Genome-wide identification and drought-responsive root expression profiling of the VQ gene family in proso millet (Panicum miliaceum L.)
- Published
- Accepted
- Received
- Academic Editor
- Nicole Nogoy
- Subject Areas
- Bioinformatics, Genomics, Molecular Biology, Plant Science
- Keywords
- Proso millet, VQ gene family, Genome-wide analysis, Drought stress, Expression analysis, Germplasm resources
- Copyright
- © 2026 Tan et al.
- Licence
- This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits using, remixing, and building upon the work non-commercially, as long as it is properly attributed. For attribution, the original author(s), title, publication source (PeerJ) and either DOI or URL of the article must be cited.
- Cite this article
- 2026. Genome-wide identification and drought-responsive root expression profiling of the VQ gene family in proso millet (Panicum miliaceum L.) PeerJ 14:e20792 https://doi.org/10.7717/peerj.20792
Abstract
Background
Proso millet (Panicum miliaceum L.) is a drought-tolerant cereal crop cultivated in arid and semi-arid regions. VQ proteins, a class of plant-specific proteins characterized by a conserved VQ motif (FxxxVQxxTG), are known to play critical roles in plant responses to abiotic stress. To elucidate the genetic basis of drought tolerance in proso millet, a genome-wide identification and characterization of the VQ gene family was undertaken.
Methods
This study involved the identification of all VQ family members from the proso millet genome, followed by comprehensive analyses including chromosomal localization, phylogenetic relationships, gene structure, conserved motifs, collinearity, and promoter cis-acting elements. Expression profiling was conducted using transcriptomics and Quantitative Real-Time Reverse Transcription Polymerase Chain Reaction (qRT-PCR) to investigate potential gene functions. Additionally, the drought tolerance of various germplasm materials was systematically assessed.
Results
A total of 70 VQ genes (PmVQ1–PmVQ70) were identified and classified into four distinct subfamilies. Based on the screening of root-preferential and drought-responsive candidate genes, combined with phenotypic and qRT-PCR analyses of drought-tolerant and drought-sensitive materials, four key candidate genes were identified. qRT-PCR analysis revealed that the four genes exhibited differential expression patterns between drought-tolerant and drought-sensitive materials, suggesting their potential roles as core regulators in proso millet’s drought response, particularly in root-specific regulatory pathways under drought stress. This study provides a systematic analysis of the VQ gene family in proso millet and offers valuable genetic resources for elucidating drought tolerance mechanisms and advancing molecular breeding.
Introduction
Climate change-induced environmental stresses, particularly drought, pose serious threats to global agriculture. Against the backdrop of a continuously growing global population, it has become increasingly critical to maintain crop yields in order to meet fundamental demands (Boyer, 1982; Chaves & Oliveira, 2004; Gupta, Rico-Medina & Cano-Delgado, 2020). Therefore, elucidating the molecular mechanisms underlying drought tolerance and developing high-yielding, water-efficient crop varieties have emerged as urgent priorities in agricultural research.
Proso millet (Panicum miliaceum L.), an annual C4 grass species, is characterized by its short growth cycle, high water-use efficiency, and remarkable tolerance to abiotic stresses, especially drought (Zou et al., 2019). Coupled with features such as a compact genome, self-pollination, and a short reproductive period, proso millet has emerged as an ideal model species for studying abiotic stress tolerance—particularly drought adaptation mechanisms—in cereal crops (Diao, 2017). It is also recognized as a promising candidate crop for addressing future challenges in food and nutritional security. Therefore, systematically elucidating the genetic basis and molecular mechanisms underlying drought tolerance in proso millet is of strategic importance for crop improvement and global food security. Unraveling the genetic basis of drought tolerance and deciphering the molecular mechanisms involved will be crucial for genetic improvement of crops and enhancement of global food security (Chen et al., 2023; Cong et al., 2025; Shan et al., 2020).
Plants initiate a series of complex molecular regulatory networks in response to environmental stress. Among these, the VQ gene family, a class of plant-specific transcriptional regulatory cofactors, plays a critical role. VQ proteins contain a highly conserved “FxxxVQxTG” amino acid motif. While they typically lack intrinsic DNA-binding capacity, they exert their functions by interacting with other key regulatory proteins. Extensive research has shown that VQ proteins are central players in plant responses to abiotic stresses such as drought (Cheng et al., 2012). By interacting with diverse partner proteins, VQ proteins participate in various physiological processes, including plant growth and development, immune responses, and abiotic stress adaptation (Hu et al., 2013; Lai et al., 2011; Tan et al., 2024). Current research on VQ proteins has largely focused on their interaction with WRKY transcription factors and mitogen-activated protein kinases (MAPKs) (Pecher et al., 2014; Wang et al., 2015).
Accumulating evidence underscores the importance of VQ genes in modulating drought tolerance across species. For instance, overexpression of TaVQ4-D in Arabidopsis and wheat significantly improves drought tolerance, whereas silencing TaVQ4-D reduces drought tolerance. TaVQ4-D physically interacts with MPK3 and MPK6 and serves as their phosphorylation substrate in the drought response (Zhang et al., 2023). Overexpression of MdVQ37 upregulates SA catabolism genes, leading to reduced endogenous SA levels under drought and negatively regulating drought tolerance in apple (Dong et al., 2022). IbWRKY2 enhances drought and salt tolerance and interacts with IbVQ4, whose expression is significantly induced by PEG (polyethylene glycol) treatment, suggesting that IbVQ4 may be a key regulator of drought tolerance in sweet potato (Zhu et al., 2020). In maize, ZmVQ56 negatively regulates drought and low-nitrogen stress tolerance by modulating root development and nitrogen metabolism pathways (Xu et al., 2025).
