Unraveling microbial diversity and physicochemical hazard level in Thai traditional fermented shrimp paste (Kapi)
- Published
- Accepted
- Received
- Academic Editor
- Zothanpuia
- Subject Areas
- Agricultural Science, Food Science and Technology, Microbiology
- Keywords
- Fermented shrimp paste, Bacterial community, Histamine, Food safety, Food quality
- Copyright
- © 2026 Mahidsanan et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, reproduction and adaptation in any medium and for any purpose provided that it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2026. Unraveling microbial diversity and physicochemical hazard level in Thai traditional fermented shrimp paste (Kapi) PeerJ 14:e20864 https://doi.org/10.7717/peerj.20864
Abstract
Fermented shrimp paste (Kapi) is a culturally significant condiment valued for its flavor and nutritional value. However, inconsistent production practices may lead to microbial contamination and histamine accumulation, posing health risks. Understanding microbial diversity, salt-tolerant pathogens, and the effectiveness of hygienic controls is essential for improving product safety and quality consistency. This study assessed the microbial and physicochemical characteristics of seven Kapi produced under varying hygienic conditions: a certified commercial export product (M1), a traditionally fermented product using 25% (w/w) salt (M2), wet-market bulk products (M3, M4), and sealed community-enterprise products (M5, M6, M7). Microbiological hazards were characterized using Oxford Nanopore sequencing, and physicochemical analyses, including histamine content, were conducted. M1 exhibited the highest safety and quality, with no detected pathogens. In contrast, M3 and M5 contained high microbial loads, including Staphylococcus aureus, Bacillus cereus, Clostridium perfringens, and fungi. Bacterial diversity varied significantly across samples, with M3 and M4 showing the greatest richness, whereas M6 had the lowest. Dominant species identified were Lentibacillus salinarum, Lentibacillus amyloliquefaciens, and Lentibacillus kimchi in M1, M3, and M4, and Staphylococcus sciuri in M2, M5, and M6, all of which possessed histamine-degrading potential. Alkalibacterium kapii dominated M7, while histamine-producing Jeotgalicoccus halotolerans was found in M5–M7. Physicochemical variations corresponded with production practices. M6 showed the highest histamine level (39.4 mg/kg), while M1 and M4 had the lowest (12.7 and 9.5 mg/kg, respectively), indicating differences in hygiene and salt management. Overall, microbial and physicochemical hazard levels were closely associated with production environments. To enhance the safety and consistency of Kapi, we propose a risk-based framework that includes standardizing salt concentration (25% w/w) to control microbial dynamics and histamine formation, strengthening hygienic practices from Good Hygiene Practice (GHP) to Good Manufacturing Practices (GMP) compliance to ensure adequate safety control, and applying defined starter cultures to stabilize fermentation and reduce variability. These strategies collectively address how hygiene, salt concentration, and environmental control influence microbial and physicochemical hazards in traditional fermented shrimp paste.
Introduction
Fermented shrimp paste, known as Kapi in Thailand, is a traditional seafood condiment produced through the salt fermentation of small shrimp (Acetes spp.) or krill (Mesopodopsis orientalis). Similar products are found across Asia under various local names such as Terasi (Indonesia), Belacan (Malaysia), Bagoong-alamang (Philippines), and Mam ruoc (Vietnam) (Pamungkaningtyas, 2023). Although Kapi is widely consumed for its umami flavor and nutritional value (Kleekayai et al., 2015; Chotechuang, 2012), spontaneous fermentation often results in inconsistent quality and potential safety risks. Fermentation typically involves salt concentrations of 25–35% (w/w) under anaerobic conditions for several months, allowing halophilic bacteria such as Bacillus, Tetragenococcus, Alkalibacterium, Lactobacillus, Lentibacillus and Oceanobacillus to dominate (Li et al., 2021; Pongsetkul et al., 2017; Pamungkaningtyas, 2023; Che et al., 2021). However, these conditions can also permit the survival of salt-tolerant pathogenic bacteria including Staphylococcus aureus, Salmonella enterica serovar and Vibrio spp. (Lee et al., 2024; Gaffar, Jatmiko & Prihanto, 2022; Chhetri, Prakitchaiwattana & Settachaimongkon, 2019).
Inadequate hygiene practices, particularly in wet markets, which are traditional open-air markets commonly found in Southeast Asia where fresh foods, including fermented products, are sold in unpackaged or minimally processed form and have limited hygienic control (Zhou et al., 2025), contribute to contamination with Escherichia coli, coliforms, and Bacillus cereus (Nakamura et al., 2022), further increasing microbial contamination risks. During extended fermentation, microbial decarboxylation of amino acids can produce biogenic amines (BAs), especially histamine, which remains stable under normal household cooking conditions and cause allergic or toxic reactions (Vinci & Maddaloni, 2020; Shi, Kim & Moon, 2023). Histamine levels are therefore regarded as critical indicators of both microbial activity and product safety (Li et al., 2021). Recent evidence indicates that maintaining a 25% (w/w) salt concentration effectively reduces microbial loads and histamine accumulation in fermented shrimp products (Li et al., 2021). However, the influence of salt concentration on microbial diversity and hygienic management remains an important area for further investigation. In recent years, studies have shown that reducing the traditional salt concentration from 25% (w/w) to 6-18% maintains fermentation efficiency, with Tetragenococcus halophilus remaining active in inhibiting spoilage microbes. Higher salt levels, by contrast, reduce lactic acid bacteria (LAB) growth, moisture, and bioactivity, leading to lighter color and lower functionality (Kurnianto et al., 2025).
While these studies provide valuable insights under controlled fermentation conditions, further research is needed to understand how microbial population dynamics and salt concentration interact within real-world production environments. This includes systems operating under varying levels of Good Manufacturing Practices (GMP) compliance, small-scale community processing, and wet-market conditions. Similarly, Wang et al. (2023) emphasized the importance of integrating microbial community profiling with chemical-safety assessment in fermented seafood products. Building on these advances, the present study investigates Kapi produced under diverse production and hygiene conditions, ranging from commercial Good Manufacturing Practices/Hazard Analysis and Critical Control Point (GMP/HACCP) systems to unregulated markets, using Oxford Nanopore sequencing to elucidate the relationship between microbial diversity, histamine formation, and physicochemical safety. Despite the long-standing tradition and economic relevance of Kapi production, comprehensive studies linking microbial diversity, histamine risk, and hygienic practices across different production systems remain scarce. Prior research has primarily focused on either fermentation chemistry or individual bacterial isolates, without integrating food-safety frameworks such as GMP and Hazard Analysis and Critical Control Point (HACCP). Consequently, the influence of production environment and safety-management levels on microbial ecology and physicochemical safety has not been fully elucidated. To address these gaps, this study employed Oxford Nanopore Technologies (ONT), an advanced metagenomic tool capable of identifying both culturable and non-culturable microorganisms at the species level, as well as functional genes related to histidine decarboxylation and biogenic amine formation (Leggett et al., 2020; Yogeswara et al., 2024). Integrating ONT-based microbial profiling with physicochemical and histamine analyses enables a comprehensive evaluation of microbial diversity and food safety. This approach also supports risk assessment, process optimization, and regulatory compliance to ensure product quality and consumer protection (Tyler et al., 2018; Jabeen et al., 2022).
