor growth.
While the current study focused on EpCAM gain-of-function using an EpCAM-null model, complementary loss-of-function approaches will be essential to fully delineate EpCAM dependency in ESCA. In particular, targeted silencing of EpCAM using CRISPR-mediated gene knockout or RNA interference (shRNA/siRNA) in EpCAM-high ESCA cell models (e.g., OE19 or OE33) would enable direct assessment of EpCAM requirement for tumor cell fitness, invasiveness, and downstream signalling. Such strategies would also allow functional validation of the EpCAM-associated transcriptomic and PPI networks identified in this study. Accordingly, future studies integrating EpCAM loss-of-function approaches with biochemical and functional assays, such as protein interaction analyses (e.g., co-immunoprecipiation) and pathway-specific perturbation experiments, will be essential to define the causal relationships between EpCAM and its downstream effectors in ESCA.
The dependency of our ESCA in vitro model on EpCAM underscores the potential of this protein as a therapeutic target in ESCA. Several EpCAM-targeting monoclonal antibodies have been developed and tested in multiple epithelial cancers, including colorectal and breast cancer (Schmidt et al., 2010; Satofuka et al., 2023; Lee et al., 2023), highlighting the translational relevance of EpCAM-directed strategies. In addition to its therapeutic potential, EpCAM has also been widely exploited as a diagnostic marker, particularly for the detection of circulating tumor cells (CTCs). Alongside EpCAM, the identification of additional ESCA-specific surface or extracellular markers could further improve diagnostic and prognostic stratification. Recent multi-omics studies have indeed confirmed EpCAM overexpression in ESCA and identified additional enriched candidates, including GPA33, SLC25A30, TAOK2, and AGMAT (O’Neill et al., 2024).
A holistic understanding of EpCAM-associated molecular networks is critical for identifying downstream dependencies that may be exploited clinically. Reintroduction of exogenous EpCAM into EpCAM-null FLO-1 cells resulted in widespread transcriptional reprogramming, with 571 genes upregulated and 226 genes downregulated relative to controls. These changes are consistent with previously described EpCAM signaling mechanisms mediated through its intracellular (EpICD) and extracellular (EpEX) domains (Mohtar et al., 2020). Functional enrichment analyses revealed strong over-representation of pathways related to cell adhesion, migration, extracellular matrix organization, and signal transduction, providing a molecular basis for the enhanced migratory, adhesive, clonogenic, and invasive phenotypes observed upon EpCAM overexpression.
Among the upregulated genes, NNMT and SOX2 emerged as prominent candidates in the transcriptomic and protein–protein interaction (PPI) network analyses. NNMT has been implicated in metabolic and epigenetic reprogramming in several malignancies (Wang et al., 2022), and recent work has linked NNMT to metastasis-associated metabolic remodeling in esophageal squamous cell carcinoma (Huang et al., 2024). Similarly, SOX2 is a lineage-defining transcription factor essential for esophageal development and has been associated with poor prognosis and oncogenic signaling in esophageal cancer (Bass et al., 2009). While these findings suggest potential connections between EpCAM and NNMT or SOX2-driven programs, their involvement in EpCAM signaling in ESCA remains predictive, as these targets were not experimentally validated in the present study. Consequently, these associations should be interpreted as hypothesis-generating and warrant future mechanistic interrogation.
To prioritize candidates for experimental validation, we focused on genes that combined high network connectivity within the EpCAM-associated PPI network with functional relevance to cell adhesion and extracellular matrix (ECM) dynamics. Based on these criteria, COL1A1, PXDN, and NCAM1 were selected for qPCR validation. Notably, PXDN occupied a central position within the PPI network, interacting with multiple EpCAM-associated nodes, suggesting a key role in maintaining network integrity. qPCR analysis confirmed significant upregulation of COL1A1 and PXDN in EpCAM-overexpressing FLO-1 cells, whereas NCAM1 showed a modest but non-significant increase, indicating selective activation of downstream targets.