The root system serves as a crucial organ for perceiving and adapting to drought, coordinating morphological, physiological, and molecular responses to enhance plant drought tolerance. Roots function not only as water-absorbing organs but also as regulatory hubs that integrate stress signals and coordinate whole-plant drought resilience (Chen et al., 2021; Marik & Sadhukhan, 2025). Understanding these root-mediated mechanisms provides an essential foundation for breeding drought-tolerant crops and developing sustainable agricultural practices.
Although the VQ gene family has been relatively well studied in crops such as Arabidopsis, rice, wheat, and maize, research on this gene family in proso millet remains notably limited. To date, the genome-wide identification, evolutionary characteristics, and expression regulation patterns of the VQ gene family in proso millet under drought stress, particularly its role in roots as a key organ for stress perception, are still unclear. In this study, we conducted a comprehensive bioinformatic analysis of the VQ gene family in proso millet, combined with drought phenotyping of multiple accessions and gene expression validation. We identified four key VQ genes with root-specific expression that are closely associated with drought tolerance. These findings systematically reveal the molecular characteristics of the VQ gene family in proso millet and its potential role in drought response, providing new insights and key genetic resources for a deeper understanding of drought tolerance mechanisms in proso millet.
Materials and Methods
Plant materials and drought treatment
Plant materials. Twelve proso millet germplasm accessions with varying degrees of drought tolerance were used in this study, including both wild and cultivated types. All accessions originated from China and were obtained from the germplasm accessions bank of the Institute of Millet Crops, Hebei Academy of Agriculture and Forestry Sciences. Detailed information including full names, abbreviations, Chinese names, collection locations, and collection dates is provided in Table S1.
Seed sterilization and germination. Fifty plump seeds from each accession were surface-sterilized in 10% (v/v) sodium hypochlorite solution for 15 min, rinsed thoroughly five times with distilled water, and then placed on petri dishes lined with two layers of moistened filter paper (supplemented with 10 mL distilled water). The dishes were incubated in a climate-controlled growth chamber set at 28 °C, 60% relative humidity, 30,000 lux light intensity, and a 16-h light/8-h dark photoperiod.
Drought treatment and sample collection. At the three-leaf stage, 20 uniformly grown seedlings per accession were transferred into a hydroponic system containing either 20% (w/v) PEG-6000 solution (simulated drought stress) or 1/2 Hoagland nutrient solution (control). After 5 days of treatment, whole root systems (from the root cap to the root tip) were harvested separately from the drought stress and control groups immediately flash-frozen in liquid nitrogen, and stored at −80 °C for subsequent RNA extraction and qRT-PCR analysis. Leaf tissues were collected in parallel for physiological measurements. All experiments were conducted with three independent biological replicates.
Identification and chromosomal localization of VQ gene family in proso millet
Genomic and annotation data for proso millet (accession number: GWHAAEZ00000000.1) were retrieved from the Genome Warehouse of the BIG Data Center (http://bigd.big.ac.cn/gwh). Arabidopsis thaliana, Oryza sativa, and Zea mays were selected as reference species for comparative genomic analysis based on the following considerations: (1) these species are well-annotated model plants with systematically characterized VQ gene families; (2) they represent both dicot (Arabidopsis) and monocot (rice and maize) lineages, providing broad evolutionary coverage; (3) as cereal crops closely related to proso millet, rice and maize facilitate the identification of conserved and lineage-specific VQ genes within the grass family; and (4) maize shares a C4 photosynthetic pathway with proso millet, offering functional insights relevant to drought adaptation. Comparative genomic data for Arabidopsis, rice, and maize, including genome annotations and protein sequences, were acquired from the EnsemblPlants database (https://plants.ensembl.org/index.html). To identify VQ family proteins, the hidden Markov model (HMM) profile for the VQ domain (PF05678) was retrieved from the Pfam database (http://pfam.xfam.org/). This profile was then employed to screen the proso millet genome using TBtools, with a threshold E-value of 1e−5 to identify candidate VQ genes (Chen et al., 2020).
Key physicochemical properties of the identified VQ proteins—such as amino acid length, theoretical isoelectric point (pI), and molecular weight (MW)—were predicted via the ProtParam online tool (http://web.expasy.org/protparam/). Chromosomal positions of the PmVQ genes were determined using TBtools, and a physical map illustrating their genomic distribution was constructed.
Phylogenetic relationships, gene structure, and conserved motif analysis of VQ genes
To elucidate the evolutionary relationships and systematic classification of VQ genes, a phylogenetic tree was generated from the full-length amino acid sequences of VQ proteins from proso millet, Arabidopsis, maize, and rice. Multiple sequence alignment was performed using MUSCLE, followed by tree construction in IQ-TREE under the Neighbor-Joining algorithm with 1,000 ultrafast bootstrap replicates. The resulting phylogenetic tree was visualized and annotated in iTOL (https://itol.embl.de). Conserved motifs within the PmVQ proteins were identified using the MEME online tool, with the maximum number of motifs specified as 10.
Analysis of gene duplication events in the VQ family
To investigate the duplication mechanisms of VQ genes in proso millet, intra-specific duplication events were examined through the Advanced Circos function in TBtools. Furthermore, inter-species collinearity analysis was performed among proso millet, Arabidopsis, rice, and maize using the One-Step MCScanX tool in TBtools. The synonymous substitution rate (Ks), non-synonymous substitution rate (Ka), and their ratio (Ka/Ks) were calculated using both MEGA11 and TBtools to assess selective pressures on duplicated gene pairs.