Therefore, this study was designed to evaluate Kapi products produced under varying levels of quality control and compliance with GMP and HACCP principles. Four representative production types were examined: (i) a certified commercial export product (process control under GMP/HACCP), (ii) a traditionally fermented product with 25% (w/w) salt, (iii) a wet-market bulk product (uncontrolled hygienic conditions), and (iv) a sealed community-enterprise product (limited Good Hygiene Practice (GHP) compliance). The study investigates bacterial diversity at both genus and species levels, with particular focus on histamine-producing bacteria, and further assesses histamine content, physicochemical properties, and pathogenic contamination as indicators of product safety. This approach clarifies how variations in production practices, packaging environments, and safety-management systems affect microbial diversity, histamine formation, and overall safety. The findings offer baseline data to support standardized production and enhance the microbial and physicochemical safety of fermented shrimp paste and related products.
Materials and Methods
Collection of samples
Fermented shrimp paste (Kapi) samples used in this study were obtained from various production practices in Chanthaburi Province, Thailand, representing distinct levels of quality control and hygiene management. Seven samples were categorized into four representative production types: (i) certified commercial export product manufactured under GMP/HACCP standards (M1), which served as the process control; (ii) a traditionally fermented product prepared with 25% (w/w) salt concentration (M2); (iii) bulk Kapi sold in wet-markets, representing traditional wet-market products with uncontrolled salt concentrations and minimal hygienic conditions (M3 and M4); and (iv) sealed products from community enterprises, reflecting small-scale production with a limited GHP compliance (M5, M6, and M7). All commercial Kapi samples were collected within their labeled manufacturing (MFG) date of 05 June 2024 and best-before (BBF) date of 05 June 2026 to ensure product freshness and validity prior to analysis. This sampling framework was designed to capture the diversity of real-world production environments and hygiene practices, thereby enabling comparative evaluation of microbial diversity and food safety among distinct manufacturing systems.
Sample collection was conducted according to standard procedures employed in food analytical laboratories and the food industry, following the AOAC method described by Feldsine, Abeyta & Andrews (2002). All samples were aseptically collected into clean and sterile bags, with a total weight of 10 kg to allow for testing, backup, and reference purposes. To ensure uniformity and representativeness, portions were collected from the upper, middle, and lower sections, then thoroughly mixed before analysis. All samples were sealed and labeled with the date and time of collection, name of the sample collector, and the source. Samples were then stored at −20 °C during transportation to the laboratory. A portion of each collected sample was allocated for immediate analysis.
Analysis of microbiological hazards in fermented shrimp paste
Enumeration of bacterial counts
A 25 g sample of fermented shrimp paste sample was mixed with 225 mL of 0.1% (w/v) peptone water in a stomacher bag. The mixture was homogenized using a stomacher (Karlsruhe, Germany) at a 200-rpm for 60 s to obtain a uniform initial dilution. Serial dilutions were then prepared for further microbial analysis (US-FDA, 2023).
To enumerate total viable count (TVC), a 0.1 mL aliquot from each dilution level was pipetted onto plate count agar (PCA) (HiMedia, Maharashtra, India), and evenly spread using the spread plate technique. The plates were incubated at 35 °C for 24 h. To enumerate lactic acid bacteria (LAB), 0.1 mL of each diluted sample was pipetted onto De Man Rogosa Sharpe (MRS) agar (HiMedia) plate. The plates were incubated in an anaerobic jar at 35 °C for 48 h in an inverted position. The number of colonies on agar plates were then counted and expressed as log CFU/g (US-FDA, 2023).
To count B. cereus and S. aureus, selective agar media were used to inhibit the growth of non-target microorganisms and ensure accurate enumeration. For B. cereus, Mannitol Egg Yolk Polymyxin (MYP) agar (HiMedia) containing polymyxinB was used as the selective agent. A 0.1 mL aliquot of the diluted fermented shrimp paste sample was spread onto separate plates containing MYP agar. The plates were then incubated at 35 °C for 24 h. Colonies appearing pink to red with a surrounding precipitate were counted. For S. aureus, Baird-Parker Agar (BPA) with egg yolk tellurite was used as the selective agent (HiMedia). A 0.1 mL aliquot of each diluted sample was spread onto separate plates containing BPA and incubated at 35 °C for 24 h. S. aureus appeared as black colonies with a halo, which were then counted (US-FDA, 2023).
To assess the levels of E. coli and coliforms, Petrifilm™E. coli/coliforms count plates were used. A 25 g of sample was diluted in 0.1%(w/v) peptone water, and one mL of the diluted sample was transferred onto the surface of the petrifilm plate. The sample was then evenly spread across the film to ensure uniform distribution, after which it was incubated at 35 °C for 24 h. Coliform colonies typically appeared as red colonies, while E. coli colonies were identified by their blue or blue–green color on a red background.
To detect Clostridium perfringens, a one mL aliquot of the homogenized sample was transferred into 10 mL of cooked meat medium (HiMedia) and incubated anaerobically at 35 °C for 24 h to promote bacterial growth. The cultured samples were then transferred onto perfringens agar base (HiMedia) using spread plating techniques. The plates were incubated anaerobically at 35 °C for 24 h. C. perfringens colonies were then identified by their black coloration resulting from sulfite reduction (US-FDA, 2023).
To detect Salmonella spp., Xylose Lysine Deoxycholate (XLD) Agar (HiMedia) was used as the selective medium. A 25 g of the sample was mixed with 225 mL of pre-enriched lactose broth, homogenized using a stomacher for 2 min, and incubated at 35 °C for 24 h. Subsequently, one mL of the overnight pre-enriched sample was transferred into 10 mL of selenite cystine broth and incubated at 35 °C for 24 h. The broth culture was aseptically streaked onto XLD agar and incubated at 35 °C for 24 h. Colonies that appeared red with a black center were identified as Salmonella spp. (US-FDA, 2023).