PXDN (peroxidasin) is a secreted extracellular matrix–associated peroxidase that catalyzes the formation of sulfilimine crosslinks in collagen IV networks, thereby contributing to basement membrane stabilization and tissue architecture (Cheng & Shi, 2022). Increasing evidence implicates PXDN in cancer progression, where altered basement membrane remodeling can promote tumor cell invasion, survival, and interaction with the surrounding stroma (Bhave et al., 2012; Tian et al., 2024). Importantly, PXDN has been linked to cell adhesion, focal adhesion signaling, and invasive niche formation, processes that closely align with the phenotypic effects observed upon EpCAM overexpression in our model (Zhu & Jin, 2024; Li et al., 2024). Given that EpCAM is known to regulate cell–cell adhesion and modulate cytoskeletal and signaling dynamics, the upregulation of PXDN suggests a mechanism whereby EpCAM signaling may indirectly remodel the extracellular matrix and basement membrane to facilitate invasive behavior. This interpretation is further supported by enrichment of focal adhesion and ECM–receptor interaction pathways in our transcriptomic analyses.
COL1A1 major fibrillar collagens that constitute the structural backbone of the tumor extracellular matrix and are increasingly recognized as active regulators of tumor progression rather than passive scaffolding components (Kuivaniemi & Tromp, 2019; Li et al., 2022). Overexpression of COL1A1 has been linked to enhanced tumor stiffness, focal adhesion signaling, and activation of pro-tumorigenic pathways such as FAK–PI3K–AKT signaling in multiple epithelial cancers, including gastrointestinal malignancies (Provenzano et al., 2008; Levental et al., 2009; Kai, Drain & Weaver, 2019). Importantly, ECM remodeling driven by collagen deposition has been shown to promote cancer cell invasion, survival, and therapeutic resistance. These are phenotypes that closely mirror those induced by EpCAM overexpression in our model. Mechanistically, EpCAM has been implicated in regulating cell–cell and cell–matrix interactions through modulation of adhesion complexes and cytoskeletal dynamics (Gires et al., 2020). The observed upregulation of COL1A1 therefore suggests that EpCAM signaling may indirectly remodel the extracellular matrix to reinforce adhesive and invasive cellular states, providing a plausible mechanistic link between EpCAM overexpression and the aggressive phenotypes observed in ESCA cells. This notion is further supported by enrichment of focal adhesion and ECM–receptor interaction pathways in our transcriptomic analyses.
Together, these findings position PXDN and COL1A1 as validated downstream effectors of EpCAM signaling, reinforcing the concept that EpCAM promotes ESCA progression, at least in part, through modulation of extracellular matrix structure and tumor–microenvironment interactions.
Conclusions
This study demonstrates that EpCAM promotes aggressive cancer-associated phenotypes in ESCA, including enhanced migration, adhesion, clonogenic survival, and invasion. Using an EpCAM-null ESCA model, we show that EpCAM overexpression drives transcriptional programs enriched for cell adhesion, focal adhesion signaling, and extracellular matrix organization. Importantly, targeted validation confirmed upregulation of the extracellular matrix genes COL1A1 and COL3A1, linking EpCAM signaling to collagen-mediated matrix remodeling and invasive cellular behavior. Collectively, these findings position EpCAM as a key regulator of tumor cell–extracellular matrix interactions and highlight its potential as a therapeutic and biomarker target in ESCA. Further studies incorporating EpCAM loss-of-function approaches and in vivo validation will be essential to fully define the EpCAM signaling axis and its clinical relevance.
Supplemental Information
Fluorescence microscopy images of MCF-7 cell lines transfected with EpCAM-GFP construct showing cell surface expression.
Left panel: nucleus staining with DAPI; Middle panel: EpCAM-GFP; Right panel: Merge image of DAPI and EpCA
Significant DEGs from RNA-seq quantification of EpCAM-GFP in comparison with GFP-FLO1 cells.
RNA-seq raw data.
The aligned RNA-seq reads in BAM format are quantified using featureCounts V2.0.3 (Liao, Smyth, and Shi 2014). This gene-level quantification approach utilizes a gene transfer format (GTF v36) containing gene models and count the number of reads that align to each gene (read count).