Expression patterns of PmVQ genes in different tissues and under drought stress
Transcriptomic data mining and visualization. Publicly available transcriptome datasets from proso millet were utilized to analyze tissue-specific and drought-responsive expression profiles of PmVQ genes. The transcriptomic data of drought treatment were derived from the roots of the cultivar ‘Yumi 2’ under five time points: control (0 h, labeled CK) and drought-stressed seedlings at 0.5, 1.0, 1.5, and 3.0 h (labeled D1–D4), each with three biological replicates. Expression heatmaps were generated using TBtools based on normalized log2 FC values (Zhang et al., 2024; Zou et al., 2019).
qRT-PCR validation under drought stress. To validate the expression patterns of selected PmVQ genes in response to drought, roots from accessions subjected to the drought treatment described in “Plant materials and drought treatment” were used for qRT-PCR analysis. Total RNA was extracted from proso millet roots using FreeZol Reagent (R711-01; Vazyme, Nanjing, China) and reverse transcribed into cDNA using HiScript III RT SuperMix (R323-01; Vazyme, Nanjing, China). The qRT-PCR analysis was performed on a Bio-Rad CFX96 with ChamQ Universal SYBR qPCR Master Mix (Q711-03; Vazyme, Nanjing, China). The qRT-PCR primers for analysis are listed in Table S2. PmActin served as the internal control. The fold change in the expression levels of target genes was calculated via relative quantification (2−ΔΔCT). Statistical differences among groups in the qRT-PCR data were assessed by one-way analysis of variance (ANOVA) using GraphPad Prism 9 software, with the significance level set at P ≤ 0.05. Different letters above the bars indicate statistically significant differences.
Identification of cis-regulatory elements
To predict cis-regulatory elements, promoter sequences (2,000 bp upstream of the start codon) of the PmVQ genes were obtained from the Genome Warehouse of the BIG Data Center. These sequences were subsequently analyzed using the PlantCARE (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/) tool to identify putative cis-regulatory elements. The distribution of the identified elements was visualized using TBtools to infer their potential regulatory functions.
Measurement of chlorophyll content and ion leakage rate
Chlorophyll content in proso millet leaves was determined using a SPAD chlorophyll meter (SPAD-502 Plus; Minolta, Tokyo, Japan). The relative change in chlorophyll content was calculated as follows: (SPAD value before treatment − SPAD value after treatment)/SPAD value before treatment.
For the ion leakage rate, detached leaves were immersed in 5 mL of deionized water and subjected to a vacuum for 30 min. The initial conductivity (C1) was then measured. Subsequently, the leaves were placed in a boiling water bath for 30 min, and after cooling to room temperature, the conductivity (C2) was measured again. The ion leakage rate was calculated using the formula (C1/C2) × 100%. The relative change in ion permeability was determined as (Ion permeability before treatment − Ion permeability after treatment)/Ion permeability before treatment.
For each proso millet material, Student’s t-tests were performed separately using GraphPad Prism 9 to analyze the statistical differences in SPAD values and ion leakage rates between post-drought-treatment and control measurements. Comparisons were not conducted across different materials. Significant differences are indicated by asterisks (*P < 0.05, ***P < 0.001).
Results
Identification and basic characteristics of the VQ gene family in proso millet
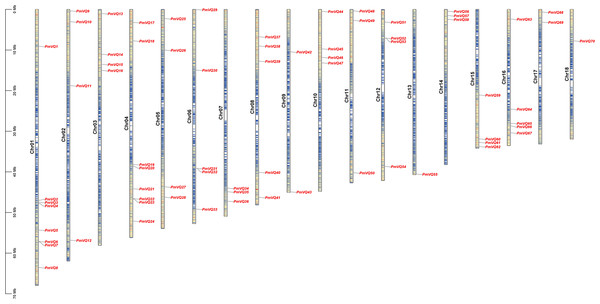
A total of 70 non-redundant VQ genes were identified in proso millet and named PmVQ1 to PmVQ70 according to their physical positions on the chromosomes. Chromosomal localization revealed that these genes were unevenly distributed across 18 chromosomes, with chromosomes 1 and 4 containing the most members (eight), while chromosomes 13 and 18 each contained only one member (Fig. 1). Physicochemical analysis showed that PmVQ proteins ranged from 85 to 447 amino acids in length, with molecular weights between 8.99 and 43.58 kDa and isoelectric points between 4.11 and 11.72. Subcellular localization predictions indicated that 48 PmVQ proteins were located in the nucleus, 16 resided in the chloroplast, and six were found in the mitochondria (Table S3).
Figure 1: Chromosomal distribution of the PmVQ gene family in proso millet.