Enumeration of yeasts and molds
A 25 g of the fermented shrimp paste sample was mixed with 225 mL of 0.1%(w/v) peptone water. Serial dilutions were then performed to obtain an appropriate dilution. A 0.1 mL of the diluted sample was spread onto Potato Dextrose Agar (Himedia) supplemented with 10% (w/v) tartaric acid to inhibit bacterial growth, and incubated at 25 °C for 3–5 days. The total count of yeasts and molds were counted and expressed as log CFU/g (US-FDA, 2023).
Microbial community assessment using Oxford nanopore technologies
Metagenomic DNA extraction
Prokaryotic metagenomic DNA was isolated using QIAamp PowerFecal Pro DNA Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Briefly, 0.25 g of pooled samples, taken from ten bags per treatment, was used for extraction. The quality and concentration of the extracted DNA was then analyzed using DeNovix QFX Fluorometer.
16S rRNA library sequencing
The prokaryotic 16S rRNA gene was amplified using a Native barcoding kit, targeting the full length 16S rRNA. PCR amplification was carried out under the following cycling conditions: initial denaturation at 95 °C for 1 min; followed by 40 cycles of 95 °C for 20 s, 55 °C for 30 s, and 65 °C for 2 min; with a final extension at 65 °C for 5 min. The resulting 16S rRNA amplicons were purified using AMPure XP magnetic beads and ligated with distinct Native barcodes for sequencing. DNA libraries with different barcodes (approximately 1,500 bp) were further purified using AMPure XP beads. The quality and quantity of DNA libraries were evaluated using DeNovix QFX Fluorometer and QIAxcel Advanced (Qiagen, Hilden, Germany), respectively. Sequencing was performed using Oxford nonopore MinION Mk1C platform following the manufacturer’s protocol (Oxford Nanopore Technologies).
Bioinformatics analysis
Basecalling and adapter trimming of raw sequences were performed using Dorado program version 0.7.3. Sequencing reads were filtered using DADA2 pipeline (minlength = 1,000, maxlength = 1,600). High-quality reads were then processed using Emu algorithm with the Emu database (5) to establish microbial community profiles. Alpha diversity indices (Chao1 richness and Shannon) and beta diversity were computed using R software.
The nucleic acid sequences generated in this study have been deposited in an open access Sequence Read Archive database of NCBI under accession number PRJNA1221651.
Analysis of physiochemical factors in fermented shrimp paste
pH, salinity, and titratable acidity measurement
A 10 g of each sample was thoroughly homogenized with 90 mL of distilled water (w/v) prior to determining pH, salt concentration, and titratable acidity. The pH value was measured using a pH meter (Mettler Toledo, Zurich, Switzerland). Salt concentration was measured using refractometer meter (Mettler Toledo Easy R40 Refractometer). Titratable acidity was measured following the method described by Tomovska, Gjorgievski & Makarijoski (2016). Briefly, each sample solution was titrated with 0.1 M NaOH until a stable, slightly pink color was reached. The results were expressed as a percentage of lactic acid.
Measurement of total nitrogen and protein content
The total nitrogen content of all fermented shrimp paste samples was determined using the classical Kjeldahl method (Kjeldahl, 1883). Protein content was subsequently estimated based on the measured nitrogen values.
Histamine analysis
The concentration of histamine in Kapi samples was analyzed using high performance liquid chromatography (HPLC) according to the method described by Li et al. (2021), with slight modifications. The analysis was performed using a Symmetry C18 (3.9 mm × 150 mm, 5 µm). The mobile phase was composed of (A) acetonitrile and (B) 10 mM KH2PO4 (30:70%). The run time was 5 min, with a constant flow rate of 0.6 mL/min. The column temperature was set at 25 °C, with the injection volume set to 20 μL. A fluorescence detector (with an excitation 350 nm and an emission wavelength of 444 nm) was used.
Statistical analysis
All experiments were performed using three independent biological replicates, with each measurement within a replicate performed in technical triplicate to ensure accuracy and reproducibility. Data were analyzed using one-way analysis of variance (ANOVA) with SPSS Statistics (Version 26; IBM Co., Armonk, NY, USA). Duncan’s multiple range test was applied to determine significant differences among means, with statistical significance set at P < 0.05. Results are expressed as mean ± standard deviation (SD).
Results
Microbiological quality and hazard index
Table 1 presents the microbiological assessment, which revealed substantial variations in microbial quality and hazard levels among Kapi samples from different production systems. The commercial GMP/HACCP-certified product (M1) demonstrated the highest safety level, with a very low total viable count (TVC, 1.97 log CFU/g), well below the 5 log CFU/g hygienic threshold, and the absence of pathogenic organisms. These results reflect effective hygienic and environmental control throughout the production process. In contrast, the sample traditionally produced with a controlled 25% (w/w) salt concentration (M2) exhibited the highest TVC (7.31 log CFU/g), exceeding both other production practices and the recommended safety limit. Nevertheless, this process effectively inhibited the growth and survival of E. coli, Salmonella spp., and C. perfringens at bactericidal levels. However, it was less effective against Gram-positive bacteria such as B. cereus (3.45 log CFU/g) and S. aureus, which remained detectable, although the latter was present at a low and acceptable level (<2 log CFU/g).
| Sources | LAB | TVC | Yeasts and molds | Foodborne pathogens | ||||
|---|---|---|---|---|---|---|---|---|
| E. coli | S. aureus | B. cereus | Salmonella | C. perfringens | ||||
| M1 | 1.96 ± 0.10c | 1.97 ± 0.35d | ND | ND | ND | ND | ND | ND |
| M2 | 3.38 ± 0.12a | 7.31 ± 0.20a | 1.81 ± 0.40c | ND | 1.94 ± 0.10d | 3.45 ± 0.14a | ND | ND |
| M3 | 3.29 ± 0.22a | 5.38 ± 0.34b | 4.08 ± 0.09a | ND | 4.47 ± 0.06a | 3.59 ± 0.10a | ND | Detected |
| M4 | 2.68 ± 0.31b | 5.71 ± 0.21b | 2.55 ± 0.20b | ND | 2.75 ± 0.65c | 3.19 ± 0.17b | ND | ND |
| M5 | 2.73 ± 0.09b | 5.63 ± 0.17b | 2.15 ± 0.25bc | ND | 3.49 ± 0.16b | 1.96 ± 0.07d | ND | Detected |
| M6 | 2.49 ± 0.60bc | 3.48 ± 0.60c | ND | ND | 1.89 ± 0.11d | ND | ND | ND |
| M7 | 2.31 ± 0.36bc | 5.36 ± 0.10b | ND | ND | 2.03 ± 0.10d | 2.52 ± 0.05c | ND | ND |
Notes:
M1 (commercial export product, control), M2 (local enterprise producers utilizing the recommended 25% salt concentration), M3 and M4 (samples from local wet-markets production practices, market 1 and 2, respectively), M5, M6 and M7 (samples from different local enterprise producers production practices). Different lowercase letters within the same column indicate statistically significant differences P < 0.05. A TVC below 5 log CFU/g indicates that the product meets hygienic standards. The presence of Salmonella sp. and C. perfringens, which are foodborne pathogens, was assessed using detection/non-detection methods, with ‘ND’ indicating that the pathogen was not detected in the sample.