The scale on the left indicates the base pair lengths (Mb) of the chromosomes. The color gradient on the chromosome from red to blue represents a decrease in gene density.Phylogenetic, gene structure, and conserved motif analysis of VQ gene family
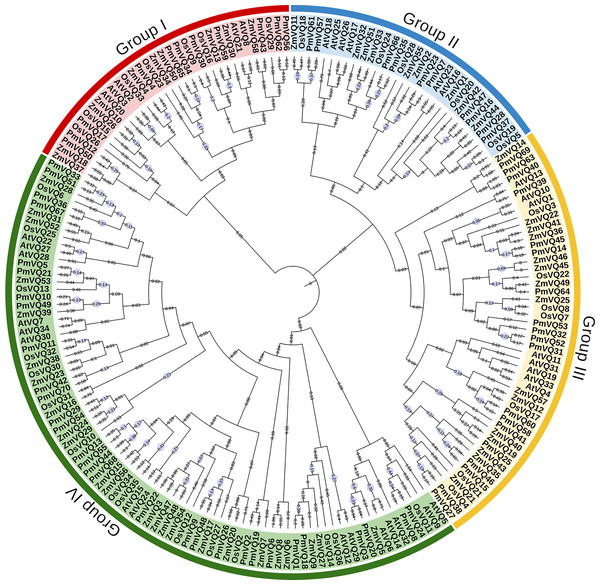
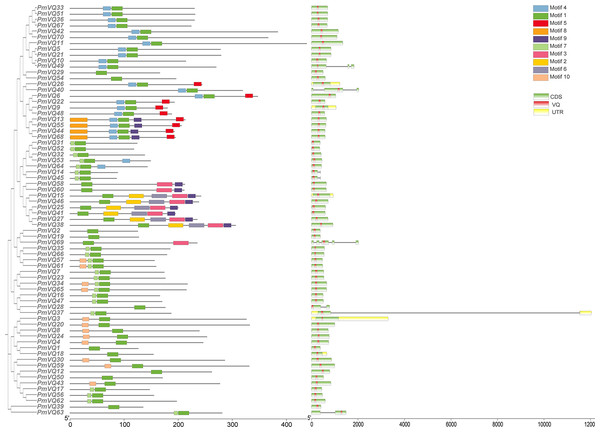
The phylogenetic tree, based on a total of 198 VQ protein sequences from four species (70 from proso millet, 34 from Arabidopsis, 36 from rice, and 58 from maize), reveals the evolutionary relationships within VQ gene family. All VQ proteins were divided into four subfamilies (Group I–IV). PmVQs were distributed across all subfamilies and showed closer relationships with homologs from Arabidopsis, maize and rice (Fig. 2). Gene structure analysis revealed that only six PmVQ genes contained introns, while the remaining 64 were intron-less. Conserved motif analysis identified ten motifs, with members within the same clade generally exhibiting similar motif compositions, all of which contained the conserved VQ domain (Fig. 3).
Figure 2: Phylogenetic analysis of VQ family genes.
A neighbor-joining tree was constructed based on the VQ genes identified from proso millet, Arabidopsis, rice, and maize. These genes were clustered into four major groups based on their evolutionary relationships, as indicated by the solid-line boxes in different colors.Figure 3: The gene structure, and conserved motifs of the PmVQ gene in proso millet.
Ten conserved motifs of the PmVQ gene family. Modules of different colors represent distinct conserved motifs. Gene structure and domains of PmVQ. In this representation, green boxes correspond to exons, black lines denote introns, yellow boxes indicate untranslated regions (UTRs), and red boxes represent the VQ domains.Collinearity analysis and VQ gene family evolution
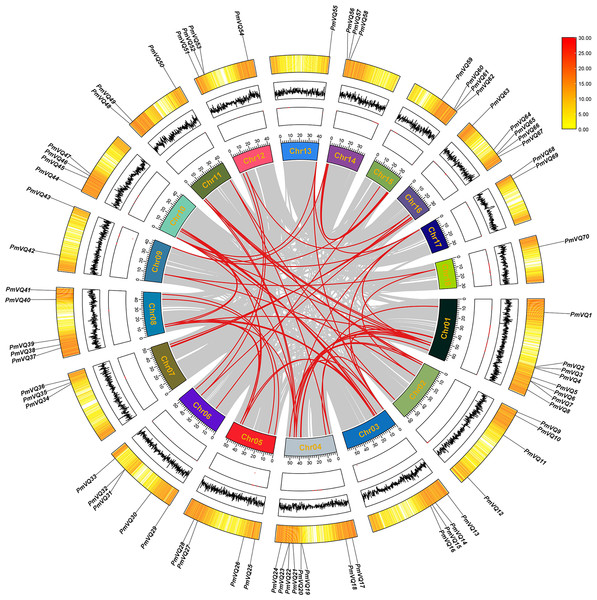
Chromosomal localization revealed an uneven distribution of the 70 PmVQ genes across the 18 chromosomes of proso millet. To investigate the expansion mechanism of PmVQ gene family, an intra-species collinearity analysis was performed. The results identified 63 duplicated PmVQ gene pairs in the proso millet genome, all of which were segmental duplications. Chromosome 4 exhibited the highest density of these gene pairs (15 pairs), while chromosome 18 had the lowest (only 2 pairs) (Fig. 4). In the PmVQ collinearity diagram, the outer ring displays the chromosomal map of proso millet, annotated with physical distance (Mb), GC content, and gene density. The inner ring uses colored arcs to connect collinear PmVQ gene pairs, with different colors representing distinct collinear blocks. Ka/Ks analysis indicated that 59 of the duplicated gene pairs had ratios less than 1.00, suggesting that they have predominantly undergone purifying selection. Collectively, these findings demonstrate that segmental duplication has been the primary driver of the expansion of the VQ gene family in proso millet (Table S4).
Figure 4: Intraspecies collinearity analysis of the PmVQ family genes.