When compared with M3 and M4, which were produced without processing control, packaging, or environmental protection, the microbial patterns were largely consistent with those observed in M2. However, the counts of S. aureus were approximately 2 log CFU/g higher in the uncontrolled samples, and C. perfringens was detected exclusively in these products, indicating a complete lack of inhibition under open and unhygienic production conditions. In the case of M5, M6, and M7, which were produced under basic control measures following the minimum standards typically applied by small community enterprises, considerable variation in microbial profiles was observed. Although E. coli and Salmonella spp. were not detected, as in the other samples, C. perfringens was found in M5, accompanied by total viable counts (TVC) exceeding the acceptable safety limit (5 log CFU/g). Notably, yeasts and molds were not detected in the GMP/HACCP-controlled sample (M1) or in the minimally controlled community-enterprise products (M6 and M7). However, a low level of fungal contamination (1.81 log CFU/g) was observed in the salt-controlled sample (M2). Certain lactic acid bacteria (LAB) species may function as natural starter cultures that initiate spontaneous fermentation and enhance product quality. In this study, LAB counts were low in the GMP/HACCP-controlled sample (M1) (<2 log CFU/g) but ranged between 2–3 log CFU/g in samples M2–M7. Therefore, 16S rRNA gene profiling was employed to characterize bacterial populations and elucidate the roles of these naturally occurring LAB and other bacteria in fermentation dynamics and product safety.
Bacterial community profile of shrimp paste from different production practices
Sequencing analysis
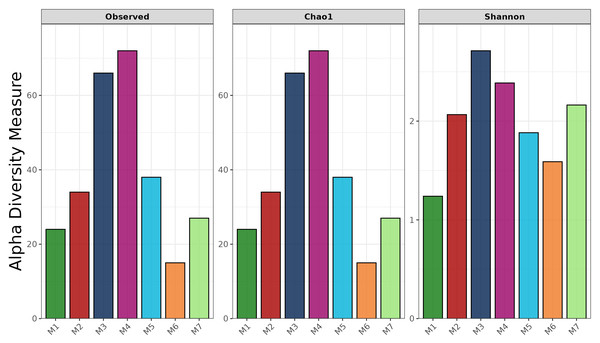
The full-length 16S rRNA gene was amplified and sequenced for all fermented shrimp paste samples using Oxford Nanopore Technologies. The observed ASV (Amplicon Sequence Variant), Chao 1, and Shannon indices are shown in Fig. 1. The observed ASV counts in M3 and M4 were 66 and 72, respectively, which were the highest among all samples. In contrast, the ASV count in M6 was 15, indicating markedly lower richness compared to other samples. Moreover, similar results were also observed in Chao1 richness index. This pattern was mirrored in the Chao1 richness index, which also ranked M6 lowest. The Shannon index showed that M1 had the lowest evenness, indicating a less evenly distributed bacterial community compared to other samples. Additionally, M5, M6, and M7 all showed lower observed ASVs, Chao1 richness, and Shannon diversity than M3 and M4.
Figure 1: Alpha-diversity indices of bacterial communities in fermented shrimp paste from different production practices: observed ASVs, Chao 1 richness, and Shannon diversity.
Bacterial composition in fermented shrimp paste
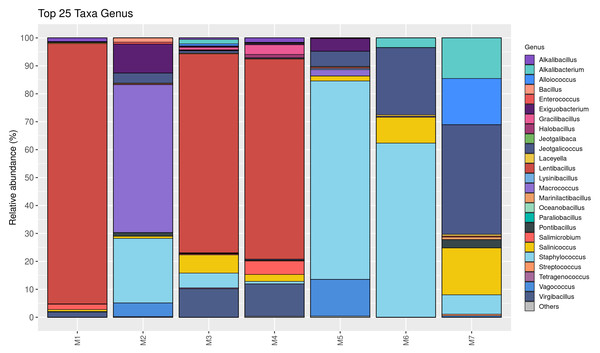
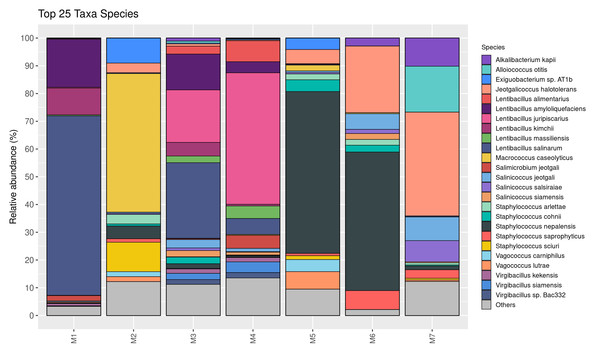
The relative abundance of bacterial genera is shown in Fig. 2. Twenty-five taxa were identified at the genus level in this study. Lentibacillus (93.2%) was the predominant microbiota in M1 sample, while Macrococcus (52.9%), Staphylococcus (23.1%), and Exiguobacterium (10.4%) were threemost abundant genera in M2 sample. Lentibacillus (≈ 71%) and Virgibacillus (≈ 10%) were significantly abundant in M3 and M4. Staphylococcus was the dominant genus found in M5 and M6, accounting for 71.1% and 62.4%, respectively. However, Jeotgalicoccus (39.3%), Salinicoccus (16.8%), and Alloiococcus (16.6%) were the dominant genera in M7. Figure 3 displays the relative abundance of different species in each sample. Twenty-five taxa were identified at the species level. Lentibacillus salinarum, Lentibacillus amyloliquefaciens, and Lentibacillus kimchi were the three most abundant species in M1 sample. Macrococcus caseolyticus (50.0%), Staphylococcus sciuri (10.7%) and Exiguobacterium sp. AT1b (9.0%) were dominant in M2. Lentibacillus juripiscarius, Lentibacillus alimentarius, and L. salinarum were found in both M3 and M4. In addition, Staphylococcus nepalensis and Jeotgalicoccus halotolerans were identified in both M5 and M6. J. halotolerans (37.4%), Alloiococcus otitis (16.6%), and Alkalibacterium kapii (10.1%) were the three most dominant species in M7.