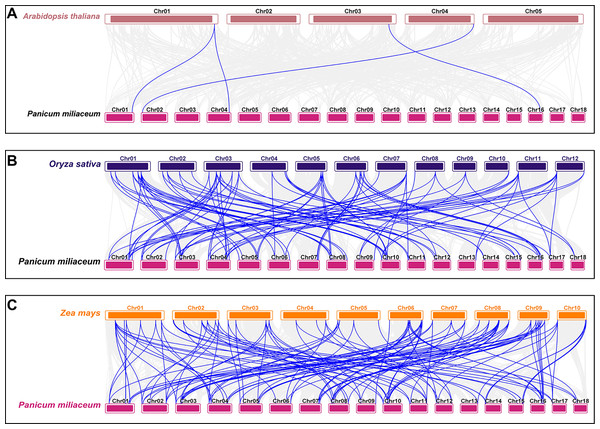
From the outermost to the innermost layers, the figure shows the gene density on chromosomes, the GC content, and the gap distribution in the genome. The red lines indicate collinear blocks within the whole genome.To investigate the origin and evolutionary mechanisms of the VQ genes in proso millet, a comparative collinearity analysis was conducted with Arabidopsis, rice, and maize. The analysis identified four orthologous gene pairs between proso millet and Arabidopsis, 99 with rice, and 129 with maize (Table S5). The evolutionary relationship of the PmVQ gene family is relatively conserved with maize but shows greater divergence with Arabidopsis from the Brassicaceae family (Fig. 5).
Figure 5: Interspecies collinearity analysis of VQ family genes between proso millet and other plant species.
(A) The interspecies collinearity analysis of the VQ genes between proso millet and Arabidopsis. (B) The interspecies collinearity analysis of the VQ genes between proso millet and rice. (C) The interspecies collinearity analysis of the VQ genes between proso millet and maize. The gray lines represent collinear blocks between the proso millet genome and the other species, while the blue lines indicate collinearity among VQ genes.Identification of cis-acting elements in the PmVQ gene promoters
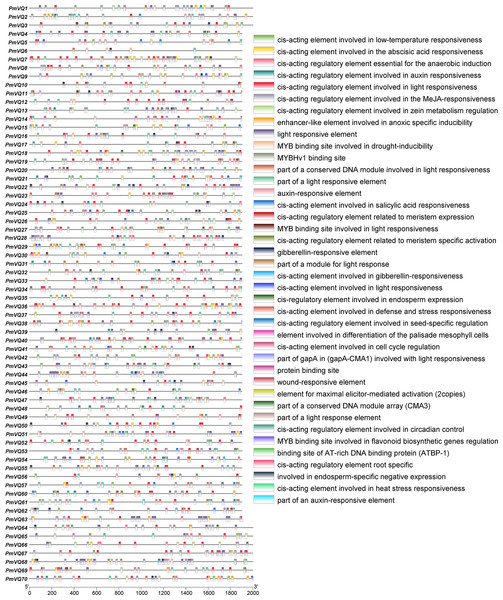
To investigate the potential role of PmVQ genes in drought stress response and regulation, cis-acting elements analysis was performed on the 2,000 bp promoter sequences upstream of the start codon. A total of 2,248 cis-acting elements were identified in the promoters of the 70 PmVQ genes, including 313 abscisic acid (ABA)-responsive elements, 60 drought-induced elements, and 20 stress-related elements. With the exception of PmVQ58, PmVQ59, and PmVQ70, the remaining 67 PmVQ genes contained at least one type of the stress-related elements described above (Fig. 6). Notably, 48 PmVQ genes harbored two or more drought-related elements, suggesting that most members of this gene family may play potential roles in drought stress response.
Figure 6: Promoter cis-regulatory elements in the PmVQ gene family.
The right panel displays the cis-regulatory elements identified in the promoter regions, while the left panel illustrates their positional distribution within the PmVQ genes promoters.Expression pattern analysis of PmVQ genes
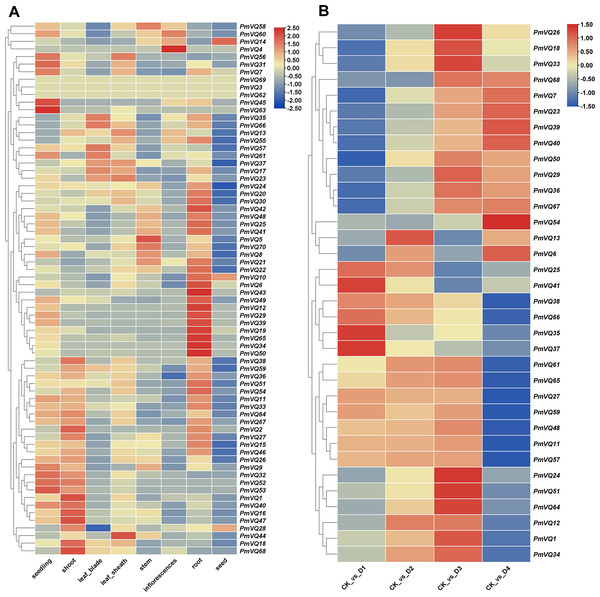
Given that roots are the primary organs for perceiving and responding to drought stress, tissue expression heatmap analysis revealed that 26 PmVQ genes exhibited root-specific or high expression in roots (Fig. 7A and Table S6). Based on transcriptome data from roots under PEG-simulated drought stress, 34 differentially expressed PmVQ genes were identified (Fig. 7B and Table S7). Integration with promoter cis-acting elements analysis led to the final screening of 14 candidate PmVQ genes that were significantly regulated by drought stress in roots, including 11 up-regulated and three down-regulated genes (Figs. 8A, 8B). These genes may function as early responders in root signaling pathways and play important roles in drought stress response.
Figure 7: Expression patterns of PmVQ family genes.