Figure 2: Bacterial composition in fermented shrimp paste from different production practices: genus-level profile.
Figure 3: Bacterial composition in fermented shrimp paste from different production practices: species-level profile.
Prediction of enzyme classification and metabolic pathways in fermented shrimp paste
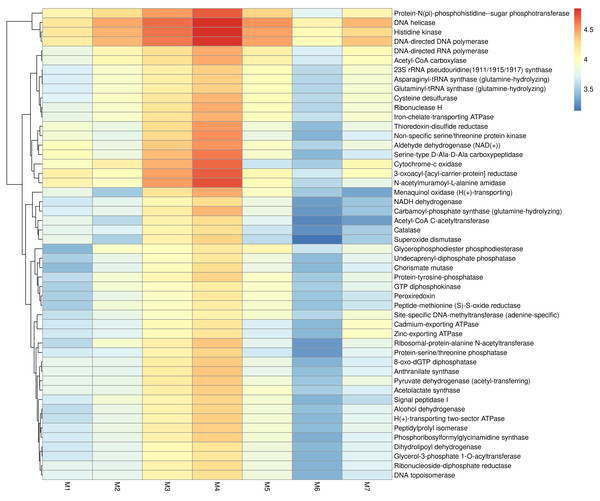
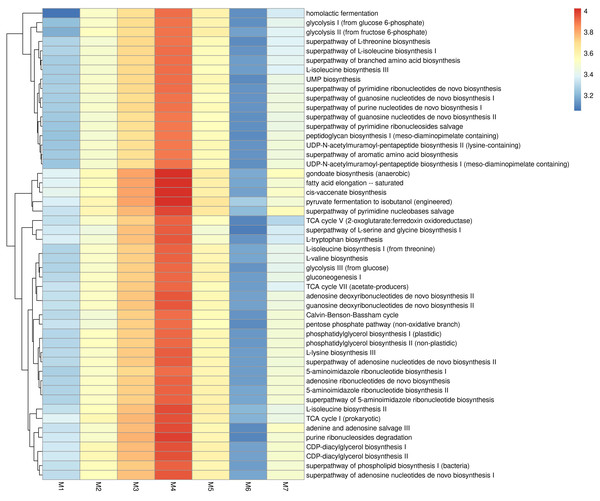
Figure 4 presents the predicted KEGG enzyme classifications derived from microbiota, based on comparisons of their 16S rRNA profiles. Among all samples, M4 showed the highest metabolic potential, while M6 showed the lowest. Three enzymes, DNA helicase, histidine kinase, and DNA-directed DNA polymerase, were found across all samples. In addition, Protein-N(pi)-phosphohistidine sugar phosphotransferase, DNA helicase, histidine kinase, DNA-directed DNA polymerase, cytochrome-c oxidase, 3-oxoacyl-[acyl-carrier-protein] reductase, and N-acetylmuramoyl-Lalanine amidase were highly prevalent in M4. The metabolic potential profile of M3 was relatively similar to that of M4. As shown in Fig. 5, M3 and M4 showed the highest KEGG metabolic pathways, while M6 showed the lowest. Metabolic pathways such as gondoate biosynthesis (anaerobic), fatty acid elongation-saturated, cis-vaccenate biosynthesis, pyruvate fermentation to isobutanol (engineered), superpathway of pyrimidine nucleobases salvage, adenine and adenosine salvage III, TCA cycle I (prokaryotic), CDP-diacylglycerol biosynthesis I and II, purine ribonucleosides degradation, and L-lysine biosynthesis III were highly prevalent in M3 and M4.
Figure 4: KEGG enzyme classifications identified in fermented shrimp paste samples collected from different production practices.
Figure 5: KEGG metabolic pathways identified in fermented shrimp paste samples collected from different production practices.
Physicochemical characteristics in fermented shrimp paste from different production practices
Table 2 presents the physicochemical characteristics of fermented shrimp paste obtained from various production practices. The samples showed statistically significant differences (P < 0.05) in pH, titratable acidity (%), salt concentration (%), nitrogen content, protein content, and histamine content. The pH values ranged from 6.76 to 7.58. The highest pH was observed in sample M4 (7.58), suggesting either lower acid production or more advanced proteolysis during fermentation. Conversely, M6 showed the lowest pH (6.76), indicating higher acidity, which was consistent with its higher acid content (25.4%). Titratable acidity content varied significantly among the samples. The highest acidity percentage was detected in M6 (25.4%) and M7 (24.0%), which could indicate more intense fermentation activity or the presence of acid-producing bacteria. In contrast, M4 exhibited the lowest acid content (14.7%), aligning with its higher pH value. Salt concentrations ranged from 20.6% (M3) to 35.5% (M7). Samples M5 and M7 had the highest salt levels, likely due to production practices aimed at preservation. Salt concentration not only influences microbial activity but also impacts texture and flavor of shrimp paste. Nitrogen content and protein percentage were highest in M4 (5.83% N and 36.5% protein). Histamine content ranged from 9.5 mg/kg to 39.4 mg/kg. M6 exhibited the highest histamine concentration (39.4 mg/kg), while M4 had the lowest (9.5 mg/kg). The higher histamine levels observed in M6 and M7 could be attributed to prolonged fermentation, temperature conditions, or the presence of histamine-producing microorganisms.
| Sources | pH value | Acid (%) | Salt concentration (%) | N-content (%) | Protein (%) | Histamine content (mg/kg) |
|---|---|---|---|---|---|---|
| M1 | 7.48 ± 0.04b | 18.9 ± 0.25d | 24.9 ± 0.73d | 3.9 ± 0.02e | 24.9 ± 0.14e | 12.7 ± 0.12f |
| M2 | 7.35 ± 0.05c | 20.4 ± 0.10c | 20.8 ± 0.75e | 4.3 ± 0.04d | 27.1 ± 0.27d | 22.9 ± 0.05d |
| M3 | 7.56 ± 0.04a | 17.7 ± 0.16e | 20.6 ± 0.96e | 4.0 ± 0.02e | 25.3 ± 0.13e | 25.8 ± 0.07c |
| M4 | 7.58 ± 0.10a | 14.7 ± 0.69f | 27.3 ± 0.47c | 5.8 ± 0.17a | 36.5 ± 1.09a | 9.5 ± 0.05g |
| M5 | 7.02 ± 0.01d | 20.3 ± 0.72c | 31.8 ± 0.54b | 5.5 ± 0.17b | 34.1 ± 1.07b | 18.4 ± 0.05e |
| M6 | 6.76 ± 0.02e | 25.4 ± 0.40a | 21.9 ± 0.82e | 5.1 ± 0.11c | 31.9 ± 0.69c | 39.4 ± 0.02a |
| M7 | 7.06 ± 0.01d | 24.0 ± 0.06b | 35.5 ± 1.67a | 5.2 ± 0.03c | 32.7 ± 0.19c | 32.0 ± 0.06b |
Notes:
M1 (commercial export product, control), M2 (local enterprise producers utilizing the recommended 25% salt concentration), M3 and M4 (samples from local wet-markets production practices, market 1 and 2, respectively), M5, M6 and M7 (samples from different local enterprise producers production practices).