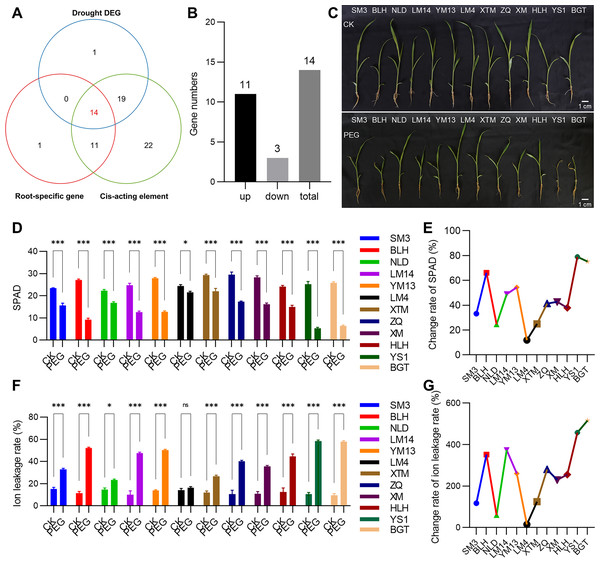
(A) Heatmap analysis of transcript levels of PmVQ genes across different tissues. (B) Drought stress response expression patterns of PmVQ genes. The heatmap based on log2 FC values displays differential expression between plants subjected to drought treatment at 0 h (CK), 0.5 h (D1), 1 h (D2), 1.5 h (D3), and 3 h (D4) relative to the control group.Figure 8: Drought tolerance evaluation of proso millet germplasm accessions.
(A) Venn diagram displaying PmVQ genes possessing drought-associated cis-regulatory elements, exhibiting root-specific high expression, and showing differential expression after drought treatment. (B) Numbers of upregulated and downregulated genes from the 14 candidate PmVQ genes in (A). (C) Phenotypic comparison of 12 proso millet germplasm accessions after 5 days of PEG-6000 treatment. (D and E) Chlorophyll content in the 12 proso millet germplasm accessions after PEG treatment and its relative change rate. (F and G) Ion leakage rate in the 12 proso millet germplasm accessions after PEG treatment and its relative change rate. Asterisks indicate significant differences. Student’s t-test, (*P < 0.05, ***P < 0.001).Validation of key PmVQ genes in drought stress response in roots
Based on a previously established collection of proso millet germplasm with varying drought tolerance, 12 representative accessions were selected for phenotypic analysis under PEG-simulated drought stress (5 days) (Fig. 8C). Significant variation in drought response was observed among the 12 proso millet accessions based on measurements of SPAD values, ion leakage rates, and their relative changes after drought stress treatment (Figs. 8D–8G). The drought-tolerant materials LM4 and NLD exhibited SPAD value decreases of 11.89% and 24.66%, respectively, and ion leakage increases of 14.96% and 59.60%, demonstrating stronger drought tolerance. In contrast, the sensitive material YS1 showed the most pronounced changes in physiological indicators, with a SPAD value decrease of 78.97% and an ion leakage increase of 457.71%, and was thus identified as a drought-sensitive accession.
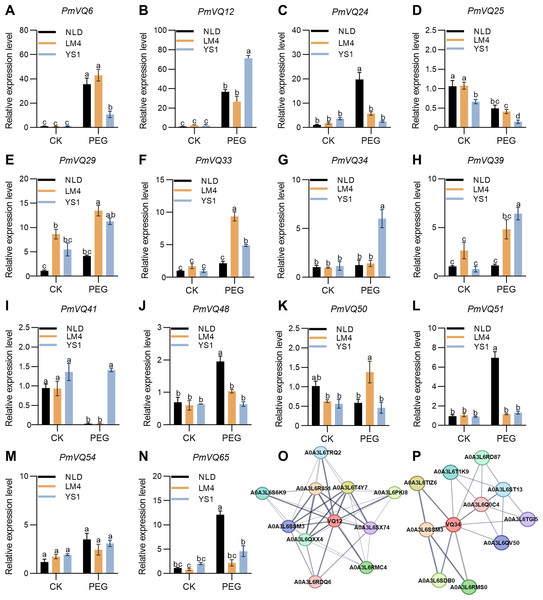
qRT-PCR analysis was further performed to examine the expression dynamics of 14 candidate genes in the roots of LM4, NLD, and YS1 before and after PEG treatment. The results showed distinct expression patterns for four genes—PmVQ6, PmVQ12, PmVQ34, and PmVQ41—between drought-tolerant and -sensitive materials. After drought treatment, the transcript level of PmVQ6 was significantly up-regulated, with markedly higher expression in tolerant materials than in sensitive ones. Conversely, the expression of PmVQ12, PmVQ34, and PmVQ41 was significantly lower in tolerant materials. The transcript levels of the other genes showed no significant differences between the drought-tolerant and drought-sensitive materials before and after drought treatment (Figs. 9A–9N). These findings collectively demonstrate that these four genes are key candidates involved in the drought stress response in proso millet roots.
Figure 9: Expression patterns of the 14 PmVQ genes after drought treatment.
(A–N) Transcript levels of PmVQ genes in drought-tolerant accessions (NLD and LM4) and drought-sensitive accession (YS1) after drought treatment. (O, P) Prediction of potential interacting proteins for PmVQ12 and PmVQ34. All experiments were repeated at least three times with similar results. Different letters above the columns indicate significant differences as determined by one‐way analysis of variance (P < 0.05).Subsequently, the potential protein-protein interaction networks of PmVQ12 and PmVQ34 were predicted using STRING. Based on the prediction results from the STRING database (https://cn.string-db.org/cgi/input), PmVQ12 may interact with MAPK and WRKY transcription factors, while PmVQ34 is potentially involved in regulatory pathways mediated by WRKY proteins (Figs. 9O, 9P). The prediction results revealed a shared interacting protein, A0A3L6SSM3, between PmVQ12 and PmVQ34, which encodes a protein containing a conserved WRKY domain. The results suggesting that the drought response and regulatory process in proso millet involves a complex network.
Discussion
A total of 70 VQ genes were identified in the proso millet genome, and these genes were unevenly distributed across all chromosomes. The number of VQ genes in proso millet was significantly greater than that in Arabidopsis (34), rice (36), and maize (58). Phylogenetic analysis classified the VQ genes from proso millet, Arabidopsis, rice, and maize into four major clades. The VQ genes within the same clade exhibited close evolutionary relationships, reflecting the evolutionary conservation of this gene family across species and suggesting that PmVQ genes may possess more diverse functions. Among the 70 PmVQ genes, 64 were intronless, a feature consistent with previous study (Jing & Lin, 2015).
Whole-genome duplication, tandem duplication, and segmental duplication are important evolutionary mechanisms driving the expansion and functional diversification of gene families. In this study, a total of 63 duplication events were identified in the proso millet VQ gene family, all of which were segmental duplications, indicating that segmental duplication served as the primary driving force for the expansion of the PmVQ gene family. Among the PmVQ homologous gene pairs, 59 pairs exhibited Ka/Ks ratios of less than 1.00, suggesting that PmVQ gene family has been under strong purifying selection during evolution.
The 70 PmVQ genes were widely distributed across all 18 chromosomes of proso millet, with a notable enrichment in telomeric regions. Intra-genomic collinearity analysis revealed that chromosome 4 contained the densest distribution of collinear gene pairs. Cross-species collinearity comparisons indicated the presence of 4, 99, and 129 collinear gene pairs between proso millet and Arabidopsis, rice, and maize, respectively. These conserved collinear relationships underscore the important role of VQ genes in maintaining the structural and functional stability of the gene family. Most orthologous relationships did not follow the typical one-to-one correspondence pattern, implying multiple rounds of gene duplication and subsequent functional differentiation during evolution. Notably, the extensive collinear regions observed between proso millet and maize further support their close evolutionary relationship within the VQ gene family.
In this study, four key PmVQ genes (PmVQ6, PmVQ12, PmVQ34, and PmVQ41) were identified through analysis of promoter cis-regulatory elements, tissue-specific expression, and differentially expressed genes between drought stress and control. These genes share common sequence characteristics but also exhibit notable differences in expression patterns.
Phylogenetic analysis revealed that the four key candidate genes, PmVQ6, PmVQ12, PmVQ34, and PmVQ41, are distributed across distinct evolutionary subgroups (Group IV, Group I, Group I, and Group III, respectively). This distribution pattern suggests that they may participate in divergent regulatory networks and contribute to sub-functional differentiation in drought response. Specifically, although PmVQ12 and PmVQ34, both belonging to Group I, are closely related phylogenetically, they exhibit different expression trends under drought stress (PmVQ12 is upregulated in both drought-tolerant and drought-sensitive materials, whereas PmVQ34 is specifically induced only in drought-sensitive materials). This may reflect complementary or antagonistic functional roles within the same regulatory module. In contrast, PmVQ41, which belongs to Group III, is significantly down-regulated in drought-tolerant materials, a pattern distinct from that of Group I members, implying that it may be involved in drought adaptation through independent or intersecting signaling pathways. Notably, PmVQ6, the sole representative from Group IV, displays the strongest drought-induced expression in drought-tolerant materials, suggesting that this subgroup may have evolved specialized functions in drought tolerance regulation. The observed expression divergence across subgroups, combined with the diversity in promoter cis-regulatory elements (PmVQ6 contains only drought-responsive elements, while PmVQ34 specifically harbors ABA-responsive elements), further supports the notion that the PmVQ gene family achieves precise regulation of drought response through sub-functional differentiation and integration of multiple signaling pathways.
The four key PmVQ genes exhibited clearly divergent expression patterns between drought-tolerant and drought-sensitive materials in root. Under drought treatment, both PmVQ6 and PmVQ12 were significantly up-regulated. However, PmVQ6 showed stronger induction in resistant materials than in sensitive ones, whereas PmVQ12 was more highly induced in sensitive materials. PmVQ34 was strongly induced only in sensitive materials after drought stress, with no significant change in resistant materials. In contrast, PmVQ41 was significantly down-regulated only in resistant materials. Based on the expression profiles presented above, we propose a preliminary hypothesis that PmVQ6 may function as a positive regulator of drought tolerance in proso millet, whereas PmVQ12, PmVQ34, and PmVQ41 may act as negative regulators. This inference remains to be validated through subsequent experimentation.
Based on the findings of this study and previous research, we speculate that VQ genes in proso millet may be involved in the drought stress response through multiple mechanisms. First, the presence of transcription factor binding sites such as MYB and MYC in their promoter regions suggests that these genes may be directly regulated by key transcription factors in drought signaling pathways (Lei, Ma & Yu, 2018). Second, as transcriptional regulatory factors, VQ proteins may form complexes with WRKY transcription factors to jointly regulate the expression of downstream stress-responsive genes. Previous studies have demonstrated that specific VQ–WRKY modules play critical roles in stress signal transduction in model plants (Cheng et al., 2012; Wang et al., 2015). Furthermore, the root-preferential expression pattern of these genes implies that they may enhance drought tolerance by modulating root development or rhizosphere signaling perception.