Different lowercase letters within the same column indicate statistically significant differences P < 0.05. According to the Codex Alimentarius Commission, eleven seafood commodity standards include a histamine safety provision under Section 5 (Hygiene), stating that no sample unit may exceed 200 mg/kg.
Discussion
Intrinsic factors such as pH, acidity, salt, and protein content play key roles in determining microbial growth, whereas histamine serves as a microbial metabolite and an indicator of microbial activity and food safety (Jay, Loessner & Golden, 2005). The findings of this study were consistent with those reported by Nakamura et al. (2022), particularly for samples M2-M7, which were produced under different production practices from the wet markets and local enterprises. These samples exhibited high levels of LAB and TVC, comparable to values reported in other fermented shrimp pastes sold in supermarket and wet-markets. S. aureus was detected in M2-M7 (1.89–4.47 log CFU/g). This pathogen is known to sense external salt stress, activating its adaptive mechanisms that enable survival under high-salinity conditions (Feng et al., 2022). Likewise, Sumardianto, Purnamayati & Rianingsih (2024) revealed that sodium content regulates water activity (<0.60), thereby restricting S. aureus proliferation. However, the viable count of S. aureus in M1, M2 and M6 remained within acceptable safety limits defined by the Thai Community Product Standard, which specifies a maximum of 2.00 log CFU/g (Thai Community Product Standard, 2018). B. cereus was detected in M2, M3, M4, M5, and M7, with counts ranging from 1.96−3.59 log CFU/g. Although below the threshold considered hazardous to human health (>6 log CFU/g) (Batt, 2000), the viable counts in M2, M3, and M4 exceeded the product safety standards (Thai Community Product Standard, 2018). In addition, C. perfringens was detected in both M3 and M5. According to the microbiological quality standards for fermented shrimp paste, C. perfringens must not be present, as it can produce five distinct toxigenic types (A-E) and is known to inhabit the human gastrointestinal tract (Brynestad & Granum, 2002; TIS, 1992). In this study, its detection in M3 and M5 therefore indicates non-compliance with safety criteria, suggesting insufficient hygienic control during production processes, including raw material handling, packaging, and post-packaging management. Regarding yeasts and molds, samples M1, M2, M4, M5, M6, and M7 fell within acceptable safety limits, but M3 exceeded the permissible standard (Thai Community Product Standard, 2018). This finding aligns with Pongsetkul, Benjakul & Boonchuen (2022), who reported the presence of yeast and mold in fermented shrimp paste stored in various packaging containers. The lack of GMP implementation and the open selling conditions observed at wet-markets (M3 and M4) likely contributed to TVC exceeding 5 log CFU/g, indicating an increased risk of contamination from poor hygiene practices and environmental expose. Furthermore, the warm temperatures and high humidity characteristic of Thailand and the broader Southeast Asian region create ideal conditions for the survival and growth of bacteria. Most bacteria thrive in warm, moist environments, typically between 30–40 °C, where metabolic activity and cell reproduction are optimized. Under such conditions, food left at room temperature can quickly enter the danger zone (5–60 °C), allowing pathogens such as S. aureus, B. cereus, Salmonella, C. preeminence, and mold with to multiply rapidly. These environmental conditions significantly increase the risk of food spoilage and foodborne illness if proper handling and storage practices are not followed. Consequently, Anal et al. (2020) suggested that contamination in fermented products is primarily attributed to high humidity and temperature conditions that support the growth of microorganisms and toxins. Additionally, the wet or fresh market environment significantly contributes to microbial contamination due to inadequate hygiene practices in these settings (Karanth et al., 2023). The findings of this study further demonstrate that even in uncontrolled production environments lacking full GMP implementation, packaging control, and standardized salt regulation, the survival and proliferation of Gram-negative bacteria such as E. coli and Salmonella spp. can be effectively inhibited. However, these conditions proved insufficient to suppress or eliminate Gram-positive bacteria including S. aureus, B. cereus, and C. perfringens as well as fungal contaminants. Maintaining a salt concentration of approximately 25% (w/w) resulted in a noticeable reduction of C. perfringens contamination; nonetheless, this level remained inadequate for controlling S. aureus and B. cereus, highlighting a persistent microbial safety challenge that cannot be resolved through salinity adjustment alone. Similarly, production systems operating under minimal GHP conditions, representing the baseline standard typically applied by small-scale or local enterprises, exhibited significant reductions in S. aureus and B. cereus counts, alongside effective control of fungi, E. coli, and Salmonella spp. These results collectively indicate that neither salinity regulation nor hygiene management alone can ensure microbiological safety at a bactericidal level. Achieving a non-hazardous safety threshold therefore requires an integrated approach that simultaneously regulates both salinity and hygienic production factors. This combined framework provides essential evidence supporting the need for enhanced GMP/GHP compliance, environmental control, and process standardization to ensure microbial safety in community-based Kapi production.
Moreover, despite the completion of fermentation, LAB and several pathogenic bacteria remained detectable across samples, suggesting microbial persistence under different production conditions. To further elucidate these microbial dynamics, the 16S rRNA gene profiling results provided deeper insights into microbial diversity patterns across different production systems. Alpha diversity indices based on ASV-level analysis serve as useful tools for evaluating the richness and diversity of species in the microbial community. A higher value of Chao index reflects a greater species abundance within a sample, while the Shannon index quantifies species diversity, with higher values indicating more diverse microbial populations (Sun et al., 2016). In this study, the highest Shannon and Chao1 indices were observed in samples M3 and M4, suggesting that these samples - obtained from local fresh markets- exhibited the highest species diversity. These results suggest that naturally fermented, unpackaged shrimp paste from fresh markets, which are typically subject to less environmental control, harbor highly diverse and rich microbial community. This observation is likely a direct consequence of the uncontrolled fermentation conditions and exposure to environmental microorganisms (Palmnäs-Bédard et al., 2023; Deka et al., 2021). Notably, M2, which was produced by a local enterprise using the traditional salt concentration (25%), exhibited greater bacterial diversity than M1, a commercial product. This finding suggests that M2 underwent a more active and diverse fermentation process, likely influenced by semi-controlled microbiota. This phenomenon can be attributed to the fact that the raw materials used are derived from marine environments, where microorganisms are naturally adapted to survive under high-salt conditions. As a result, they can survive and remain viable under the high-salt conditions used in shrimp paste production, persisting for up to 3–9 months after the fermentation process. This is consistent with the findings of Du et al. (2019), who reported that in traditional fermented seafood containing 30% salt, bacterial diversity remained detectable up to the 15th month post-fermentation. Notably, Tetragenococcus was positively correlated with the fermentation process and contributed to improving the quality of fermented seafood. In contrast, the observed ASVs, Chao1, and Shannon indices of M5, M6, and M7 were lower than those of M3 and M4, indicating reduced microbial diversity. This pattern aligns with previous reports suggesting that industrial-scale production, which generally applies stricter microbial control and sanitation practices, tends to limit microbial variation (Tamang et al., 2020).
A detailed examination of the identified bacterial genera and species clarified their critical roles in the natural fermentation process of fermented shrimp paste from different production practices, highlighting how dominant species influence product safety. As shown in Figs. 2 and 3, Lentibacillus spp. and A. kapii were the most abundant bacteria in the samples. Lentibacillus spp. is a halophilic microorganism capable of growing in high NaCl content (5%–30%), across a wide pH range (5–9), and producing protease during fermentation. On the other hand, A. kapii is highly halotolerant, slightly halophilic, and alkaliphilic organism, with an optimal growth range of 0.5–4.0% (w/v) NaCl and a pH between 8.0–9.5, though it can survive in a broader pH range of 6.0–11.0, depending on the strain (Pakdeeto et al., 2007; Phewpan et al., 2020; Ishikawa et al., 2009). Furthermore, Macrococcus spp. are gram-positive cocci belonging to the Staphylococcaceae family. They are closely related to genus Staphylococci, but are generally not considered human food borne pathogens. M. caseolyticus has been proposed for use in starter culture consortia due to its proteolytic and lipolytic activities, which contribute to the aroma and flavor of fermented foods (Ramos, Vigoder & Nascimento, 2021). However, Zhou et al. (2012) and Sang et al. (2020) reported M. caseolyticus, J. halotolerans and S. sciuri as histamine-producing bacteria, suggesting that inhibiting these species during fermentation may help reduce histamine accumulation. Notably, these same bacteria are also responsible for the formation of desirable flavor in the shrimp paste. When co-cultured with Virgibacillus, Exiguobacterium promotes the growth of salt-tolerant bacteria while simultaneously reducing histamine content (Liu et al., 2024).
Overall, the coexistence of pathogenic and beneficial taxa revealed by 16S rRNA profiling reflects a complex ecological balance in fermented shrimp paste. While pathogens such as S. aureus, B. cereus, and C. perfringens survived the high-salinity conditions, halophilic genera such as Tetragenococcus, Lentibacillus, and Alkalibacterium likely contributed to product safety and quality through antimicrobial and proteolytic activities. These interactions are consistent with the detected functional pathways, suggesting that active genes involved in stress adaptation, energy metabolism, and amino-acid turnover support microbial survival and activity during fermentation. This functional overlap links the taxonomic composition to the metabolic potential, providing a basis for interpreting the predicted enzyme and pathway profiles observed in the following section. All samples were found to possess a wide range of microbial enzyme activities, including DNA helicase, histidine kinase, and DNA-directed DNA polymerase, which are key enzymes associated with the activity of viable microorganisms. Helicases are a class of enzymes essential to all microorganisms, functioning to unpack DNA strands during key genetic processes (Tuteja & Tuteja, 2004). Histidine kinases are multifunctional enzymes, typically transmembrane and protein transferases, which play an important role in signal transduction across the bacterial cell membrane (Kim & Forst, 2001). Furthermore, DNA-directed DNA polymerase plays a significant role in bacterial DNA replication by synthesizing new DNA strands using the existing DNA as a template (Bassam, Caetano-Anollés & Gresshoff, 1992). The highest levels of KEGG metabolic pathway activity were observed in M3 and M4, which aligns with the alpha diversity results, indicating that these samples exhibited the greatest bacterial diversity.
The integration of microbiological and functional analyses provides a holistic view of fermented shrimp paste. The coexistence of halophilic beneficial and pathogenic taxa explains the enzymatic and metabolic variations observed, which are closely influenced by physicochemical parameters such as pH, salinity, and total protein. These factors govern microbial succession, proteolysis, and amino-acid decarboxylation leading to histamine formation. The following section further elaborates on how these parameters reflect microbial activity and chemical safety across production systems. In the present study, the pH of all Kapi samples ranged from 6.76 to 7.58, while the salt concentration ranged from 20.6 to 35.5%(w/w), reflecting the slightly alkaline conditions typical of traditional high-salt fermentation. These results are consistent with those reported by Li et al. (2021), who found that Huanghua shrimp paste from China exhibited pH values of 6.77–8.25, with dominant bacterial phyla Firmicutes and Proteobacteria, and genera such as Tetragenococcus, Lactobacillus, and Halanaerobium, which were positively correlated with histamine and amino-nitrogen levels. Similarly, Pilapil et al. (2016) observed that both traditional and commercial shrimp pastes from the Philippines maintained pH values of 6.8–7.7, contributing to microbial stability with low water activity. In contrast, Li et al. (2022) examined low-salt shrimp paste (10% NaCl) fermented at 10 °C–20 °C and reported a gradual pH decrease from 8.50 to 8.01, attributed to organic acid formation under reduced salinity and cold conditions. Compared with these low-temperature fermentations, the lower pH range (6.76–7.58) observed in the present study likely resulted from both higher salt content and tropical ambient temperatures. These conditions favor mixed halophilic populations, particularly Lentibacillus and Tetragenococcus, which produce lactic and acetic acids that maintain a balanced acid–base stability throughout fermentation.