Protein-protein interaction predictions revealed that both PmVQ12 and PmVQ34 interact with a common protein, “A0A3L6SSM3”. This protein contains a conserved WRKY domain and is homologous to OsWRKY24 in rice and AtWRKY26 in Arabidopsis. Functional studies have shown that OsWRKY24 is not only involved in regulating developmental traits in rice, such as grain size, leaf angle, and plant height, but also participates in various physiological processes through the brassinosteroid (BR) signaling pathway and the MAPK cascade (Jang & Li, 2018; Tang et al., 2022). In contrast, the expression of AtWRKY26 is modulated by diverse environmental stimuli, including heat stress, salt, abscisic acid, and osmotic stress (Fu & Yu, 2010). Therefore, by interacting with a common PmWRKY transcription factor, PmVQ12 and PmVQ34 may coordinately regulate drought tolerance in proso millet, potentially through both the MAPK signaling cascade and the integration of plant hormone signaling pathways. These predictions provide direct support for the hypothesis that VQ proteins participate in signal transduction by forming diverse protein complexes. The distinct expression dynamics of the four key VQ genes further suggest that they may constitute a multi-layered and temporally coordinated drought response network, in which early-responsive genes are responsible for rapid signal perception and transduction, while late-responsive genes contribute to long-term adaptive regulation.
The research on the VQ gene family in proso millet has revealed a complex regulatory network involving key genes such as PmVQ6, PmVQ12, PmVQ34, and PmVQ41, which coordinately regulate drought stress responses. The differential expression of these genes is a critical factor underlying the variation in drought tolerance among germplasm accessions. Future research should not only focus on elucidating their precise functions and roles in signaling pathways but also prioritize advancing their breeding applications. This includes developing functional molecular markers linked to superior alleles for efficient genotype-assisted selection, using gene-editing technologies to precisely modulate key genes and create novel germplasms with desired drought-resistant traits, and deciphering their interaction networks with factors such as WRKY to enable multi-gene pyramiding and modular design for coordinated improvement of drought tolerance and related agronomic traits. Ultimately, integrating multi-omics analyses and large-scale germplasm characterization will establish a systematic breeding framework linking genes to phenotypes. This will drive proso millet breeding from traditional empirical selection toward molecular design, providing both theoretical insights and technological support for developing high-yielding, water-efficient cultivars adapted to climate change.
Conclusions
A total of 70 VQ genes were identified in the proso millet genome and classified into four distinct subfamilies. The expansion of this gene family was primarily driven by segmental duplication events, followed by strong purifying selection during evolution. Promoter cis-regulatory element analysis and transcriptomic profiling revealed widespread involvement of PmVQ genes in drought stress responses, leading to the identification of 14 root-specific, drought-responsive candidate genes. Systematic phenotyping of multiple germplasm accessions combined with qRT-PCR validation further identified four key candidate genes (PmVQ6, PmVQ12, PmVQ34, and PmVQ41). These genes exhibited high basal expression levels in the roots and showed markedly divergent expression patterns between drought-tolerant and drought-sensitive materials under drought stress. This suggests their potential key role in mediating drought response and tolerance regulation in proso millet via root-specific mechanisms. Furthermore, they are likely involved in a complex regulatory network through interactions with proteins such as MAPK and WRKY, thereby coordinately fine-tuning the drought tolerance mechanisms in proso millet. This study lays a solid foundation for systematically elucidating the molecular functions of VQ genes in proso millet drought tolerance, particularly in the root-specific response to and regulation of drought stress, and provides valuable genetic resources for developing drought-tolerant cultivars.
This study lays a foundation for elucidating the functions of VQ genes in proso millet, particularly their root-specific regulatory roles in drought tolerance. In the future, functional molecular markers can be developed based on the key genes identified, enabling the establishment of an efficient molecular-assisted breeding system. Drought-tolerant germplasms can be further developed through multi-gene pyramiding breeding strategies. Concurrently, comparative genomic studies of VQ gene families in Poaceae crops will help uncover the evolutionary characteristics of drought resistance-related genes, providing a theoretical basis for the cross-species utilization of stress resistance genes. These efforts will deepen the understanding of drought tolerance mechanisms in proso millet and offer new genetic resources and strategies for improving stress tolerance in Poaceae crops.
Supplemental Information
The details of the 12 proso millet germplasm accessions.
The details of the 12 proso millet germplasm accessions including full names, abbreviations, Chinese names, collection locations, and collection dates.
qRT‐PCR primers.
The names and sequences of primers used for qRT-PCR in this study.
The subcellular localization and physicochemical analysis of PmVQs proteins.
Predicted results of coding protein length, molecular size, isoelectric point, and subcellular localization for 70 PmVQ genes.
Ka/Ks analysis of PmVQ gene family.
The KA/KS ratios of VQ genes with interspecific collinearity in promo millet.
Homologous gene pairs from inter-species collinearity analysis.
The names of the collinear gene pairs identified in the interspecific collinearity analysis between broomcorn millet and Arabidopsis, rice, and maize.
Raw data of transcriptome data from different tissues of proso millet, and the expression of VQ genes in different tissues.
Raw data of the transcriptome data for different tissues of proso millet and the pmVQ gene are provided in Sheet1 and Sheet2, respectively.
Raw data of up- and down-regulated differentially expressed genes under drought treatment in root.
Raw data of differential expression genes (DEGs) in proso millet roots under drought stress at various time points.
The raw data of relative expression level.
Relative expression data from qRT-PCR in the drought treatment experiment.