Furthermore, the elevated titratable acidity detected in all Kapi samples reflects a high accumulation of organic acids generated by microbial metabolic activity, further confirming the contribution of halotolerant LAB to acid regulation. This observation aligns with Yusof, Zulkifli & Jusoh (2023), who found that commercial shrimp paste with higher salinity exhibited higher titratable acidity and lower pH values (7.00–7.32), demonstrating a direct correlation between salt concentration, organic acid formation, and microbial fermentation activity. Together, these consistent findings emphasize that pH and titratable acidity are key indicators of fermentation dynamics and microbial balance in traditional shrimp paste. Building on this, the acid-base equilibrium maintained by halotolerant LAB not only stabilizes the pH of the product but also promotes proteolytic activity, leading to increased nitrogen release and protein breakdown during fermentation. In the present study, the nitrogen and protein contents across all Kapi production practices ranged from 3.98–5.83% and 24.9–36.5%, respectively, reflecting active proteolysis under high-salt fermentation conditions. These microbial activities are also responsible for the generation of biogenic amines and nitrites, which are formed through the microbial decarboxylation of amino acids and nitrogenous compounds (Lee, Jung & Jeon, 2015). The presence of these compounds suggests a high degree of protein hydrolysis and may indicate either advanced fermentation progress or superior raw material quality. Thus, an important finding from the risk assessment is the clear relationship between histamine levels and the presence of halophilic and salt-tolerant pathogenic bacteria, representing both chemical and microbiological hazards. These dual hazards, histamine as a toxic compound and the associated pathogens, highlight the need for integrated control strategies that simultaneously manage contamination across different practices and protect consumer health. To safeguard food safety and ensure trade compliance, international authorities such as the Codex Alimentarius, European Regulation (EC) No. 2073/2005, and FDA/WHO have established strict limits for histamine in fishery products (El Hariri, Bouchriti & Bengueddour, 2018). In this context, our study further classified the combined chemical and microbiological risks of fermented shrimp paste based on histamine concentration and dominant bacterial profiles. This approach provides a practical and evidence-based framework for evaluating safety levels across different production systems.
Thus, in this study, both chemical and microbiological hazards in fermented shrimp paste were successfully categorized into three distinct histamine concentration ranges, each associated with different dominant bacterial species. The lowest histamine levels were found in samples M1, M4, and M5 (9.5–18.4 mg/kg), low-moderate levels in samples M2 and M3 (22.9–25.8 mg/kg), and moderate levels in samples M6 and M7 (32.0–39.9 mg/kg). These results represent the study’s original findings, indicating that M1, a GMP-certified product, showed the best safety profile characterized by low histamine levels and the absence of major pathogens. Samples M4 and M5, from wet-market and local enterprise production practices, respectively, also exhibited similarly low histamine concentrations. This suggests that M1 and M4 samples were dominated by the halophilic bacterium L. salinarum, which has been reported to enhance proteolysis and amino acid production during fermentation, contributing to umami flavor without histamine or otherbioginic amines (Udomsil et al., 2022; Du et al., 2019). Therefore, L. salinarum demonstrates potential as a beneficial starter culture for producing high-quality fermented shrimp paste while simultaneously reducing overall hazard levels. At the low-moderate histamine level, samples M2 (25% w/w salt concentration), M5, and M6 (produced under local small enterprise practices), showed histamine levels that remained within the safe limit (<50 mg/kg), although a slight upward trend was observed compared to M1 and M4. Notably, the detection of S. sciuri, a halophilic bacterium previously described by Ma et al. (2022) for its histamine-degrading potential in fermented products, suggests that this species may serve as a beneficial starter culture for controlling histamine accumulation in traditional fermentation. At the medium histamine level, samples M6 and M7, produced by different local enterprises exhibited concentrations ranging from 32.0–39.9 mg/kg. These levels may be attributed to the predominance of J. halotolerans, a species reported to possess strong tendency for biogenic amine production (Sang et al., 2020). Although these levels were higher than those observed in other samples, they still remained within safety limits and did not exceed the threshold value of 50 mg/kg. According to the FAO/WHO Expert Committee, the NOAEL (No Observed Adverse Effect Level) is defined as 50 mg histamine per 250 g seafood portion, equivalent to 200 mg/kg, while the FDA considers levels above 50 mg/kg as decomposed and over 500 mg/kg as a health hazard (Food and Agriculture Organization of the United Nations and World Health Organization, 2012; Lo Magro et al., 2020). This study provides new insights into the combined chemical and microbiological hazards of fermented shrimp paste. Although histamine was detected across all production systems, its concentration remained below 50 mg/kg. This level is well within the Codex safety limit, indicating low chemical risk. However, the detection of C. perfringens in samples from the wet-market (M3) and local enterprise (M5) revealed a potential microbiological hazard, as this toxigenic species is strictly prohibited under safety standards. The coexistence of low chemical risk and sporadic pathogenic contamination underscores the need to strengthen safety management. Community-based Kapi production should advance from basic GHP to GMP-level operations to ensure process control and product consistency. Such advancement would also enable the use of selected halophilic starter cultures, such as Lentibacillus or Tetragenococcus, to stabilize fermentation, suppress pathogens, and minimize biogenic amine formation. Furthermore, maintaining an optimal salinity level (25% w/w) with continuous hazard monitoring will further enhance the microbial and chemical safety of fermented shrimp paste.
Conclusions
This study evaluated seven Kapi samples produced under different hygienic conditions, revealing a clear link between production environment, microbial diversity, and histamine accumulation. Although histamine was detected in all samples, concentrations remained below international safety limits (≤50 mg/kg). Based on an integrated chemical–microbiological assessment, histamine risk was classified as low (9.5–18.4 mg/kg; M1, M4, M5), low-moderate (22.9–25.8 mg/kg; M2, M3), and moderate (32.0–39.9 mg/kg; M6, M7). Lentibacillus salinarum and Staphylococcus sciuri exhibited potential as histamine-degrading starter cultures that could stabilize fermentation and reduce biogenic amine formation. Wet-market products (M3, M4) showed the highest microbial loads and diversity, reflecting insufficient hygiene and environmental control. These findings demonstrate that maintaining 25% (w/w) salt alone cannot ensure product safety; effective control requires GMP, hygienic processing, and environmental regulation particularly of temperature and humidity during fermentation and storage. An integrated, risk-based framework is therefore proposed, combining salt calibration, hygienic process design, and the use of beneficial halophilic microorganisms to enhance microbial and physicochemical stability. This scalable approach enables small-scale and community producers to progress toward GMP compliance, improving product safety, consistency, and market competitiveness. Although enzyme classifications and metabolic pathways were predicted from 16S rRNA profiles, functional validation through experimental studies remains necessary to confirm these activities and clarify their roles in the fermentation process.
Supplemental Information
Complete raw dataset
Microbial diversity indices, physicochemical quality parameters, and safety-related measurements of fermented shrimp paste samples